Abstract
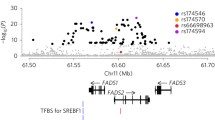
Since ancient times, meat and fat of seals and whales, rich in omega-3 polyunsaturated fatty acids, prevailed in the traditional “Arctic” diet of the indigenous people of the far Northeast of Siberia (Eskimos, Chukchi, Koryaks). This was reflected in the peculiarities of lipid metabolism of northern aborigines—for example, among them, variants of the genes for fatty acid desaturases (FADS1 and FADS2) encoding enzymes with reduced activity are predominantly widespread. In this study, we investigated the distribution of 22 bp insertion of the FADS2 gene (rs66698963), which affects the expression of the FADS1 gene, in populations of the northeastern (Koryaks, Evens) and southern (Buryats) parts of Siberia. The Koryaks had a minimum frequency of 22 bp insertions (5.6%)—almost an order of magnitude less frequently than Buryats (45.3%). In addition, a high frequency (10.9%) of nonsense mutation in the CYB5R2 gene (position 7 694 023 of chromosome 11) was detected in the Koryaks. This mutation leads to premature termination of the synthesis of NADH cytochrome b5 reductase, which is necessary for the functioning of desaturases encoded by FADS genes. The revealed genetic features of the indigenous population of the Far North are explained by long-term adaptation to a traditional diet very rich in lipids, which is a reason that prevents the need for additional synthesis of polyunsaturated fatty acids using fatty acid desaturases.

Similar content being viewed by others
REFERENCES
Marquardt, A., Stöhr, H., White, K., and Weber, B.H., cDNA cloning, genomic structure, and chromosomal localization of three members of the human fatty acid desaturase family, Genomics, 2000, vol. 66, pp. 175–183. https://doi.org/10.1006/geno.2000.6196
Los’, D.A., Structure, regulation of expression and functioning of fatty acid desaturases, Usp. Biol. Khim., 2001, vol. 41, pp. 163–198.
Malyarchuk, B.A., Long-term gene—environment interactions and genetics of metabolic disorders in aboriginal populations of Northeast Asia, Ecol. Genet., 2018, vol. 16, no. 2, pp. 30–35. https://doi.org/10.17816/ecogen16230-35
Ameur, A., Enroth, S., Johansson, A., et al., Genetic adaptation of fatty-acid metabolism: a human specific haplotype increasing the biosynthesis of long-chain omega-3 and omega-6 fatty acids, Am. J. Hum. Genet., 2012, vol. 90, pp. 809–820. https://doi.org/10.1016/j.ajhg.2012.03.014
Reardon, H.T., Zhang, J., Kothapalli, K.S., et al., Insertion-deletions in a FADS2 intron 1 conserved regulatory locus control expression of fatty acid desaturases 1 and 2 and modulate response to simvastatin, Prostaglandins, Leukotrienes Essent. Fatty Acids, 2012, vol. 87, pp. 25–33. https://doi.org/10.1016/j.plefa.2012.04.011
Voruganti, V.S., Higgins, P.B., Ebbesson, S.O., et al., Variants in CPT1A, FADS1, and FADS2 are associated with higher levels of estimated plasma and erythrocyte delta-5 desaturases in Alaskan Eskimos, Front. Genet., 2012, vol. 3, p. 86. eCollection 2012.https://doi.org/10.3389/fgene.2012.00086
Mathieson, I., Lazaridis, I., Rohland, N., et al., Genome-wide patterns of selection in 230 ancient Eurasians, Nature, 2015, vol. 528, pp. 499–503. https://doi.org/10.1038/nature16152
Ye, K., Gao, F., Wang, D., et al., Dietary adaptation of FADS genes in Europe varied across time and geography, Nat. Ecol. Evol., 2017, vol. 1, p. 0167. https://doi.org/10.1038/s41559-017-0167
Mathieson, S. and Mathieson, I., FADS1 and the timing of human adaptation to agriculture, Mol. Biol. Evol., 2018, vol. 35, pp. 2957–2970. https://doi.org/10.1093/molbev/msy180
Mathias, R.A., Fu, W., Akey, J.M., et al., Adaptive evolution of the FADS gene cluster within Africa, PLoS One, 2012, vol. 7. e44926. https://doi.org/10.1371/journal.pone.0044926
Amorim, C.E., Nunes, K., Meyer, D., et al., Genetic signature of natural selection in first Americans, Proc. Natl. Acad. Sci. U.S.A., 2017, vol. 114, pp. 2195–2199. https://doi.org/10.1073/pnas.1620541114
Malyarchuk, B.A. and Derenko, M.V., Polymorphism of polyunsaturated fatty acid metabolism genes (FADS1 and FADS2) in the indigenous population of Siberia, Vestn. Sev.-Vost. Nauchn. Tsentra Dal’nevost. Otd. Ross. Akad. Nauk, 2018, no. 3, pp. 106–111.
