Abstract
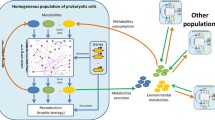
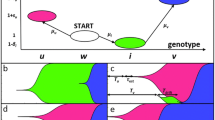
The study focuses on the mechanisms of crossing the valleys of fitness by a population of haploid microorganisms, whose fitness depends on allelic values at two different loci and is determined by a complex landscape, the shape of which corresponds to the pattern “a mountain in the field surrounded by a trench”; these mechanisms are analyzed using computer modeling. We have studied the influence of various biological factors on the evolutionary perspective of microbial colonies, the reproduction rate of which is controlled by a protein consisting of two subunits encoded at different loci. Molecular genetic (mutation rate, affinity of subunits), population (fitness function landscape and population density), and ecological (concentration of available substrate in the habitat, flow intensity) factors have been considered. Our results demonstrate that the difference in fitness for various allelic combinations, while determining the shape of the fitness landscape, sets the optimal mutation rates to overcome its valleys and opens a window of opportunity for the evolution of the population toward the state of the highest average fitness. Moreover, depending on the fitness landscape type, either gradual or saltational evolutionary regimes are optimal for reaching the peak.





Similar content being viewed by others
REFERENCES
Epistasis, Moore, J.H. and Williams, S.M., Eds., New York: Springer-Verlag, 2015, vol. 1253. https://doi.org/10.1007/978-1-4939-2155-3
Kimura, M., The role of compensatory neutral mutations in molecular evolution, J. Genet., 1985, vol. 64, no. 1, pp. 7—19. https://doi.org/10.1007/BF02923549
Wright, S., Evolution and the Genetics of Population, vol. 3: Experimental Results and Evolutionary Deductions, Chicago: University of Chicago Press, 1984.
Whitlock, M.C., Phillips, P.C., Moore, F.B.-G., and Tonsor, S.J., Multiple fitness peaks and epistasis, Annu. Rev. Ecol. Syst., 1995, vol. 26, pp. 601—629.
Altukhov, Yu.P., Geneticheskie protsessy v populyatsiyakh (Genetic Processes in Populations), Moscow: Akademkniga, 2003, 3rd ed.
Kuchner, O. and Arnold, F.H., Directed evolution of enzyme catalysts, Trends Biotechnol., 1997, vol. 15, no. 12, pp. 523—530. https://doi.org/10.1016/S0167-7799(97)01138-4
Gatenby, R.A. and Vincent, T.L., Application of quantitative models from population biology and evolutionary game theory to tumor therapeutic strategies, Mol. Cancer Ther., 2003, vol. 2, no. 9, pp. 919—927.
Turelli, M., Barton, N.H., and Coyne, J.A., Theory and speciation, Trends Ecol. Evol., 2001, vol. 16, no. 7, pp. 330—343. https://doi.org/10.1016/S0169-5347(01)02177-2
Domingo, E. and Holland, J.J., RNA virus mutations and fitness for survival, Annu. Rev. Microbiol., 1997, vol. 51, no. 1, pp. 151—178. https://doi.org/10.1146/annurev.micro.51.1.151
Szendro, I.G., Schenk, M.F., Franke, J. et al., Quantitative analyses of empirical fitness landscapes, J. Stat. Mech. Theory Exp., 2013, vol. 2013, no. 1, p. P01005. https://doi.org/10.1088/1742-5468/2013/01/P01005
Meer, M.V., Kondrashov, A.S., Artzy-Randrup, Y., and Kondrashov, F.A., Compensatory evolution in mitochondrial tRNAs navigates valleys of low fitness, Nature, 2010, vol. 464, no. 7286, pp. 279—282. https://doi.org/10.1038/nature08691
Mackay, T.F.C., Epistasis and quantitative traits: using model organisms to study gene—gene interactions, Nat. Rev. Genet., 2013, vol. 15, no. 1, pp. 22—33. https://doi.org/10.1038/nrg3627
Handel, A. and Rozen, D.E., The impact of population size on the evolution of asexual microbes on smooth versus rugged fitness landscapes, BMC Evol. Biol., 2009, vol. 9, no. 1, p. 236. https://doi.org/10.1186/1471-2148-9-236
Whitlock, M.C., Griswold, C.K., and Peters, A.D., Compensating for the meltdown: the critical effective size of a population with deleterious and compensatory mutations, Ann. Zool. Fennici, 2003, vol. 40, no. 2, pp. 169—183.
