Abstract
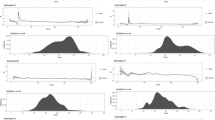
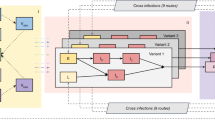
This study proposes a mathematical model to analyze the transmission mechanism of SARS-CoV-2 using a reaction-diffusion framework. The importance of incorporating diffusion in mathematical models of SARS-CoV-2 is emphasized, as it can lead to a more accurate understanding of disease spread and inform effective public health interventions. The impact of diffusion effects on the SARS-CoV-2 epidemic model has been significant, as the virus has rapidly spread across the globe since it was first identified in late 2019. The speed and efficiency of transmission have been a key factor in the severity of the pandemic, as many individuals may be asymptomatic or experience mild symptoms, making it difficult to identify and control the spread of the virus. The model includes six classes, namely Susceptible, Exposed to strain 1 SARS-CoV-2 and strain 2 SARS-CoV-2, infected by strain 1 SARS-CoV-2 and strain 2 SARS-CoV-2, and Recovered or Removed (\(\mathbb {S}\) \(\mathbb {E}_1\) \(\mathbb {E}_2\) \(\mathbb {I}_1\) \(\mathbb {I}_2\) \(\mathbb {R}\)), which are dependent on both time and space. This study uses the next-generation matrix approach to calculate the threshold number \(R_0\) and estimates parameter values through the use of least squares curve fitting tools. A combination of the operator splitting approach, finite difference method, and Crank–Nicolson method is used to simulate the model. The study looks at how stable the disease-free and endemic equilibrium points are, and it performs sensitivity analysis to look at the effects of different parameters. The simulation results of the model are compared in detail with and without diffusion and are verified through mutual comparison and theoretical analysis to ensure the accuracy of the solution. This study provides valuable insights into the transmission mechanism of SARS-CoV-2 and has important implications for public health policy.









Similar content being viewed by others
Data Availability Statement
This manuscript has associated data in a data repository. [Authors’ comment: All data generated or analyzed during this study are included in this article].
References
B. Hu, H. Guo, P. Zhou, Z.L. Shi, Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 19, 141–154 (2021)
F. Rahimi, A.T.B. Abadi, Is Omicron the last SARS-CoV-2 Variant of Concern? Archiv. Med. Res. (2022). https://doi.org/10.1016/j.arcmed.2022.01.001
Centers for Disease Control and Prevention. “Interim clinical considerations for use of COVID-19 vaccines currently authorized in the United States.” (2021)
P. Tang, M.R. Hasan, H. Chemaitelly, H.M. Yassine, F.M. Benslimane, H.A. Al Khatib, S. AlMukdad, P. Coyle, H.H. Ayoub, Z. Al Kanaani, E. Al Kuwari, A. Jeremijenko, A.H. Kaleeckal, A.N. Latif, RM. Shaik, H.F. Abdul Rahim, G.K. Nasrallah, M.G. Al Kuwari, H.E. Al Romaihi, A.A. Butt, M.H. Al-Thani, A. Al Khal, R. Bertollini, L.J. Abu-Raddad, BNT162b2 and mRNA-1273 COVID-19 vaccine effectiveness against the SARS-CoV-2 Delta variant in Qatar. Nat Med. 2021 Nov 2. https://doi.org/10.1038/s41591-021-01583-4. Epub ahead of print. PMID: 34728831
S. Nasreen et al., Effectiveness of COVID-19 vaccines against variants of concern. Canada. Preprint at medRxiv (2021). https://doi.org/10.1101/2021.06.28.21259420
A.E. Samoilov, V.V. Kaptelova, A.Y. Bukharina et al., Case report: change of dominant strain during dual SARS-CoV-2 infection. BMC Infect. Dis. 21, 959 (2021)
https://www.cnbc.com/2021/07/12/belgian-woman-infected-with-two-covid-variants-at-the-same-time.html
https://edition.cnn.com/2021/03/11/americas/brazil-variants-simultaneous-infection-intl/index.html
P. Combes, M. Bisseux, A. Bal, et al. Evidence of co-infection during delta and omicron variants of concern co-circulation, weeks 49-2021 to 02-2022, France. medRxiv 2022; published online March 3. https://doi.org/10.1101/2022.03.02.22271694 (preprint)
R.J. Rockett, J. Draper, M. Gall, et al. Co-infection with SARS-COV-2 omicron and delta variants revealed by genomic surveillance. medRxiv 2022; published online Feb 15. https://doi.org/10.1101/2022.02.13.22270755(preprint)
