Abstract
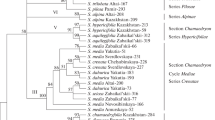
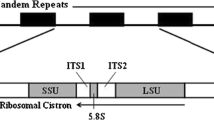
The variability of nucleotide sequences of ribosomal genes and internal transcribed spacers located in the ITS1–5.8S rRNA–ITS2a–2S rRNA–ITS2 region separating the 18S and 28S rRNA genes in the rRNA gene cluster was studied in order to estimate the possibility of using these sequences as molecular markers for species identification and reconstruction of the phylogeny of the family Chironomidae. It was established that, in species of the family Chironomidae, the content of AT base pairs in the sequences of the 5.8S rRNA and 2S rRNA genes is lower (50–57%) than in the sequences of the ITS1, ITS2a, and ITS2 transcribed spacers (64–71%). The sizes of nucleotide sequences of the 5.8S rRNA (123 bp) and 2S rRNA (30 bp) genes in species of the family Chironomidae are conservative, while the sizes of the ITS1, ITS2a, and ITS2 internal transcribed spacers vary both within the species and between species. Neither intraspecific, nor interspecific, nor intergeneric nucleotide differences were found in the sequences of the 2S rRNA gene in 27 species from five genera of the family Chironomidae. Nucleotide sequences of the 5.8S rRNA gene are identical in most studied species of the Chironomus genus. Fixed interspecific nucleotide substitutions were found only in two sites of this gene sequence in three species: C. piger, C. riparius (473С>T), and C. annularius (474A>G). The number of insertions, deletions, and nucleotide substitutions in the sequences of the ITS1, ITS2a, and ITS2 internal transcribed spacers and the whole ITS1–5.8S rRNA–ITS2a–2S rRNA–ITS2 region is generally low within species and increases 5–6 times in sibling species and more than 10 times in morphologically distinct species; that is, it increases in accordance with the growth of taxonomic status of species from the family Chironomidae. The data obtained indicate that nucleotide sequences of ribosomal genes and internal transcribed spacers from the ITS1–5.8S rRNA–ITS2a–2S rRNA–ITS2 region of the rRNA gene cluster can be used as molecular markers for species identification and reconstruction of the phylogeny of the family Chironomidae.

Similar content being viewed by others
REFERENCES
Ferrington, L.C., Global diversity of non-biting midges (Chironomidae; Insecta—Diptera) in freshwater, Hydrobiologia, 2008, vol. 595, no. 1, pp. 447—455. https://doi.org/10.1007/s10750-007-9130-1
Saether, O.A., Phylogeny of the subfamilies of Chironomidae (Diptera), Syst. Entomol., 2000, vol. 25, no. 3, pp. 393—403. https://doi.org/10.1046/j.1365-3113.2000.00111.x
Wiederholm, T., Chironomidae of the Holarctic Region: Keys and Diagnoses, Part 1: Larvae, in Ent. Scand. Suppl., 1983, vol. 19.
Shobanov, N.A., Shilova, A.I., and Belyanina, S.I., Volume and structure of the genus Chironomus Meig. (Diptera, Chironomidae): a review of the world’s fauna, in Ekologiya, evolyutsiya i sistematika khironomid (Ecology, Evolution, and Taxonomy of Chironomids), Tol’yatti: Inst. Biol. Vnutrennikh Vod and Inst. Ekol. Volzh. Basseina Russ. Akad. Nauk, 1996, pp. 44—96.
Lindeberg, B. and Wiederholm, T., Notes on the taxonomy of European species of Chironomus (Diptera Chironomidae), Ent. Scand. Suppl., 1979, vol. 10, pp. 99—116.
Keyl, H.-G., Chromosomenevolution bei Chironomus: II. Chromosomenumbauten und phylogenetische Beziehungen der Arten, Chromosoma, 1962, vol. 13, no. 4, pp. 464—514. https://doi.org/10.1007/BF00327342
Kiknadze, I.I., Shilova, A.I., Kerkis, I.E., et al., Kariotipy i morfologiya lichinok triby Chironomini: atlas (Karyotypes and Larval Morphology of the Midges of the Tribe Chironomini: An Atlas), Novosibirsk: Nauka, 1991.