Szendro, I.G., Franke, J., de Visser, J.A.G.M., and Krug, J., Predictability of evolution depends nonmonotonically on population size, Proc. Natl. Acad. Sci. U.S.A., 2013, vol. 110, no. 2, pp. 571—576. https://doi.org/10.1073/pnas.1213613110
Gokhale, C.S., Iwasa, Y., Nowak, M.A., and Traulsen, A., The pace of evolution across fitness valleys, J. Theor. Biol., 2009, vol. 259, no. 3, pp. 613—620. https://doi.org/10.1016/j.jtbi.2009.04.011
Weissman, D.B., Feldman, M.W., and Fisher, D.S., The rate of fitness-valley crossing in sexual populations, Genetics, 2010, vol. 186, no. 4, pp. 1389—1410. https://doi.org/10.1534/genetics.110.123240
Weissman, D.B., Desai, M.M., Fisher, D., and Feldman, M.W., The rate at which asexual populations cross fitness valleys, Theor. Popul. Biol., 2009, vol. 75, no. 4, pp. 286—300. https://doi.org/10.1016/j.tpb.2009.02.006
Haarsma, L.L., Nelesen, S., VanAndel, E., et al., Simulating evolution of protein complexes through gene duplication and co-option, J. Theor. Biol., 2016, vol. 399, pp. 22—32. https://doi.org/10.1016/j.jtbi.2016.03.028
Aita, T. and Husimi, Y., Fitness spectrum among random mutants on Mt. Fuji-type fitness landscape, J. Theor. Biol., 1996, vol. 182, no. 4, pp. 469—485. https://doi.org/10.1006/jtbi.1996.0189
Iordanskii, N.N., Evolyutsiya zhizni (The Evolution of Life), Moscow: Akademia, 2001.
Lashin, S.A., Suslov, V.V., and Matushkin, Y.G., Theories of biological evolution from the viewpoint of the modern systemic biology, Russ. J. Genet., 2012, vol. 48, no. 5, pp. 481— 496. https://doi.org/10.1134/S1022795412030064
Afonnikov, D.A., Oshchepkov, D.Y., and Kolchanov, N.A., Detection of conserved physico-chemical characteristics of proteins by analyzing clusters of positions with co-ordinated substitutions, Bioinformatics, 2001, vol. 17, no. 11, pp. 1035—1046. https://doi.org/10.1093/bioinformatics/17.11.1035
Dayhoff, M.O., Schwartz, R.M., and Orcutt, B.C., A model of evolutionary change in proteins, Atlas of Protein Sequence and Structure, vol. 5, suppl. 3, Dayhoff, M.O., Ed., Washington, DC: Natl. Biomed. Res. Found., 1978, pp. 345—352.
Ewens, W., Mathematical Population Genetics: 1. Theoretical Introduction, New York: Springer-Verlag, 2004. https://doi.org/10.1007/978-0-387-21822-9
Drake, J.W., Charlesworth, B., Charlesworth, D., and Crow, J.F., Rates of spontaneous mutation, Genetics, 1998, vol. 148, no. 4, pp. 1667—1686.
Lynch, M., Evolution of the mutation rate, Trends Genet., 2010, vol. 26, no. 8, pp. 345—352. https://doi.org/10.1016/j.tig.2010.05.003
Markel’, A.L., Stress and evolution, Russ. J. Genet. Appl. Res., 2008, vol. 12, nos. 1—2, pp. 206—215.
