Abstract
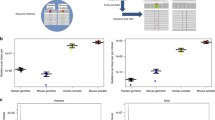
The emergence of genetic diseases and evolutionary processes are associated with the flow of genetic information from one generation to another, in which genetic information carried by gametes can be changed by the appearance of de novo germline mutations. The rate of germline mutations determines the rate of evolution and the incidence of heritable disorders. Despite the great theoretical and practical importance, the problem of establishing mutation rates and their dependence on different factors remains scarcely studied, and the mutation rate values obtained by different methods vary considerably. The review discusses different ways of estimating the rate of these mutations and makes an attempt to explain the reasons for discrepancies in the data obtained. Three levels of the mutation formation are considered: (1) mutations that are formed during the development of a given individual during gametogenesis (basic mutations); (2) mutations transmitted to offspring and determining differences in the genomes of consecutive generations (parents and offspring), which include basic mutations and possible changes resulting from complex processes of sperm transfer to oocyte, fertilization, and subsequent events that lead to only one viable offspring of hundreds of millions spermatozoa and oocytes; (3) mutations which are formed at level 2, fixed in evolution, and determine evolutionary processes and differences between genomes, in particular, of hominoids, hominids, and hominins.


Similar content being viewed by others
REFERENCES
Scally, A., Mutation rates and the evolution of germline structure, Philos. Trans. R. Soc. Lond., B, 2016, vol. 371, no. 1699. https://doi.org/10.1098/rstb.2015.0137
Lynch, M., Rate, molecular spectrum, and consequences of human mutation, Proc. Natl. Acad. Sci. U.S.A., 2010, vol. 107, no. 3, pp. 961—968. https://doi.org/10.1073/pnas.0912629107
Campbell, I.M., Yuan, B., Robberecht, C., et al., Parental somatic mosaicism is underrecognized and influences recurrence risk of genomic disorders, Am. J. Hum. Genet., 2014, vol. 95, no. 2, pp. 173—182. https://doi.org/10.1016/j.ajhg.2014.07.003
Rahbari, R., Wuster, A., Lindsay, S.J., et al., Timing, rates and spectra of human germline mutation, Nat. Genet., 2016, vol. 48, no. 2, pp. 126—133. https://doi.org/10.1038/ng.3469
Tang, W.W., Kobayashi, T., Irie, N., et al., Specification and epigenetic programming of the human germ line, Nat. Rev. Genet., 2016, vol. 17, no. 10, pp. 585—600. https://doi.org/10.1038/nrg.2016.88
Alekseenko, I.V., Kuzmich, A.I., Pleshkan, V.V., et al., The cause of cancer mutations: improvable bad life or inevitable stochastic replication errors?, Mol. Biol. (Moscow), 2016, vol. 50, no. 6, pp. 906—921. https://doi.org/10.7868/S0026898416060033
Sverdlov, E.D. and Mineev, K., Mutation rate in stem cells: an underestimated barrier on the way to therapy, Trends Mol. Med., 2013, vol. 19, no. 5, pp. 273—280. https://doi.org/10.1016/j.molmed.2013.01.004
Kondrashov, F.A. and Kondrashov, A.S., Measurements of spontaneous rates of mutations in the recent past and the near future, Philos. Trans. R. Soc. Lond., B, 2010, vol. 365, no. 1544, pp. 1169—1176. https://doi.org/10.1098/rstb.2009.0286
Lynch, M., Evolution of the mutation rate, Trends Genet., 2010, vol. 26, no. 8, pp. 345—352. https://doi.org/10.1016/j.tig.2010.05.003
Wang, J. and Song, Y., Single cell sequencing: a distinct new field, Clin. Transl. Med., 2017, vol. 6, no. 1, p. 10. https://doi.org/10.1186/s40169-017-0139-4
Golov, A.K., Razin, S.V., and Gavrilov, A.A., Single-cell genome-wide studies give new insight into nongenetic cell-to-cell variability in animals, Histochem. Cell Biol., 2016, vol. 146, no. 3, pp. 239—254. https://doi.org/10.1007/s00418-016-1466-z
Junker, J.P. and van Oudenaarden, A., Every cell is special: genome-wide studies add a new dimension to single-cell biology, Cell, 2014, vol. 157, no. 1, pp. 8—11. https://doi.org/10.1016/j.cell.2014.02.010
Wang, J., Fan, H.C., Behr, B., et al., Genome-wide single-cell analysis of recombination activity and de novo mutation rates in human sperm, Cell, 2012, vol. 150, no. 2, pp. 402—412. https://doi.org/10.1016/j.cell.2012.06.030
Kirkness, E.F., Grindberg, R.V., Yee-Greenbaum, J., et al., Sequencing of isolated sperm cells for direct haplotyping of a human genome, Genome Res., 2013, vol. 23, no. 5, pp. 826—832. https://doi.org/10.1101/gr.144600.112
Alex, B. and Moorjani, P., DNA dating: how molecular clocks are refining human evolution’s timeline, Conversation, 2017.