Kiknadze, I., Istomina, A., Golygina, V., and Gunderina, L., Karyotypes of Palearctic and Holarctic Species of the Genus Chironomus, Novosibirsk: GEO, 2016.
Martin, J., Australian, North American, New Zealand and Oriental Chironomus species. http://www.chironomidae.net/Martin/JMRes.html. Accessed May 11, 2019.
Wuelker, W., Devai, Gy., and Devai, I., Computer assisted studies of chromosome evolution in the genus Chironomus (Dipt.) comparative and integrated analysis of chromosome arms A, E and F, Acta Biol. Debr. Oecol. Hung., 1989, vol. 2, pp. 373—387.
Gunderina, L.I., Kiknadze, I.I., Istomina, A.G., et al., Divergence of the polytene chromosome banding sequences as a reflection of evolutionary rearrangements of the genome linear structure, Russ. J. Genet., 2005, vol. 41, no. 2, pp. 130—137. https://doi.org/10.1007/s11177-005-0036-6
Shobanov, N.A., Evolution of the genus Chironomus (Diptera, Chironomidae): 2. A phylogenetic model, Zool. Zh., 2002, vol. 81, no. 6, pp. 711—718.
Kiknadze, I.I., Gunderina, L.I., Butler, M.G., et al., Chromosomes and continents, Biosphere Origin and Evolution, Dobretsov, N., Kolchanov, N., Rozanov, A., and Zavarin, G., Eds., New York: Springer-Verlag, 2008, pp. 349—369.
Guryev, V., Makarevitch, I., Blinov, A., and Martin, J., Phylogeny of the genus Chironomus (Diptera) inferred from sequences mitochondrial cytochrome b and cytochrome oxidase 1, Mol. Phylogenet. Evol., 2001, vol. 19, no.1, pp. 9—21. https://doi.org/10.1006/mpev.2001.0898
Martin, J., Guryev, V., and Blinov, A., Population variability in Chironomus (Camptochironomus) species (Diptera, Nematocera) with a Holarctic distribution: evidence of mitochondrial gene flow, Insect Mol. Biol., 2002, vol. 11, pp. 387—397. https://doi.org/10.1046/j.1365-2583.2002.00348.x27
Proulx, I., Martin, J., Carew, M., and Hare, L., Using various lines of evidence to identify Chironomus species (Diptera: Chironomidae) in eastern Canadian lakes, Zootaxa, 2013, vol. 3741, pp. 401—458. https://doi.org/10.11646/zootaxa.3741.4.1
Gunderina, L.I. and Katokhin, A.V., Variation and divergence of the rDNA ITS-1 region in species of the genus Chironomus (Diptera: Chironomidae), Contemporary Chironomid Studies (Proc. XVII Int. Symp. on Chironomidae, 2009, Nankai University, China), Nankai: Nankai University Press, 2011, pp. 22—35.
Asari, H., Kasuya, S., Kobayashi, T., et al., Identification of closely related Hydrobaenus species (Diptera: Chironomidae) using the second internal transcribed spacer (ITS2) region of ribosomal DNA, Aquat. Insects, 2004, vol. 26, nos. 3—4, pp. 207—213. https://doi.org/10.1080/01650420412331327295
Kaga, K., Kasuya, S., Kobayashi, T., and Ohtaka, A., Identification of chironomid species by DNA sequence—especially genus Hydrobaenus including H. sp. “Tsugaru”, in Contemporary Chironomid Studies (Proc. XVII Int. Symp. on Chironomidae, 2009, Nankai University, China), Nankai: Nankai University Press, 2011, pp. 36—40.
Schlötterer, Ch., Hauser, M.-T., von Haeseler, A., and Tautz, D., Comparative evolutionary analysis of rDNA ITS regions in Drosophila, Mol. Biol. Evol., 1994, vol. 11, no. 3, pp. 513—522.
Tautz, D., Hancock, J.M., Webb, D.A., et al., Complete sequences of the rRNA genes of Drosophila melanogaster, Mol. Biol. Evol., 1988, vol. 5, no. 4, pp. 366—376.
