Abstract
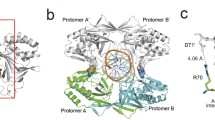
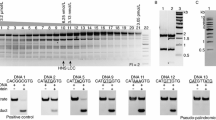
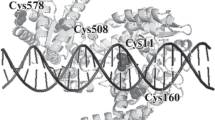
The photoactivatable modified oligonucleotides were used to investigate direct contacts formed by the type IIE EcoRII restriction endonuclease and the T/A bases of its recognition site (5'-CCT/AGG). EcoRII dimer consists of a central catalytic core, made of two C-terminal endonuclease-like domains (EcoRII-C) from different subunits, and two N-terminal effector DNA binding domains (EcoRII-N). According to co-crystal structure of isolated EcoRII-C with DNA catalytic dimer EcoRII-C flips nucleotides of the central T/A pair into the enzyme binding pockets. Нere, photocross-linking technique was used to investigate the direct contacts formed by extrahelical T/A bases in the protein pockets of full-length EcoRII within the pre-reactive EcoRII–DNA complex obtained in the presence of Ca2+ in solution. Photoreactive zero-length agent 5-iodo-2'-deoxyuridine (IdU) was introduced as single substituent into the central T/A position of EcoRII recognition site or into the flanking nucleotide sequences of 14-mer DNA substrate. The substitution of only dT or dA residues in EcoRII recognition site resulted in formation of photocross-links upon irradiation only in the presence of Ca2+. Proteolytic digestion of the enzyme-oligonucleotide conjugates followed by MALDI-MS analysis have allowed to identify the 224VEYD227 EcoRII region involved in the formation of the cross-links. This region belongs to the central part of H-10 α-helix. Y226 residue was suggested to form cross-link with T or A bases of EcoRII site replaced by IdU within the pre-reactive complex. The flipped base pair protein pockets of EcoRII seem to accommodate equally well both A and T bases of the DNA substrate. Altogether, IdU-containing photoactivatable DNA substrates have allowed to trap the flipped bases in complex with full-length EcoRII before DNA cleavage in the solution and to identify direct enzyme–DNA contacts important for high specificity of EcoRII for the Т/A nucleotides providing a highly specific cleavage reaction.






Similar content being viewed by others
REFERENCES
Pingoud, A. and Jeltsch, A., Nucleic Acids Res., 2001, vol. 29, pp. 3705–3727. https://doi.org/10.1093/nar/29.18.3705
Roberts, R.J., Belfort, M., Bestor, T., et al., Nucleic Acids Res., 2003, vol. 31, pp. 1805–1812. https://doi.org/10.1093/nar/gkg274
Mücke, M., Krüger, D.H., and Reuter, M., Nucleic Acids Res., 2003, vol. 31, pp. 6079–6084. https://doi.org/10.1093/nar/gkg836
Kirsanova, O.V., Baskunov, V.B., and Gromova, E.S., Mol. Biol. (Moscow), 2004, vol. 38, pp. 752–764. https://doi.org/10.1023/B:MBIL.0000043944.45429.aa
Petrauskene, O.V., Karpova, E.A., Gromova, E.S., and Guschlbauer, W., Biochem. Biophys. Res. Commun., 1994, vol. 198, pp. 885–890. https://doi.org/10.1006/bbrc.1994.1126
Tamulaitis, G., Mücke, M., and Siksnys, V., FEBS Lett., 2006, vol. 580, pp. 1665–1671. https://doi.org/10.1016/j.febslet.2006.02.010
Tamulaitis, G., Sasnauskas, G., Mücke, M., and Siksnys, V., J. Mol. Biol., 2006, vol. 358, pp. 406–419. https://doi.org/10.1016/j.jmb.2006.02.024
Shlyakhtenko, L.S., Gilmore, J., Portillo, A., Tamulaitis, G., Siksnys, V., and Lyubchenko, Y.L., Biochemistry, 2007, vol. 46, pp. 11128–11136. https://doi.org/10.1021/bi701123u
Zhou, X.E., Wang, Y., Reuter, M., Mücke, M., Krüger, D.H., Meehan E.J., and Chen, L., J. Mol. Biol., 2004, vol. 335, pp. 307–319. https://doi.org/10.1016/j.jmb.2003.10.030
Golovenko, D., Manakova, E., Tamulaitiene, G., Grazulis, S., and Siksnys, V., Nucleic Acids Res., 2009, vol. 37, pp. 6613–6624. https://doi.org/10.1093/nar/gkp699