Lu, S., Zong, C., Fan, W., et al., Probing meiotic recombination and aneuploidy of single sperm cells by whole-genome sequencing, Science, 2012, vol. 338, no. 6114, pp. 1627—1630. https://doi.org/10.1126/science.1229112
Hou, Y., Fan, W., Yan, L., et al., Genome analyses of single human oocytes, Cell, 2013, vol. 155, no. 7, pp. 1492—1506. https://doi.org/10.1016/j.cell.2013.11.040
Wang, Y. and Navin, N.E., Advances and applications of single-cell sequencing technologies, Mol. Cell, 2015, vol. 58, no. 4, pp. 598—609. https://doi.org/10.1016/j.molcel.2015.05.005
Keightley, P.D., Rates and fitness consequences of new mutations in humans, Genetics, 2012, vol. 190, no. 2, pp. 295—304. https://doi.org/10.1534/genetics.111.134668
Acuna-Hidalgo, R., Veltman, J.A., and Hoischen, A., New insights into the generation and role of de novo mutations in health and disease, Genome Biol., 2016, vol. 17, no. 1, p. 241. https://doi.org/10.1186/s13059-016-1110-1
Segurel, L., Wyman, M.J., and Przeworski, M., Determinants of mutation rate variation in the human germline, Annu. Rev. Genomics Hum. Genet., 2014, vol. 15, pp. 47—70. https://doi.org/10.1146/annurev-genom-031714-125740
Kondrashov, A.S., Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases, Hum. Mutat., 2003, vol. 21, no. 1, pp. 12—27. https://doi.org/10.1002/humu.10147
de Ligt, J., Veltman, J.A., and Vissers, L.E., Point mutations as a source of de novo genetic disease, Curr. Opin. Genet. Dev., 2013, vol. 23, no. 3, pp. 257—263. https://doi.org/10.1016/j.gde.2013.01.007
Smith, T., Ho, G., Christodoulou, J., et al., Extensive variation in the mutation rate between and within human genes associated with Mendelian disease, Hum. Mutat., 2016, vol. 37, no. 5, pp. 488—494. https://doi.org/10.1002/humu.22967
Campbell, C.D. and Eichler, E.E., Properties and rates of germline mutations in humans, Trends Genet., 2013, vol. 29, no. 10, pp. 575—584. https://doi.org/10.1016/j.tig.2013.04.005
Lipson, M., Loh, P.R., Sankararaman, S., et al., Calibrating the human mutation rate via ancestral recombination density in diploid genomes, PLoS Genet., 2015, vol. 11, no. 11. e1005550. https://doi.org/10.1371/journal.pgen.1005550
Moorjani, P., Gao, Z., and Przeworski, M., Human germline mutation and the erratic evolutionary clock, PLoS Biol., 2016, vol. 14, no. 10. e2000744. https://doi.org/10.1371/journal.pbio.2000744
Conrad, D.F., Keebler, J.E., DePristo, M.A., et al., Variation in genome-wide mutation rates within and between human families, Nat. Genet., 2011, vol. 43, no. 7, pp. 712—714. https://doi.org/10.1038/ng.862
Kong, A., Frigge, M.L., Masson, G., et al., Rate of de novo mutations and the importance of father’s age to disease risk, Nature, 2012, vol. 488, no. 7412, pp. 471—475. https://doi.org/10.1038/nature11396
Jonsson, H., Sulem, P., Kehr, B., et al., Parental influence on human germline de novo mutations in 1548 trios from Iceland, Nature, 2017, vol. 549, no. 7673, pp. 519—522. https://doi.org/10.1038/nature24018
Itsara, A., Wu, H., Smith, J.D., et al., De novo rates and selection of large copy number variation, Genome Res., 2010, vol. 20, no. 11, pp. 1469—1481. https://doi.org/10.1101/gr.107680.110
Forsberg, L.A., Gisselsson, D., and Dumanski, J.P., Mosaicism in health and disease—clones picking up speed, Nat. Rev. Genet., 2017, vol. 18, no. 2, pp. 128—142. https://doi.org/10.1038/nrg.2016.145
Kloosterman, W.P., Francioli, L.C., Hormozdiari, F., et al., Characteristics of de novo structural changes in the human genome, Genome Res., 2015, vol. 25, no. 6, pp. 792—801. https://doi.org/10.1101/gr.185041.114
Moorjani, P., Amorim, C.E., Arndt, P.F., et al., Variation in the molecular clock of primates, Proc. Natl. Acad. Sci. U.S.A., 2016, vol. 113, no. 38, pp. 10607—10612. https://doi.org/10.1073/pnas.1600374113
