Abstract
Complex network is an effective approach to studying the characteristics and interactions of complex systems, which can be used to analyze the core functions and global behavior of complex biological systems. Type 2 diabetes mellitus (T2DM), the most common type of diabetes mellitus, is a complex polygenic metabolic disease associated with genetic and environmental factors. How the complex interactions between T2DM-related genes affect the pathogenesis and treatment of T2DM is not yet fully understood. By applying the network approach to biological data, this study constructs a pathway-based network model of T2DM-related genes to explore the interrelationships between genes. Analysis of statistical and topological characteristics shows that the network exhibits the small-world rather than scale-free property, with a high average degree of 99.22, revealing close and complex connections between these genes. To determine the key hub genes of the network, an integrated centrality is used to comprehensively reflect the contribution of the three centrality indices (degree centrality, betweenness centrality and closeness centrality) of nodes; by taking the threshold of 0.70 for integrated centrality, nine key hub genes are identified: PIK3CD, PIK3CA, MAPK1, PIK3R1, PRKCA, AKT2, AKT1, TNF and KRAS. These genes should play an important role in the occurrence and development of T2DM, and their identification will provide relevant and useful knowledge for further biological and medical research on their functions in T2DM (especially in the development of multi-target drugs for T2DM). This further provides clues for exploring the pathogenesis and treatment of T2DM.
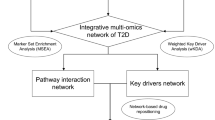
Graphic abstract






Similar content being viewed by others
Data Availability Statement
The datasets of T2DM-related genes and related biological pathways were collected from the NCBI Gene Database and the KEGG PATHWAY Database, respectively. The main data supporting the findings of this study are included in this article. Other data generated during and/or analyzed during this study are available from the corresponding author on reasonable request.
Notes
NCBI (National Center for Biotechnology Information) website: https://www.ncbi.nlm.nih.gov/.
KEGG (Kyoto Encyclopedia of Genes and Genomes) website: https://www.kegg.jp/ or https://www.genome.jp/kegg/.
References
H. Freeman, R.D. Cox, Type-2 diabetes: a cocktail of genetic discovery. Hum. Mol. Genet. 15(Suppl. 2), R202–R209 (2006)
Z. Shen, Q. Chen, H. Ying, Z. Ma, X. Bi, X. Li, M. Wang, C. Jin, D. Lai, Y. Zhao, G. Fu, Identification of differentially expressed genes in the endothelial precursor cells of patients with type 2 diabetes mellitus by bioinformatics analysis. Exp. Ther. Med. 19(1), 499–510 (2020)
International Diabetes Federation, IDF Diabetes Atlas, 10th edn. (2021)
A. Saxena, N. Wahi, A. Kumar, S.K. Mathur, Functional interactomes of genes showing association with type-2 diabetes and its intermediate phenotypic traits point towards adipo-centric mechanisms in its pathophysiology. Biomolecules 10(4), 601 (2020)
Y. Zheng, S.H. Ley, F.B. Hu, Global aetiology and epidemiology of type 2 diabetes mellitus and its complications. Nat. Rev. Endocrinol. 14(2), 88–98 (2018)
Chinese Diabetes Society, Guidelines for the prevention and treatment of type 2 diabetes in China (2020 edition). Chin. J. Diabetes Mellit. 13(4), 315–409 (2021)