Klimasauskas, S., Kumar, S., Roberts, R.J., and Cheng, X.D., Cell, 1994, vol. 76, pp. 357–369. https://doi.org/10.1016/0092-8674(94)90342-5
Reinisch, K.M., Chen, L., Verdine, G.L., and Lipscomb, W.N., Cbabell., 1995, vol. 82, pp. 143–153. https://doi.org/10.1016/0092-8674(95)90060-8
Banerjee, A., Yang, W., Karplus, M., and Verdine, G.L., Nature, 2005, vol. 434, pp. 612–618. https://doi.org/10.1038/nature03458
Tamulaitis, G., Zaremba, M., Szczepanowski, R.H., Bochtler, M., and Siksnys, V., Nucleic Acids Res., 2007, vol. 35, pp. 4792–4799. https://doi.org/10.1093/nar/gkm513
Neely, R.K., Tamulaitis, G., Chen, K., Kubala, M., Siksnys, V., and Jones, A.C., Nucleic Acids Res., 2009, vol. 37, pp. 6859–6870. https://doi.org/10.1093/nar/gkp688
Babkina, O.V., Chutko, C.A., Shashkov, A.A., Dzhidzhoev, M.S., Eritja, R.I., and Gromova, E.S., Photochem. Photobiol. Sci., 2002, vol. 1, pp. 636–640. https://doi.org/10.1039/B202028A
Mücke, M., Pingoud, V., Grelle, G., Kraft, R., Krüger, D.H., and Reuter, M., J. Biol. Chem., 2002, vol. 277, pp. 14288–14293. https://doi.org/10.1074/jbc.M109311200
Tamulaitiene, G., Manakova, E., Jovaisaite, V., Tamulaitis, G., Grazulis, S., Bochtler, M., and Siksnys, V., Nucleic Acids Res., 2019, vol. 48, pp. 997–1010. https://doi.org/10.1093/nar/gky1137
Bochtler, M., Szczepanowski, R.H., Tamulaitis, G., Grazulis, S., Czapinska, H., Manakova, E., and Siksnys, V., EMBO J., 2006, vol. 25, pp. 2219–2229. https://doi.org/10.1038/sj.emboj.7601096
Tamulaitis, G., Zaremba, M., Szczepanowski, R.H., Bochtler, M., and Siksnys, V., Nucleic Acids Res., 2008, vol. 36, pp. 6101–6108. https://doi.org/10.1093/nar/gkn621
Babkina, O.V., Evstaf’eva, A.G., Chichkova, N.V., Vartapetian, A.B., Müller, S., Baskunov, V.B., Piatrauskene, O.V., Kochetkov, S.N., and Gromova E.S., Mol. Biol. (Moscow), 2000, vol. 34, pp. 913–920. https://doi.org/10.1023/A:1026688028427
Ferrer, E., Wiersma, M., Kazimierczak, B., Müller, C.W., and Eritja, R., Bioconjug. Chem., 1997, vol. 8, pp. 757–761. https://doi.org/10.1021/bc970042l
Petrauskene, O.V., Babkina, O.V., Tashlitsky, V.N., Kazankov, G.M., and Gromova, E.S., FEBS Lett., 1998, vol., 425, pp. 29–34. https://doi.org/10.1016/s0014-5793(98)00184-7
Dzhidzhoev, M.S., Platonenko, V.T., and Popov, V.K., Moscow Univ. Phys. Bull., 1984, vol. 39, pp. 118–120.
ACKNOWLEDGMENTS
We are grateful to T. Zatsepin for recording of MALDI mass spectra.
Funding
This work was supported by the Russian Foundation for Fundamental Investigations under Grant no. 19-04-00533.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This article does not contain any studies involving human participants or animals performed by any of the authors.
Conflict of Interests
The authors declare no conflict of interest.
Additional information
Abbreviations: DTT, dithiothreitol; IBA, o-iodosobenzoic acid; IdC, 5-iodo-2'-deoxycytidine; IdU, 5-iodo-2'-deoxyuridine; NCS, N-chlorosuccinimide; NTCBA, 2-nitro-5-thiocyanobenzoic acid; ON, oligodeoxynucleotide; PAGE, polyacrylamide gel electrophoresis; SDS, sodium dodecylsulfate; Tris, tris(hydroxymethyl)aminomethane.
Corresponding author: phone: +7 (495) 939-31-44; fax: +7 (495) 939-31-81.
Rights and permissions
About this article
Cite this article
Kirsanova, O.V., Subach, F.V., Loiko, A.G. et al. EcoRII Restriction Endonuclease Forms Specific Contacts to the Bases of Its Target Sequence Flipped from DNA in a Transition Complex with Photoactivatable Substrates. Russ J Bioorg Chem 47, 367–375 (2021). https://doi.org/10.1134/S106816202102014X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S106816202102014X