Fumagalli, M., Moltke, I., Grarup, N., et al., Greenlandic Inuit show genetic signatures of diet and climate adaptation, Science, 2015, vol. 349, pp. 1343–1347. https://doi.org/10.1126/science.aab2319
Harris, D.H., Ruczinski, I., Yanek, L.R., et al., Evolution of hominin polyunsaturated fatty acid metabolism: from Africa to the New World, Genome Biol. Evol., 2019, vol. 11, pp. 1417–1430. https://doi.org/10.1093/gbe/evz071
Cardona, A., Pagani, L., Antao, T., et al., Genome-wide analysis of cold adaption in indigenous Siberian populations, PLoS One, 2014, vol. 9. e98076. eCollection 2014https://doi.org/10.1371/journal.pone.0098076
Clemente, F.J., Cardona, A., Inchley, C.E., et al., A selective sweep on a deleterious mutation in the CPT1A gene in Arctic populations, Am. J. Hum. Genet., 2014, vol. 95, pp. 584–589. https://doi.org/10.1016/j.ajhg.2014.09.016
Hsieh, P., Hallmark, B., Watkins, J., et al., Exome sequencing provides evidence of polygenic adaptation to a fat-rich animal diet in indigenous Siberian populations, Mol. Biol. Evol., 2017, vol. 34, pp. 2913–2926. https://doi.org/10.1093/molbev/msx226
Reynolds, A.W., Mata-Míguez, J., Miró-Herrans, A., et al., Comparing signals of natural selection between three indigenous North American populations, Proc. Natl. Acad. Sci. U.S.A., 2019, vol. 116, pp. 9312–9317. https://doi.org/10.1073/pnas.1819467116
Mathieson, I., Limited evidence for selection at the FADS locus in Native American populations, Mol. Biol. Evol., 2020, vol. 37, pp. 2029–2033. https://doi.org/10.1093/molbev/msaa064
Kothapalli, K.S.D., Ye, K., Gadgil, M.S., et al., Positive selection on a regulatory insertion—deletion polymorphism in FADS2 influences apparent endogenous synthesis of arachidonic acid, Mol. Biol. Evol., 2016, vol. 33, pp. 1726–1739. https://doi.org/10.1093/molbev/msw049
Moltke, I., Grarup, N., Jørgensen, M.E., et al., A common Greenlandic TBC1D4 variant confers muscle insulin resistance and type 2 diabetes, Nature, 2014, vol. 512, pp. 190–193. https://doi.org/10.1038/nature13425
Grarup, N., Moltke, I., Andersen, M.K., et al., Loss-of-function variants in ADCY3 increase risk of obesity and type 2 diabetes, Nat. Genet., 2018, vol. 50, pp. 172–174.https://doi.org/10.1038/s41588-017-0022-7
Greenberg, C.R., Dilling, L.A., Thompson, G.R., et al., The paradox of the carnitine palmitoyltransferase type Ia P479L variant in Canadian aboriginal populations, Mol. Genet. Metab., 2009, vol. 96, pp. 201–207. https://doi.org/10.1016/j.ymgme.2008.12.018
Zhou, S., Xiong, L., Xie, P., et al., Increased missense mutation burden of fatty acid metabolism related genes in Nunavik Inuit population, PLoS One, 2015, vol. 10. e0128255. https://doi.org/10.1371/journal.pone.0128255
Pagani, L., Lawson, D.J., Jagoda, E., et al., Genomic analyses inform on migration events during the peopling of Eurasia, Nature, 2016, vol. 538, pp. 238–242. https://doi.org/10.1038/nature19792
Mörseburg, A., Investigating the role of demography and selection in genome scale patterns of common and rare variant diversity in humans, Ph. D. Thesis, Cambridge: Univ. Cambridge, 2018.