J.E. Shaw, R.A. Sicree, P.Z. Zimmet, Global estimates of the prevalence of diabetes for 2010 and 2030. Diabetes Res. Clin. Pract. 87(1), 4–14 (2010)
The InterAct Consortium, The link between family history and risk of type 2 diabetes is not explained by anthropometric, lifestyle or genetic risk factors: the EPIC-InterAct study. Diabetologia 56(1), 60–69 (2013)
T.A. Harrison, L.A. Hindorff, H. Kim, R.C.M. Wines, D.J. Bowen, B.B. McGrath, K.L. Edwards, Family history of diabetes as a potential public health tool. Am. J. Prev. Med. 24(2), 152–159 (2003)
I. Sethi, V. Sharma, I. Sharma, G. Singh, Gh.R. Bhat, A.J.S. Bhanwer, S. Sharma, E. Rai, Telomere maintenance genes are associated with type 2 diabetes susceptibility in northwest Indian population group. Sci. Rep. 10, 6444 (2020)
Q. Ma, Y. Li, M. Wang, Z. Tang, T. Wang, C. Liu, C. Wang, B. Zhao, Progress in metabonomics of type 2 diabetes mellitus. Molecules 23(7), 1834 (2018)
D.J. Watts, S.H. Strogatz, Collective dynamics of ‘small-world’ networks. Nature 393(6684), 440–442 (1998)
A.-L. Barabási, R. Albert, Emergence of scaling in random networks. Science 286(5439), 509–512 (1999)
S.H. Strogatz, Exploring complex networks. Nature 410(6825), 268–276 (2001)
M.E.J. Newman, The structure and function of complex networks. SIAM Rev. 45(2), 167–256 (2003)
H. Jeong, B. Tombor, R. Albert, Z.N. Oltvai, A.-L. Barabási, The large-scale organization of metabolic networks. Nature 407(6804), 651–654 (2000)
A.-L. Barabási, Z.N. Oltvai, Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5(2), 101–113 (2004)
F. Karlsson, M. Hörnquist, Order or chaos in Boolean gene networks depends on the mean fraction of canalizing functions. Phys. A 384(2), 747–757 (2007)
M. Tsuchiya, K. Selvarajoo, V. Piras, M. Tomita, A. Giuliani, Local and global responses in complex gene regulation networks. Phys. A 388(8), 1738–1746 (2009)
L. Diambra, Coarse-grain reconstruction of genetic networks from expression levels. Phys. A 390(11), 2198–2207 (2011)
F. Censi, A. Giuliani, P. Bartolini, G. Calcagnini, A multiscale graph theoretical approach to gene regulation networks: a case study in atrial fibrillation. IEEE Trans. Biomed. Eng. 58(10), 2943–2946 (2011)
R. Demicheli, D. Coradini, Gene regulatory networks: a new conceptual framework to analyse breast cancer behaviour. Ann. Oncol. 22(6), 1259–1265 (2011)
H.B. Chakrapani, S. Chourasia, S. Gupta, D. Thirumal Kumar, C. George Priya Doss, R. Haldar, Effective utilisation of influence maximization technique for the identification of significant nodes in breast cancer gene networks. Comput. Biol. Med. 133, 104378 (2021)
H. Wang, C.-Y. Xu, J.-B. Hu, K.-F. Cao, A complex network analysis of hypertension-related genes. Phys. A 394, 166–176 (2014)
H. Wang, C.-Y. Xu, J.-B. Hu, K.-F. Cao, A complex network analysis of hypertension-related genes (Revised version). arXiv:1601.07192 (2016)
H. Wang, J.-B. Hu, C.-Y. Xu, D.-H. Zhang, Q. Yan, M. Xu, K.-F. Cao, X.-S. Zhang, A pathway-based network analysis of hypertension-related genes. Phys. A 444, 928–939 (2016)
H. Wang, J.-B. Hu, C.-Y. Xu, D.-H. Zhang, Q. Yan, M. Xu, K.-F. Cao, X.-S. Zhang, Corrigendum to “A pathway-based network analysis of hypertension-related genes” [Phys. A 444, 928–939 (2016)]. Phys. A 447, 569–570 (2016)
J.-B. Hu, H. Wang, L. Wang, C.-Y. Xu, K.-F. Cao, X.-S. Zhang, Characteristic analysis of the pathway-based weighted network of hypertension-related genes. Phys. A 533, 122069 (2019)
