Abstract
Objective
Epithelial ovarian cancer (EOC) cells with CD44 and CK19 coexpression may represent a subset of ovarian cancer stem cells (OCSCs). This study was conducted to evaluate the correlation of the frequency of putative OCSCs (CD44 + CK19 + OCSCs) with the clinicopathologic features and the prognostic value in patients with recurrent advanced stage EOC.
Methods
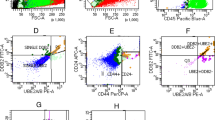
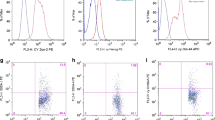
A retrospective study was carried out on 33 patients with EOC and a uniformly treated tissue microarray was constructed. A multiplexed, immunofluorescence-based method of automated in situ quantitative measurement of protein analysis was used for evaluation of the frequency or density of CD44 + CK19 + OCSCs in EOC.
Results
The mean follow-up time was 42.8 ± 27.1 months. High frequency of EOC cells with CD44+ or CD44+/CK19+ was associated with chemoresistance (P = .033 and P = .02, respectively). Using K-M analysis with log-rank test, a high frequency of putative OCSCs was associated with short disease-free interval (7.9 months vs 20.9 months, P = .019). In univariable analysis, the frequency of OCSCs, International Federation of Gynecology and Obstetrics stage and residual tumor volume were significant predictor variables and were entered into multivariable analysis (P = .019, .037, and .005, respectively). Although no independent significant predictor was found, the frequency of putative OCSCs was the most promising predictor variable compared with the other 2 variables (hazard ratio = 2.344, P = .052).
Conclusion
Our findings suggest that high frequency of OCSCs (CD44+ and CK19+) in epithelial ovarian tumors correlates with short progression-free intervals.
Similar content being viewed by others
References
Holschneider CH, Berek JS. Ovarian cancer: epidemiology, biology, and prognostic factors. Semin Surg Oncol. 2000;19(1):3–10.
Nossov V, Amneus M, Su F, et al The early detection of ovarian cancer: from traditional methods to proteomics. Can we really do better than serum CA-125? Am J Obstet Gynecol. 2008;199(3):215–223.
Jemal A, Siegel R, Xu J, Ward E. Cancer statistics, 2010. CA Cancer J Clin. 2010;60(5):277–300.
Garvalov BK, Acker T. Cancer stem cells: a new framework for the design of tumor therapies. J Mol Med (Berl). 2011;89(2):95–107.
O’Brien CA, Kreso A, Dick JE. Cancer stem cells in solid tumors: an overview. Semin Radiat Oncol. 2009;19(2):71–77.
Bao S, Wu Q, McLendon RE, et al Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature. 2006;444(7120):756–760.
Hermann PC, Huber SL, Herrler T, et al Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell. 2007;1(3):313–323.
Alvero AB, Chen R, Fu HH, et al Molecular phenotyping of human ovarian cancer stem cells unravels the mechanisms for repair and chemoresistance. Cell Cycle. 2009;8(1):158–166.
Mor G, Yin G, Chefetz I, Yang Y, Alvero A. Ovarian cancer stem cells and inflammation. Cancer Biol Ther. 2011;11(8):708–713.
Alvero AB, Fu HH, Holmberg J, et al Stem-like ovarian cancer cells can serve as tumor vascular progenitors. Stem Cells. 2009;27(10):2405–2413.
Curley MD, Therrien VA, Cummings CL, et al CD133 expression defines a tumor initiating cell population in primary human ovarian cancer. Stem Cells. 2009;27(12):2875–2883.
Kryczek I, Liu S, Roh M, et al Expression of aldehyde dehydrogenase and CD133 defines ovarian cancer stem cells. Int J Cancer. 2012;130(1):29–39.
Gao MQ, Choi YP, Kang S, Youn JH, Cho NH. CD24+ cells from hierarchically organized ovarian cancer are enriched in cancer stem cells. Oncogene. 2010;29(18):2672–2680.
Luo L, Zeng J, Liang B, et al Ovarian cancer cells with the CD117 phenotype are highly tumorigenic and are related to chemotherapy outcome. Exp Mol Pathol. 2011;91(2):596–602.
Naor D, Sionov RV, Ish-Shalom D. CD44: structure, function, and association with the malignant process. Adv Cancer Res. 1997;71:241–319.
Shipitsin M, Campbell LL, Argani P, et al Molecular definition of breast tumor heterogeneity. Cancer Cell. 2007;11(3):259–273.
Yin G, Chen R, Alvero AB, et al TWISTing stemness, inflammation and proliferation of epithelial ovarian cancer cells through MIR199A2/214. Oncogene. 2010;29(24):3545–3553.
Zoller M. CD44: can a cancer-initiating cell profit from an abundantly expressed molecule? Nat Rev Cancer. 2011;11(4):254–267.
Steffensen KD, Alvero AB, Yang Y, et al Prevalence of epithelial ovarian cancer stem cells correlates with recurrence in early-stage ovarian cancer. J Oncol. 2011;2011:620523.
Lim R, Lappas M, Ahmed N, Permezel M, Quinn MA, Rice GE. 2D-PAGE of ovarian cancer: analysis of soluble and insoluble fractions using medium-range immobilized pH gradients. Biochem Biophys Res Commun. 2011;406(3):408–413.
McCabe A, Dolled-Filhart M, Camp RL, Rimm DL. Automated quantitative analysis (AQUA) of in situ protein expression, antibody concentration, and prognosis. J Natl Cancer Inst. 2005;97(24):1808–1815.