Untergasser, A., Cutcutache, I., Koressaar, T., et al., Primer3—new capabilities and interfaces, Nucleic Acids Res., 2012, vol. 40. e115. https://doi.org/10.1093/nar/gks596
Tamura, K., Peterson, D., Peterson, N., et al., MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods, Mol. Biol. Evol., 2011, vol. 28, pp. 2731–2739. https://doi.org/10.1093/molbev/msr121
Excoffier, L., Laval, G., and Schneider, S., Arlequin (version 3.0): an integrated software package for population genetics data analysis, Evol. Bioinf. Online, 2007, vol. 1, pp. 47–50.
Vasil’evskii, R.S., Proiskhozhdenie i drevnyaya kul’tura koryakov (Origin and Ancient Culture of the Koryaks), Novosibirsk: Nauka, 1971.
Narody Severo-Vostoka Sibiri (Peoples of the North-East of Siberia), Bat’yanova, E.P. and Turaev, V.A., Eds., Moscow: Nauka, 2010.
Seo, J.S., Ju, Y.S., Lee, W.C., et al., The transcriptional landscape and mutational profile of lung adenocarcinoma, Genome Res., 2012, vol. 22, pp. 2109–2119. https://doi.org/10.1101/gr.145144.112
Martinelli, N., Girelli, D., Malerba, G., et al., FADS genotypes and desaturase activity estimated by the ratio of arachidonic acid to linoleic acid are associated with inflammation and coronary artery disease, Am. J. Clin. Nutr., 2008, vol. 88, pp. 941–949. https://doi.org/10.1093/ajcn/88.4.941
Lattka, E., Illig, T., Heinrich, J., and Koletzko, B., Do FADS genotypes enhance our knowledge about fatty acid related phenotypes?, Clin. Nutr., 2010, vol. 29, pp. 277–287. https://doi.org/10.1016/j.clnu.2009.11.005
Chen, Y., Estampador, A.C., Keller, M., et al., The combined effects of FADS gene variation and dietary fats in obesity-related traits in a population from the far north of Sweden: the GLACIER study, Int. J. Obes. (London), 2019, vol. 43, pp. 808–820. https://doi.org/10.1038/s41366-018-0112-3
Del Gobbo, L.C., Imamura, F., Aslibekyan, S., et al., ω-3 polyunsaturated fatty acid biomarkers and coronary heart disease: pooling project of 19 cohort studies, JAMA Int. Med., 2016, vol. 176, pp. 1155–1166. https://doi.org/10.1001/jamainternmed.2016.2925
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare that they have no conflict of interest.
Statement of compliance with standards of research involving humans as subjects. All procedures performed in a study involving people comply with the ethical standards of the institutional and/or national committee for research ethics and the 1964 Helsinki Declaration and its subsequent changes or comparable ethical standards.
Informed voluntary consent was obtained from each of the participants.
Rights and permissions
About this article
Cite this article
Malyarchuk, B.A., Derenko, M.V. & Denisova, G.A. Adaptive Changes in Fatty Acid Desaturation Genes in Indigenous Populations of Northeast Siberia. Russ J Genet 57, 1461–1466 (2021). https://doi.org/10.1134/S1022795421120103
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795421120103