L.K. Dahl, M. Heine, L. Tassinari, Effects of chronic excess salt ingestion: evidence that genetic factors play an important role in susceptibility to experimental hypertension. J. Exp. Med. 115(6), 1173–1190 (1962)
J.P. Rapp, Dahl salt-susceptible and salt-resistant rats: a review. Hypertension 4(6), 753–763 (1982)
M. Liang, N.H. Lee, H. Wang, A.S. Greene, A.E. Kwitek, M.L. Kaldunski, T.V. Luu, B.C. Frank, S. Bugenhagen, H.J. Jacob, A.W. Cowley Jr., Molecular networks in Dahl salt-sensitive hypertension based on transcriptome analysis of a panel of consomic rats. Physiol. Genom. 34(1), 54–64 (2008)
A.W. Cowley Jr., R.J. Roman, H.J. Jacob, Application of chromosomal substitution techniques in gene-function discovery. J. Physiol. 554(1), 46–55 (2004)
J. Qiu, J.H. Moore, C. Darabos, Studying the genetics of complex disease with ancestry-specific human phenotype networks: the case of type 2 diabetes in East Asian populations. Genet. Epidemiol. 40(4), 293–303 (2016)
H. Kitano, Systems biology: a brief overview. Science 295(5560), 1662–1664 (2002)
Y. Deville, D. Gilbert, J. van Helden, S.J. Wodak, An overview of data models for the analysis of biochemical pathways. Brief. Bioinform. 4(3), 246–259 (2003)
S. Sharma, S. Ciufo, E. Starchenko, D. Darji, L. Chlumsky, I. Karsch-Mizrachi, C.L. Schoch, The NCBI BioCollections Database. Database 2018, bay006 (2018)
S. Sharma, S. Ciufo, E. Starchenko, D. Darji, L. Chlumsky, I. Karsch-Mizrachi, C.L. Schoch, Corrigendum to “The NCBI BioCollections Database” [Database 2018, bay006 (2018)]. Database 2019, baz057 (2019)
M. Kanehisa, S. Goto, KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000)
M.E.J. Newman, Assortative mixing in networks. Phys. Rev. Lett. 89(20), 208701 (2002)
M.A. Beauchamp, An improved index of centrality. Behav. Sci. 10(2), 161–163 (1965)
L.C. Freeman, A set of measures of centrality based on betweenness. Sociometry 40(1), 35–41 (1977)
L.C. Freeman, Centrality in social networks: conceptual clarification. Social Networks 1(3), 215–239 (1978/79)
X. Du, X. Li, L. Chen, M. Zhang, L. Lei, W. Gao, Z. Shi, Y. Dong, Z. Wang, X. Li, G. Liu, Hepatic miR-125b inhibits insulin signaling pathway by targeting PIK3CD. J. Cell. Physiol. 233(8), 6052–6066 (2018)
X. Yin, Z. Xu, Z. Zhang, L. Li, Q. Pan, F. Zheng, H. Li, Association of PI3K/AKT/mTOR pathway genetic variants with type 2 diabetes mellitus in Chinese. Diabetes Res. Clin. Pract. 128, 127–135 (2017)
L. Wang, W. Diwu, N. Tan, H. Wang, J. Hu, B. Xu, X. Wang, Pathway-based protein–protein association network to explore mechanism of \(\alpha \)-glucosidase inhibitors from Scutellaria baicalensis Georgi against type 2 diabetes. IET Syst. Biol. 15(4), 126–135 (2021)
A.H. Karadoǧan, H. Arikoglu, F. Göktürk, F. İşçioǧlu, S.H. İpekçi, PIK3R1 gene polymorphisms are associated with type 2 diabetes and related features in the Turkish population. Adv. Clin. Exp. Med. 27(7), 921–927 (2018)
S. Haydar, F. Grigorescu, M. Vintilǎ, Y. Cogne, C. Lautier, Y. Tutuncu et al., Fine-scale haplotype mapping of MUT, AACS, SLC6A15 and PRKCA genes indicates association with insulin resistance of metabolic syndrome and relationship with branched chain amino acid metabolism or regulation. PLoS One 14(3), e0214122 (2019)
R. Kushi, Y. Hirota, W. Ogawa, Insulin resistance and exaggerated insulin sensitivity triggered by single-gene mutations in the insulin signaling pathway. Diabetol. Int. 12(1), 62–67 (2021)
