Abstract
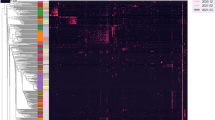
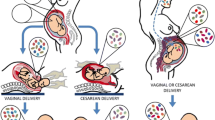
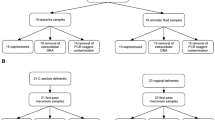
The widely accepted dogma of intrauterine sterility and initial colonization of the newborn during birth has been blurred by recent observations of microbial presence in meconium, placenta, and amniotic fluid. Given the importance of a maternal-derived in utero infant seeding, it is crucial to exclude potential environmental or procedural contaminations and to assess fetal colonization before parturition. To this end, we analyzed sterilely collected intestinal tissues, placenta, and amniotic fluid from rodent fetuses and tissues from autoptic human fetuses. Total bacterial DNA was extracted from collected samples and analyzed by Next Generation Sequencing (NGS) techniques using hypervariable 16S ribosomal RNA (rRNA) regions (V3-V4). Colonizing microbes were visualized in situ, using labeled probes targeting 16S ribosomal DNA by fluorescent in situ hybridization. The NGS analysis showed the presence of pioneer microbes in both rat and human intestines as well as in rodent placentas and amniotic fluids. Microbial communities showed fetus- and dam-dependent clustering, confirming the high interindividual variability of commensal microbiota even in the antenatal period. Fluorescent in situ hybridization analysis confirmed the microbes’ presence in the lumen of the developing gut. These findings suggest a possible antenatal colonization of the developing mammalian gut.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Brugman S, Perdijk O, van Neerven RJJ, Savelkoul HFJ. Mucosal immune development in early life: setting the stage. Arch Immunol Ther Exp. 2015;63(4):251–268.
Ardissone AN, de la Cruz DM, Davis-Richardson AG, et al. Meconium microbiome analysis identifies bacteria correlated with premature birth. PLoS One. 2014;9(3):e90784.
Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J. The placenta harbors a unique microbiome. Sci Transl Med. 2014;6(237):237ra65.
Funkhouser LJ, Bordenstein SR. Mom knows best: the universality of maternal microbial transmission. PLoS Biol. 2013;11(8):1–9.
Jiménez E, Fernández L, Marín ML, et al. Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr Microbiol. 2005;51(4):270–274.
Collado MC, Rautava S, Aakko J, Isolauri E, Salminen S. Human gut colonisation may be initiated in utero by distinct microbial communities in the placenta and amniotic fluid. Sci Rep. 2016;6:23129.
Prince AL, Ma J, Kannan PS, et al. The placental membrane microbiome is altered among subjects with spontaneous preterm birth with and without chorioamnionitis. Am J Obstet Gynecol. 2016;214(5):627.e1–627.e16.
Jiménez E, Marín ML, Martín R, et al. Is meconium from healthy newborns actually sterile? Res Microbiol. 2008;159(3):187–193.
Avagliano L, Marconi AM, Candiani M, Barbera A, Bulfamante G. Thrombosis of the umbilical vessels revisited. An observational study of 317 consecutive autopsies at a single institution. Hum Pathol. 2010;41(7):971–979.
Belkaid Y, Hand TW. Role of the microbiota in immunity and inflammation. Cell. 2014;157(1):121–141.
Fardini Y, Chung P, Dumm R, Joshi N, Han YW. Transmission of diverse oral bacteria to murine placenta: evidence for the oral microbiome as a potential source of intrauterine infection. Infect Immun. 2010;78(4):1789–1796.
Perez-Munõz ME, Arrieta MC, Ramer-Tait AE, Walter J. A critical assessment of the ‘sterile womb’ and ‘in utero colonization’ hypotheses: implications for research on the pioneer infant microbiome. Microbiome. 2017;5(1):48.
Chu DM, Ma J, Prince AL, Antony KM, Seferovic MD, Aagaard KM. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat Med. 2017;23(3):314–326.
Huttenhower C; Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486(7402):207–214.
Qin J, Li R, Raes J, et al. A human gut microbial gene catalogue established by metagenomic sequencing: commentary. Nature. 2010;11:28.
Greenhalgh K, Meyer KM, Aagaard KM, Wilmes P. The human gut microbiome in health: establishment and resilience of microbiota over a lifetime. Environ Microbiol. 2016;18(7):2103–2116.
Riva A, Borgo F, Lassandro C, et al. Pediatric obesity is associated with an altered gut microbiota and discordant shifts in Firmicutes populations. Environ Microbiol. 2016;19(1):1–29.
Turroni F, Ribbera A, Foroni E, van Sinderen D, Ventura M. Human gut microbiota and bifidobacteria: from composition to functionality. Antonie Van Leeuwenhoek. 2008;94(1):35–50.
