Abstract
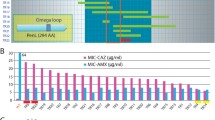
Beta-lactamases (EC 3.5.2.6) represent a superfamily containing more than 2000 members: it includes genetically and functionally different bacterial enzymes capable to degrade the beta-lactam antibiotics. Beta-lactamases of molecular class A with serine residue in the active center are the most common ones. In the context of studies of the mechanisms underlying of evolution of the resistance, TEM type beta-lactamases are of particular interest due to their broad polymorphism. To date, more than 200 sequences of TEM type beta-lactamases have been described and more than 60 structures of different mutant forms of these enzymes have been presented in the Protein Data Bank. We have considered here the main structural features of the enzymes of this type with particular attention to the analysis of key mutations determining drug resistance and the secondary mutations, their location relative to the active center and the surface of the protein globule. We have developed a BlaSIDB database (www.blasidb.org) which is an open information resource combining available data on 3D structures, amino acid sequences and nomenclature of the TEM type beta-lactamases.
Similar content being viewed by others
Abbreviations
- BlaSIDB:
-
beta-lactamase structure information database
- ESBL:
-
extended spectrum β-lactamases
- PDB:
-
Protein Data Bank
- SASA:
-
solvent accessible surface area
- WHO:
-
World Health Organization
References
World Health Organization, Antimicrobial Resistance Global Report on Surveillance, 2014. http://www.who.int/drugresistance/en/.
Abraham, E.P. and Chain, E., Nature, 1940, vol. 46, pp. 837–837.
Bonomo, R.A., Cold Spring Harb. Perspect. Med., 2017, vol. 7, pp. 1–15.
Ambler, R.P., Coulson, A.F., Frère, J.M., Ghuysen, J.M., Joris, B., Forsman, M., Levesque, R.C., Tiraby, G., and Waley, S.G., Biochem. J., 1991, vol. 276, pp. 269–270.
Bush, K. and Jacoby, G.A., Antimicrobial Agents and Chemotherapy, 2010, vol. 54, pp. 969–976.
Bush, K., Ann. N.Y. Acad. Sci., 2013, vol. 1277, pp. 84–90.
Livermore, D.M., Clin. Microbiol. Infect., 2008, vol. 14, suppl. 1, pp. 3–10.
Lee, J.H., Bae, I.K., and Lee, S.H., Medicinal Res. Rev., 2012, vol. 32, pp. 216–232.
Baquero, F., Tedim, A.P., and Coque, T.M., Front. Microbiol., 2013, vol. 4, pp. 1–15.
Medeiros, A.A., Clin. Infect. Dis., 1997, vol. 24, pp. S19–S45.
Matagne, A., Lamotte-Brasseur, J., and Frere, J.M., Biochem. J., 1998, vol. 330, pp. 581–598.
Datta, N. and Kontomichalou, P., Nature, 1965, vol. 208, pp. 239–241.
Du Bois, S.K., Marriott, M.S., and Amyes, S.G., J. Antimicrob. Chemother., 1995, vol. 35, pp. 7–22.
Pimenta, A.C., Fernandes, R., and Moreira, I.S., Mini Rev. Med. Chem., 2014, vol. 14, pp. 111–122.
Salverda, M.L., De Visser, J.A., and Barlow, M., FEMS Microbiol. Rev., 2010, vol. 34, pp. 1015–1036.
Stec, B., Holtz, K.M., Wojciechowski, C.L., and Kantrowitz, E.R., Acta Crystallogr. D. Biol. Crystallogr., 2005, vol. 61, pp. 1072–1079.
Fisette, O., Morin, S., Savard, P.Y., Lagüe, P., and Gagné, S.M., Biophys. J., 2010, vol. 98, pp. 637–645.
Guthrie, V.B., Allen, J., Camps, M., and Karchin, R., PLoS Computational Biology, 2011, vol. 7, e1002184, pp. 1–20.
Dellus-Gur, E., Toth-Petroczy, A., and Tawfik, D.S., J. Mol. Biol., 2013, vol. 425, pp. 2609–2621.
Wang, X., Minasov, G., and Shoichet, B.K., J. Mol. Biol., 2002, vol. 320, pp. 85–95.
Kather, I., Jakob, R.P., Dobbek, H., and Schmid, F.X., J. Mol. Biol., 2008, vol. 383, pp. 238–251.
Meroueh, S.O., Roblin, P., Golemi, D., Maveyraud, L., Vakulenko, S.B., Zhang, Y., Samama, J.P., and Mobashery, S., J. Am. Chem. Soc., 2002, vol. 124, pp. 9422–9430.
Stojanoski, V., Chow, D.C., Hu, L., Sankaran, B., Gilbert, H.F., Prasad, B.V., and Palzkill, T., J. Biol. Chem., 2015, vol. 290, pp. 10382–10394.
Grigorenko, V., Uporov, I., Rubtsova, M., Andreeva, I., Shcherbinin, D., Veselovsky, A., Serova, O., Ulyashova, M., Ishtubaev, I., and Egorov, A., FEBS Open Bio, 2017. doi 10.1002/2211-5463.12352
Thai, Q.K., Bős, F., and Pleiss, J., BMC Genomics, 2009, vol. 10, p. 390.
Naas, T., Oueslati, S., Bonnin, R.A., Dabos, M.L., Zavala, A., Dortet, L., Retailleau, P. and Iorga, B.I., J. Enz. Inh. Med. Chem., 2017, vol. 32, pp. 917–919.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © V.G. Grigorenko, M.Yu. Rubtsova, I.V. Uporov, I.V. Ishtubaev, I.P. Andreeva, D.S. Shcherbinin, A.V. Veselovsky, A.M. Egorov, 2018, published in Biomeditsinskaya Khimiya.
Rights and permissions
About this article
Cite this article
Grigorenko, V.G., Rubtsova, M.Y., Uporov, I.V. et al. Bacterial TEM-Type Serine Beta-Lactamases: Structure and Analysis of Mutations. Biochem. Moscow Suppl. Ser. B 12, 87–95 (2018). https://doi.org/10.1134/S1990750818020038
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1990750818020038