Kluger HM, Dolled-Filhart M, Rodov S, Kacinski BM, Camp RL, Rimm DL. Macrophage colony-stimulating factor-1 receptor expression is associated with poor outcome in breast cancer by large cohort tissue microarray analysis. Clin Cancer Res. 2004;10(1 pt 1):173–177.
Psyrri A, Kountourakis P, Yu Z, et al Analysis of p53 protein expression levels on ovarian cancer tissue microarray using automated quantitative analysis elucidates prognostic patient subsets. Ann Oncol. 2007;18(4):709–715.
Camp RL, Chung GG, Rimm DL. Automated subcellular localization and quantification of protein expression in tissue microarrays. Nat Med. 2002;8(11):1323–1327.
Gustavson MD, Bourke-Martin B, Reilly D, et al Standardization of HER2 immunohistochemistry in breast cancer by automated quantitative analysis. Arch Pathol Lab Med. 2009;133(9):1413–1419.
Gustavson MD, Bourke-Martin B, Reilly DM, et al Development of an unsupervised pixel-based clustering algorithm for compartmentalization of immunohistochemical expression using Automated QUantitative Analysis. Appl Immunohistochem Mol Morphol. 2009;17(4):329–337.
Camp RL, Chung GG, Rimm DL. Automated subcellular localization and quantification of protein expression in tissue microarrays. Nat Med. 2002;8(11):1323–1327.
Gustavson MD, Molinaro AM, Tedeschi G, Camp RL, Rimm DL. AQUA analysis of thymidylate synthase reveals localization to be a key prognostic biomarker in 2 large cohorts of colorectal carcinoma. Arch Pathol Lab Med. 2008;132(11):1746–1752.
Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bioinformatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10(21):7252–7259.
Rustin GJ, Quinn M, Thigpen T, et al Re: New guidelines to evaluate the response to treatment in solid tumors (ovarian cancer). J Natl Cancer Inst. 2004;96(6):487–488.
Rustin GJ. Can we now agree to use the same definition to measure response according to CA-125? J Clin Oncol. 2004;22(20):4035–4036.
Neumeister V, Agarwal S, Bordeaux J, Camp RL, Rimm DL. In situ identification of putative cancer stem cells by multiplexing ALDH1, CD44, and cytokeratin identifies breast cancer patients with poor prognosis. Am J Pathol. 2010;176(5):2131–2138.
Jain R, Fischer S, Serra S, Chetty R. The use of Cytokeratin 19 (CK19) immunohistochemistry in lesions of the pancreas, gastrointestinal tract, and liver. Appl Immunohistochem Mol Morphol. 2010;18(1):9–15.
Lacroix M. Significance, detection and markers of disseminated breast cancer cells. Endocr Relat Cancer. 2006;13(4):1033–1067.
Mertz KD, Demichelis F, Sboner A, et al Association of cytokeratin 7 and 19 expression with genomic stability and favorable prognosis in clear cell renal cell cancer. Int J Cancer. 2008;123(3):569–576.
Ponta H, Sherman L, Herrlich PA. CD44: from adhesion molecules to signalling regulators. Nat Rev Mol Cell Biol. 2003;4(1):33–45.
Marhaba R, Klingbeil P, Nuebel T, Nazarenko I, Buechler MW, Zoeller M. CD44 and EpCAM: cancer-initiating cell markers. Curr Mol Med. 2008;8(8):784–804.
Yin G, Alvero AB, Craveiro V, et al Constitutive proteasomal degradation of TWIST-1 in epithelial-ovarian cancer stem cells impacts differentiation and metastatic potential [published online February 20, 2012]. Oncogene. 2012.
Casagrande F, Cocco E, Bellone S, et al Eradication of chemotherapy-resistant CD44+ human ovarian cancer stem cells in mice by intraperitoneal administration of clostridium perfringens enterotoxin. Cancer. 2011;117(24):5519–5528.
Kayastha S, Freedman AN, Piver MS, Mukkamalla J, Romero-Guittierez M, Werness BA. Expression of the hyaluronan receptor, CD44S, in epithelial ovarian cancer is an independent predictor of survival. Clin Cancer Res. 1999;5(5):1073–1076.
Rodriguez-Rodriguez L, Sancho-Torres I, Mesonero C, Gibbon DG, Shih WJ, Zotalis G. The CD44 receptor is a molecular predictor of survival in ovarian cancer. Med Oncol. 2003;20(3):255–263.
Berner HS, Davidson B, Berner A, et al Expression of CD44 in effusions of patients diagnosed with serous ovarian carcinoma—diagnostic and prognostic implications. Clin Exp Metastasis. 2000;18(2):197–202.
Sillanpaa S, Anttila MA, Voutilainen K, et al CD44 expression indicates favorable prognosis in epithelial ovarian cancer. Clin Cancer Res. 2003;9(14):5318–5324.
Dolled-Filhart M, Gustavson M, Camp RL, Rimm DL, Tonkinson JL, Christiansen J. Automated analysis of tissue microarrays. Methods Mol Biol. 2010;664:151–162.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, M., Mor, G., Cheng, H. et al. High Frequency of Putative Ovarian Cancer Stem Cells With CD44/CK19 Coexpression Is Associated With Decreased Progression-Free Intervals In Patients With Recurrent Epithelial Ovarian Cancer. Reprod. Sci. 20, 605–615 (2013). https://doi.org/10.1177/1933719112461183
Published:
Issue Date:
DOI: https://doi.org/10.1177/1933719112461183