Scott KP, Gratz SW, Sheridan PO, Flint HJ, Duncan SH. The influence of diet on the gut microbiota. Pharmacol Res. 2012;69(1):52–60.
Koenig JE, Spor A, Scalfone N, et al. Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci U S A. 2011;108(suppl 1):4578–4585.
Asakuma S, Hatakeyama E, Urashima T, et al. Physiology of consumption of human milk oligosaccharides by infant gutassociated bifidobacteria. J Biol Chem. 2011;286(40):34583–34592.
Wang M, Li M, Wu S, et al. Fecal microbiota composition of breast-fed infants is correlated with human milk oligosaccharides consumed. J Pediatr Gastroenterol Nutr. 2015;60(6):825–833.
Rasmussen SO, Martin L, Østergaard MV, et al. Human milk oligosaccharide effects on intestinal function and inflammation after preterm birth in pigs. J Nutr Biochem. 2017;40:141–154.
Drago L, Toscano M, De Grandi R, Grossi E, Padovani EM, Peroni DG. Microbiota network and mathematic microbe mutualism in colostrum and mature milk collected in two different geographic areas: Italy versus Burundi. ISME J. 2016;11(4):1–10.
Doran KS, Banerjee A, Disson O, Lecuit M. Concepts and mechanisms: crossing host barriers. Cold Spring Harb Perspect Med. 2013;3(7):a010090.
Hullar MA, Fu BC. Diet, the gut microbiome, and epigenetics. Cancer J. 2014;20(3):170–175.
De Filippo C, Cavalieri D, Di Paola M, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A. 2010;107(33):14691–14696.
Strati F, Di Paola M, Stefanini I, et al. Age and gender affect the composition of fungal population of the human gastrointestinal tract. Front Microbiol. 2016;7:1227.
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R. Diversity, stability and resilience of the human gut microbiota. Nature. 2012;489(7415):220–230.
Aagaard K, J Petrosino J, Keitel W, et al. The human microbiome project strategy for comprehensive sampling of the human microbiome and why it matters. FASEB J. 2013;27(3):1012–1022.
Ma J, Coarfa C, Qin X, et al. mtDNA haplogroup and single nucleotide polymorphisms structure human microbiome communities. BMC Genomics. 2014;15:257.
Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. 2012;148(6):1258–1270.
Peterson CT, Sharma V, Elmén L, Peterson SN. Immune homeostasis, dysbiosis and therapeutic modulation of the gut microbiota. Clin Exp Immunol. 2015;179(3):363–377.
Mueller NT, Bakacs E, Combellick J, Grigoryan Z, Dominguez-Bello MG. The infant microbiome development: mom matters. Trends Mol Med. 2015;21(2):109–117.
Mecacci F, Serena C, Avagliano L, et al. Stillbirths at term: case control study of risk factors, growth status and placental histology. PLoS One. 2016;11(12):e0166514.
Salter SJ, Cox MJ, Turek EM, et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014;12:87.
Greuter D, Loy A, Horn M, Rattei T. ProbeBase-an online resource for rRNA-targeted oligonucleotide probes and primers: new features 2016. Nucleic Acids Res. 2016;44(D1):D586–D589.
Gey A, Werckenthin C, Poppert S, Straubinger RK. Identification of pathogens in mastitis milk samples with fluorescent in situ hybridization. J Vet Diagn Invest. 2013;25(3):386–394.
Berry D, Schwab C, Milinovich G, et al. Phylotype-level 16S rRNA analysis reveals new bacterial indicators of health state in acute murine colitis. ISME J. 2012;6(11):2091–2106.
Borghi E, Borgo F, Severgnini M, Savini MN, Casiraghi MC, Vignoli A. Rett syndrome: a focus on gut microbiota. Int J Mol Sci. 2017;18(2):E344.
Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD. PANDAseq: paired-end assembler for Illumina sequences. BMC Bioinformatics. 2012;13:31.
Caporaso JG, Kuczynski J, Stombaugh J, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7(5):335–336.
Lozupone C, Knight R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol. 2005;71(12):8228–8235.
Acknowledgments
We thank Raffaella Adami for her assistance in confocal laser microscopy techniques, Francesca Di Renzo and Silvia Rigamonti for technical assistance. We are grateful to Ms Dawn Savery and Dr Jon Wilson for commenting on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Borghi, E., Massa, V., Severgnini, M. et al. Antenatal Microbial Colonization of Mammalian Gut. Reproductive Sciences 26, 1045–1053 (2019). https://doi.org/10.1177/1933719118804411
Published:
Issue Date:
DOI: https://doi.org/10.1177/1933719118804411