M.L. Vatteroni, A-L. Capria, P.G. Spezia, S. Frateschi, M. Pistello, Co-infection with SARS-CoV-2 omicron BA.1 and BA.2 subvariants in a nonvaccinated woman. The Lancet 2022; https://doi.org/10.1016/S2666-5247(22)00119-7
M. Farman, M. Aslam, A. Akgul, A. Ahmad, Modeling of fractional-order COVID-19 epidemic model with quarantine and social distancing. Math. Methods Appl. Sci. 44(11), 9334–9350 (2021)
A. Ahmed, B. Salam, M. Mohammad, A. Akgul, S.H. Khoshnaw, Analysis coronavirus disease (COVID-19) model using numerical approaches and logistic model. Aims Bioeng. 7(3), 130–146 (2020)
G. Hussain, T. Khan, A. Khan, M. Inc, G. Zaman, K.S. Nisar, A. Akgul, Modeling the dynamics of novel coronavirus (COVID-19) via stochastic epidemic model. Alex. Eng. J. 60(4), 4121–4130 (2021)
A. Khan, R. Zarin, U.W. Humphries, A. Akgul, A. Saeed, T. Gul, Fractional optimal control of COVID-19 pandemic model with generalized Mittag-Leffler function. Adv. Diff. Equs. 2021, 1–22 (2021)
C. Xu, M. Farman, A. Hasan, A. Akgul, M. Zakarya, W. Albalawi, C. Park, Lyapunov stability and wave analysis of Covid-19 omicron variant of real data with fractional operator. Alex. Eng. J. 61(12), 11787–11802 (2022)
M. Farman, A. Akgul, K.S. Nisar, D. Ahmad, A. Ahmad, S. Kamangar, C.A. Saleel, Epidemiological analysis of fractional order COVID-19 model with Mittag-Leffler kernel. AIMS Math. 7(1), 756–783 (2022)
M. Amin, M. Farman, A. Akgul, R.T. Alqahtani, Effect of vaccination to control COVID-19 with fractal fractional operator. Alex. Eng. J. 61(5), 3551–3557 (2022)
A. Khan, R. Zarin, S. Khan, A. Saeed, T. Gul, U.W. Humphries, Fractional dynamics and stability analysis of COVID-19 pandemic model under the harmonic mean type incidence rate. Computer Methods in Biomechanics and Biomedical Engineering, pp. 1-22 (2021)
S. Liu, J. Shen, S. Fang, K. Li, J. Liu, L. Yang et al., Genetic spectrum and distinct evolution patterns of SARS-CoV-2. Front. Microbiol. 11, 2390 (2020). https://doi.org/10.3389/fmicb.2020.593548
H. Hashim, M. Mohammed, M. Mousa, H. Abdulameer, A. Alhassnawi, S. Hassan et al., Infection with different strains of SARS-COV-2 in patients with COVID-19. Arch. Biol. Sci. 72, 575–585 (2020)
Ndolane Sene, Analysis of the stochastic model for predicting the novel coronavirus disease. Adv. Diff. Equs. 2020(1), 1–19 (2020)
E. Yuliani, C. Alfiniyah, M.L. Juga, C.W. Chukwu, On the modeling of COVID-19 transmission dynamics with two strains: insight through caputo fractional derivative. Fractal Fractional. 6(7), 346 (2022)
P. van den Driessche, J. Watmough, Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math. Biosci. 180, 29–48 (2002)
N. Sapoukhina, Y. Tyutyunov, A. Arditi, The role of prey-taxis in biological control. The Am. Nat. 162(1), 61–76 (2003)
E. Ahmed, A.M.A. El-Sayed, H.A.A. El-Saka, On some Routh-Hurwitz conditions for fractional order differential equations and their applications in Lorenz, Rossler, Chua and Chen systems. Phys. Lett. 1, 1–4 (2006)
N. Chitnis, J.M. Hyman, J.M. Cushing, Determining important parameters in the spread of malaria through the sensitivity analysis of a mathematical model. Bull. Math. Biol. 70, 1272–1296 (2008)
J. Crank, P. Nicolson, A practical method for numerical evaluation of solutions of partial differential equations of the heat-conduction type. Proc. Camb. Philos. Soc. 43(1), 50–67 (1947)
K.W. Morton, D.F. Mayers, Numerical Solution of Partial Differential Equations: An Introduction. Cambridge University Press (2005)
H. Nadeem, Numerical solution of compartmental models by meshless and finite difference methods. Appl. Math. Comput. 238(2), 408–435 (2014)
R. Zarin, N. Haider, Numerical solution of COVID-19 pandemic model via finite difference and meshless techniques. Eng. Anal. Bound. Elem. 1(147), 76–89 (2023)
Acknowledgements
The first author appreciates the support provided by Petchra Pra Jom Klao Ph.D. Research Scholarship through grant no (50/2565), by King Mongkut’s University of Technology Thonburi, Thailand.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zarin, R., Humphries, U.W. A robust study of dual variants of SARS-CoV-2 using a reaction-diffusion mathematical model with real data from the USA. Eur. Phys. J. Plus 138, 1057 (2023). https://doi.org/10.1140/epjp/s13360-023-04631-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjp/s13360-023-04631-9