Foster, P.L., Stress responses and genetic variation in bacteria, Mutat. Res. Mol. Mech. Mutagen., 2005, vol. 569, nos. 1—2, pp. 3—11. https://doi.org/10.1016/j.mrfmmm.2004.07.017
Hoffmann, A.A. and Hercus, M.J., Environmental stress as an evolutionary force, Bioscience, 2000, vol. 50, no. 3, p. 217. https://doi.org/10.1641/0006-3568(2000)050[0217:ESAAEF]2.3.CO;2
Korogodin, V.I., Korogodina, V.L., Fajszi, C., et al., On the dependence of spontaneous mutation rates on the functional state of genes, Yeast, 1991, vol. 7, no. 2, pp. 105—117. https://doi.org/10.1002/yea.320070204
Il’ina, V.L., Korogodin, V.I., and Fajszi, C., Dependence of spontaneous reversion frequencies in adenin auxotrophic yeasts on the concentration of adenine in the medium, Genetika (Moscow), 1987, vol. 23, no. 4, pp. 637—642.
Lashin, S.A., and Matushkin, Y.G., Haploid evolutionary constructor: new features and further challenges, In Silico Biol., 2012, vol. 11, nos. 3—4, pp. 125—135. https://doi.org/10.3233/ISB-2012-0447
Lashin, S.A., Klimenko, A. I., Mustafin, Z.S., et al., HEC 2.0: improved simulation of the evolution of prokaryotic communities, Math. Biol. Bioinf., 2014, vol. 9, no. 2, pp. 585—596. https://doi.org/10.17537/2014.9.585
Lashin, S.A., Suslov, V.V., and Matushkin, Y.G., Comparative modeling of coevolution in communities of unicellular organisms: adaptability and biodiversity, J. Bioinf. Comput. Biol., 2010, vol. 08, no. 03, pp. 627—643. https://doi.org/10.1142/S0219720010004653
Lashin, S.A., Matushkin, Yu.G., Suslov, V.V., and Kolchanov, N.A., Evolutionary trends in the prokaryotic community and prokaryotic community-phage systems, Russ. J. Genet., 2011, vol. 47, no. 12, pp. 1487—1495. https://doi.org/10.1134/S1022795411110123
Klimenko, A.I., Matushkin, Yu.G., Kolchanov, N.A., and Lashin, S.A., Bacteriophages affect evolution of bacterial communities in spatially distributed habitats: a simulation study, BMC Microbiol., 2016, vol. 16, no. S1, p. S10. https://doi.org/10.1186/s12866-015-0620-4
Lashin, S.A., Matushkin, Yu.G., Suslov, V.V., and Kolchanov, N.A., Computer modeling of genome complexity variation trends in prokaryotic communities under varying habitat conditions, Ecol. Modell., 2012, vol. 224, no. 1, pp. 124—129. https://doi.org/10.1016/j.ecolmodel.2011.11.004
Klimenko, A.I., Matushkin, Yu.G., Kolchanov, N.A., and Lashin, S.A., Modeling evolution of spatially distributed bacterial communities : a simulation with the haploid evolutionary constructor, BMC Evol. Biol., 2015, vol. 15, p. S3. https://doi.org/10.1186/1471-2148-15-S1-S3
Wilke, C., Dynamic fitness landscapes in molecular evolution, Phys. Rep., 2001, vol. 349, no. 5, pp. 395—446. https://doi.org/10.1016/S0370-1573(00)00118-6
Kawashima, S., Pokarowski, P., Pokarowska, M., et al., AAindex: amino acid index database, progress report 2008, Nucleic Acids Res., 2007, vol. 36, database issue, pp. D202—D205. https://doi.org/10.1093/nar/gkm998
Funding
This study was supported by budget project no. 0324-2019-0040.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Lashin, S.A., Mustafin, Z.S., Klimenko, A.I. et al. Computer Simulation of the Evolution of Microbial Population: Overcoming Local Minima When Reaching a Peak on the Fitness Landscape. Russ J Genet 56, 242–252 (2020). https://doi.org/10.1134/S1022795420020076
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795420020076