B. Yin, Y.-M. Bi, G.-J. Fan, Y.-Q. Xia, Molecular mechanism of the effect of Huanglian Jiedu Decoction on type 2 diabetes mellitus based on network pharmacology and molecular docking. J. Diabetes Res. 2020, 5273914 (2020)
T.O. Kilpeläinen, T.A. Lakka, D.E. Laaksonen, U. Mager, T. Salopuro, A. Kubaszek, B. Todorova, O. Laukkanen, J. Lindström, J.G. Eriksson et al., Interaction of single nucleotide polymorphisms in ADRB2, ADRB3, TNF, IL6, IGF1R, LIPC, LEPR, and GHRL with physical activity on the risk of type 2 diabetes mellitus and changes in characteristics of the metabolic syndrome: The Finnish Diabetes Prevention Study. Metab. Clin. Exp. 57(3), 428–436 (2008)
W.S. Al-Qahtani, E. Al-Olayan, F.G. Albani, R.S. Suliman, N.H. Aljarba, E.M. Al-Humaidhi, A.S. Almurshedi, D.M. Domiaty, M.A. Alduwish, A.M. Al-Otaibi, A.M. Elasbali, H.G. Ahmed, B.A. Almutlaq, Utility of KRAS gene and clinicopathological features in the assessment of the risk of type 2 diabetes in the etiology of colon cancer. Glob. Med. Genet. 7(2), 35–40 (2020)
C. Nesti, A. Rubegni, D. Tolomeo, J. Baldacci, D. Cassandrini, F. D’Amore, F.M. Santorelli, Complex multisystem phenotype associated with the mitochondrial DNA m.5522G\(>\)A mutation. Neurol. Sci. 40(8), 1705–1708 (2019)
D. He, J.-H. Huang, Z.-Y. Zhang, Q. Du, W.-J. Peng, R. Yu, S.-F. Zhang, S.-H. Zhang, Y.-H. Qin, A network pharmacology-based strategy for predicting active ingredients and potential targets of LiuWei DiHuang Pill in treating type 2 diabetes mellitus. Drug Des. Dev. Ther. 13, 3989–4005 (2019)
Y.L. Konheim, J.K. Wolford, Association of a promoter variant in the inducible cyclooxygenase-2 gene (PTGS2) with type 2 diabetes mellitus in Pima Indians. Hum. Genet. 113(5), 377–381 (2003)
J. Chen, Y. Meng, J. Zhou, M. Zhuo, F. Ling, Y. Zhang, H. Du, X. Wang, Identifying candidate genes for Type 2 Diabetes Mellitus and obesity through gene expression profiling in multiple tissues or cells. J. Diabetes Res. 2013, 970435 (2013)
M.M. Kamkar, R. Nizam, S. Hasan, Genetic association of ITPR3 SNP rs999943 with Type 2 Diabetes and related metabolic traits, in International Diabetes Federation 2017 Congress (International Diabetes Federation – Abudhabi, Abudhabi, United Arab Emirates, 4–8 December 2017) (2017)
A.N. Zhu, X.X. Yang, M.Y. Sun, Z.X. Zhang, M. Li, Associations between INSR and MTOR polymorphisms in type 2 diabetes mellitus and diabetic nephropathy in a Northeast Chinese Han population. Genet. Mol. Res. 14(1), 1808–1818 (2015)
K. Rehman, M.S.H. Akash, A. Liaqat, S. Kamal, M.I. Qadir, A. Rasul, Role of interleukin-6 in development of insulin resistance and type 2 diabetes mellitus. Crit. Rev. Eukaryot. Gene Expr. 27(3), 229–236 (2017)
L. Yin, W.-J. Cai, X.-Y. Chang, J. Li, L.-Y. Zhu, X.-H. Su, X.-F. Yu, K. Sun, Analysis of PTEN expression and promoter methylation in Uyghur patients with mild type 2 diabetes mellitus. Medicine (Baltimore) 97(49), e13513 (2018)
C.-P. Kung, M.E. Murphy, The role of the p53 tumor suppressor in metabolism and diabetes. J. Endocrinol. 231(2), R61–R75 (2016)
H.-M. Liu, Y. Huang, L. Li, Y. Zhang, X. Cong, L.-L. Wu, R.-L. Xiang, MicroRNA-mRNA expression profiles and functional network of submandibular gland in type 2 diabetic db/db mice. Arch. Oral Biol. 120, 104947 (2020)
T. Hamaguchi, Y. Hirota, T. Takeuchi, Y. Nakagawa, A. Matsuoka, M. Matsumoto, H. Awano, K. Iijima, P.C. Cha, W. Satake, T. Toda, W. Ogawa, Treatment of a case of severe insulin resistance as a result of a PIK3R1 mutation with a sodium–glucose cotransporter 2 inhibitor. J. Diabetes Investig. 9(5), 1224–1227 (2018)