Friedrich, M. and Tautz, D., An episodic change of rDNA nucleotide substitution rate has occurred during the emergence of the insect order Diptera, Mol. Biol. Evol., 1997, vol. 14, no. 6, pp. 644—653.
Stage, D.E. and Eickbush, T.H., Sequence variation within the rRNA gene loci of 12 Drosophila species, Genome Res., 2007, vol. 17, no. 12, pp. 1888—1897. https://doi.org/10.1101/gr.6376807
Matsumoto, Y., Yanase, T., Tsuda, T., and Noda, H., Characterization of internal transcribed spacer (ITS1)—ITS2 region of ribosomal RNA gene from 25 species of Culicoides biting midges (Diptera: Ceratopogonidae) in Japan, J. Med. Entomol., 2009, vol. 46, no. 5, pp. 1099—1108. https://doi.org/10.1603/033.046.0517
LaRue, B., Gaudreau, C., Bagre, H.O., and Charpentier, G., Generalized structure and evolution of ITS1 and ITS2 rDNA in black flies (Diptera: Simuliidae), Mol. Phylogenet. Evol., 2009, vol. 53, no. 3, pp. 749—757. https://doi.org/10.1016/j.ympev.2009.07.032
Tatonova, Y.V., Chelomina, G.N., and Besprosvannykh, V.V., Genetic diversity of nuclear ITS1–5.8S–ITS2 rDNA sequence in Clonorchis sinensis Cobbold, 1875 (Trematoda: Opisthorchidae) from the Russian Far East, Parasitol. Int., 2012, vol. 61, pp. 664—674. https://doi.org/10.1016/j.parint.2012.07.005
Coleman, A.W., ITS2 is a double-edged tool for eukaryote evolutionary comparisons, Trends Genet., 2003, vol. 19, no. 7, pp. 370—375. https://doi.org/10.1016/S0168-9525(03)00118-5
Coleman, A.W., Pan-eukaryote ITS2 homologies revealed by RNA secondary structure, Nucleic Acids Res., 2007, vol. 35, pp. 3322—3329. https://doi.org/10.1093/nar/gkm233
Wang, X.C., Liu, C., Huang, L., et al., ITS1: a DNA barcode better than ITS2 in eukaryotes?, Mol. Ecol. Resour., 2015, vol. 15, pp. 573—586. https://doi.org/10.1111/1755-0998.12325
Rozen, S. and Skaletsky, H.J., Primer 3 on the WWW for general users and for biologist programmers, Methods Mol. Biol., 2000, vol. 132, pp. 365—386.
Edgar, R.C., MUSCLE: Multiple sequence alignment with high accuracy and high throughput, Nucleic Acids Res., 2004, vol. 32, no. 5, pp. 1792—1797. https://doi.org/10.1093/nar/gkh340
Tamura, K., Stecher, G., Peterson, D., et al., MEGA6: molecular evolutionary genetics analysis version 6.0, Mol. Biol. Evol., 2013, vol. 30, no. 12, pp. 2725—2729. https://doi.org/10.1093/molbev/mst197
Gunderina, L.I., Species-specific PCR primers for identification of the sibling species Chironomus plumosus (Linnaeus, 1758) and Chironomus balatonicus (Devai, Wuelker et Scholl, 1983) (Chironomidae, Diptera), Fauna Norv., 2012, vol. 32, pp. 151—157. https://doi.org/10.5324/fn.v31i0.1381
Gunderina, L.I., Design of molecular markers for the identification of species of the genus Chironomus (Diptera, Chironomidae), Entmol. Rev., 2014, vol. 94, no. 1, pp. 140—148. https://doi.org/10.1134/S0013873814010151
Funding
This work was supported by the budgetary project no. 0324-2019-0042.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by A. Barkhash
Rights and permissions
About this article
Cite this article
Gunderina, L.I., Katokhin, A.V. Variability of Nucleotide Sequences in the ITS1–5.8S rRNA–ITS2a–2S rRNA–ITS2 Region of rRNA Gene Cluster in Species of the Family Chironomidae. Russ J Genet 56, 916–925 (2020). https://doi.org/10.1134/S1022795420080050
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795420080050