D. Kesharwani, A. Kumar, M. Poojary, V. Scaria, M. Datta, RNA sequencing reveals potential interacting networks between the altered transcriptome and ncRNome in the skeletal muscle of diabetic mice. Biosci. Rep. 41(7), BSR20210495 (2021)
Z. Peng, R. Aggarwal, N. Zeng, L. He, E.X. Stiles, A. Debebe, J. Chen, C.-Y. Chen, B.L. Stiles, AKT1 regulates endoplasmic reticulum stress and mediates the adaptive response of pancreatic \(\beta \) cells. Mol. Cell. Biol. 40(11), e00031-20 (2020)
L. He, Y. Li, N. Zeng, B.L. Stiles, Regulation of basal expression of hepatic PEPCK and G6Pase by AKT2. Biochem. J. 477(5), 1021–1031 (2020)
L. Nigi, G.E. Grieco, G. Ventriglia, N. Brusco, F. Mancarella, C. Formichi, F. Dotta, G. Sebastiani, MicroRNAs as regulators of insulin signaling: research updates and potential therapeutic perspectives in type 2 diabetes. Int. J. Mol. Sci. 19(12), 3705 (2018)
F. Yang, Y. Chen, Z. Xue, Y. Lv, L. Shen, K. Li, P. Zheng, P. Pan, T. Feng, L. Jin, Y. Yao, High-throughput sequencing and exploration of the lncRNA-circRNA-miRNA-mRNA network in type 2 diabetes mellitus. Biomed. Res. Int. 2020, 8162524 (2020)
Z.-M. Yang, L.-H. Chen, M. Hong, Y.-Y. Chen, X.-R. Yang, S.-M. Tang, Q.-F. Yuan, W.-W. Chen, Serum microRNA profiling and bioinformatics analysis of patients with type 2 diabetes mellitus in a Chinese population. Mol. Med. Rep. 15(4), 2143–2153 (2017)
F.-C. Chen, K.-P. Shen, J.-B. Chen, H.-L. Lin, C.-L. Hao, H.-W. Yen, S.-Y. Shaw, PGBR extract ameliorates TNF-\(\alpha \) induced insulin resistance in hepatocytes. Kaohsiung J. Med. Sci. 34(1), 14–21 (2018)
I.T. Lampropoulou, M. Stangou, P. Sarafidis, A. Gouliovaki, P. Giamalis, I. Tsouchnikas, T. Didangelos, A. Papagianni, TNF-\(\alpha \) pathway and T-cell immunity are activated early during the development of diabetic nephropathy in Type II Diabetes Mellitus. Clin. Immunol. 215, 108423 (2020)
M. Nagatsuma, K. Takasawa, T. Yamauchi, R. Nakagawa, T. Mizuno, E. Tanaka, K. Yamamoto, N. Uemura, K. Kashimada, T. Morio, A postzygotic KRAS mutation in a patient with Schimmelpenning syndrome presenting with lipomatosis, renovascular hypertension, and diabetes mellitus. J. Hum. Genet. 64(2), 177–181 (2019)
S. Huang, S. Kauffman, How to escape the cancer attractor: rationale and limitations of multi-target drugs. Semin. Cancer Biol. 23(4), 270–278 (2013)
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China (NSFC) (Grant nos. 11365023 and 51866005). The authors would like to thank Professors Fukai Bao and Wen Zhang from Kunming Medical University for their helpful discussions, and the anonymous reviewer for valuable comments and suggestions.
Author information
Authors and Affiliations
Contributions
XYZ and KFC conceived and designed the research. XYZ collected the data, performed numerical calculations with the help of CYX, and drafted the original manuscript. All authors contributed to the analysis, discussion and interpretation of the results. KFC revised the manuscript and finalized the submission version of the manuscript with XSZ.
Corresponding author
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, XY., He, TY., Xu, CY. et al. Theoretical investigation of the pathway-based network of type 2 diabetes mellitus-related genes. Eur. Phys. J. B 96, 86 (2023). https://doi.org/10.1140/epjb/s10051-023-00540-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjb/s10051-023-00540-z