Wilke, T., Schultheiß, R., and Albrecht, C., As time goes by: a simple fool’s guide to molecular clock approaches in invertebrates, Am. Malac. Bull., 2009, vol. 27, pp. 25—45. https://doi.org/10.4003/006.027.0203
Chen, C., Qi, H., Shen, Y., et al., Contrasting determinants of mutation rates in germline and soma, Genetics, 2017, vol. 207, no. 1, pp. 255—267. https://doi.org/10.1534/genetics.117.1114
Amster, G. and Sella, G., Life history effects on the molecular clock of autosomes and sex chromosomes, Proc. Natl. Acad. Sci. U.S.A., 2016, vol. 113, no. 6, pp. 1588—1593. https://doi.org/10.1073/pnas.1515798113
Der Sarkissian, C., Allentoft, M.E., Avila-Arcos, M.C., et al., Ancient genomics, Philos. Trans. R. Soc. Lond., B, 2015, vol. 370, no. 1660, p. 20130387. https://doi.org/10.1098/rstb.2013.0387
Meyer, M., Kircher, M., Gansauge, M.T., et al., A high-coverage genome sequence from an archaic Denisovan individual, Science, 2012, vol. 338, no. 6104, pp. 222—226. https://doi.org/10.1126/science.1224344
Shendure, J. and Akey, J.M., The origins, determinants, and consequences of human mutations, Science, 2015, vol. 349, no. 6255, pp. 1478—1483. https://doi.org/10.1126/science.aaa9119
Moorjani, P., Sankararaman, S., Fu, Q., et al., A genetic method for dating ancient genomes provides a direct estimate of human generation interval in the last 45 000 years, Proc. Natl. Acad. Sci. U.S.A., 2016, vol. 113, no. 20, pp. 5652—5657. https://doi.org/10.1073/pnas.1514696113
Segurel, L., Thompson, E.E., Flutre, T., et al., The ABO blood group is a trans-species polymorphism in primates, Proc. Natl. Acad. Sci. U.S.A., 2012, vol. 109, no. 45, pp. 18493—18498. https://doi.org/10.1073/pnas.1210603109
Francioli, L.C., Polak, P.P., Koren, A., et al., Genome-wide patterns and properties of de novo mutations in humans, Nat. Genet., 2015, vol. 47, no. 7, pp. 822—826. https://doi.org/10.1038/ng.3292
Goriely, A., Decoding germline de novo point mutations, Nat. Genet., 2016, vol. 48, no. 8, pp. 823—824. https://doi.org/10.1038/ng.3629
Goldmann, J.M., Wong, W.S., Pinelli, M., et al., Parent-of-origin-specific signatures of de novo mutations, Nat. Genet., 2016, vol. 48, no. 8, pp. 935—939. https://doi.org/10.1038/ng.3597
Crow, J.F., The origins, patterns and implications of human spontaneous mutation, Nat. Rev. Genet., 2000, vol. 1, no. 1, pp. 40—47. https://doi.org/10.1038/35049558
Maher, G.J., Rajpert-De Meyts, E., Goriely, A., et al., Cellular correlates of selfish spermatogonial selection, Andrology, 2016, vol. 4, no. 3, pp. 550—553. https://doi.org/10.1111/andr.12185
Narasimhan, V.M., Rahbari, R., Scally, A., et al., Estimating the human mutation rate from autozygous segments reveals population differences in human mutational processes, Nat. Commun., 2017, vol. 8, no. 1, p. 303. https://doi.org/10.1038/s41467-017-00323-y
Scally, A. and Durbin, R., Revising the human mutation rate: implications for understanding human evolution, Nat. Rev. Genet., 2012, vol. 13, no. 10, pp. 745—753. https://doi.org/10.1038/nrg3295
Caldararo, N., Denisovans, Melanesians, Europeans, and Neandertals: the confusion of DNA assumptions and the biological species concept, J. Mol. Evol., 2016, vol. 83, nos. 1—2, pp. 78—87. https://doi.org/10.1007/s00239-016-9755-7
Allentoft, M.E., Collins, M., Harker, D., et al., The half-life of DNA in bone: measuring decay kinetics in 158 dated fossils, Proc. Biol. Sci., 2012, vol. 279, no. 1748, pp. 4724—4733. https://doi.org/10.1098/rspb.2012.1745
Llamas, B., Valverde, G., Fehren-Schmitz, L., et al., From the field to the laboratory: controlling DNA contamination in human ancient DNA research in the high-throughput sequencing era, Sci. Technol. Archaeol. Res., 2017, vol. 3, no. 1, pp. 1—14. https://doi.org/10.1080/20548923.2016.1258824
Vernot, B., Tucci, S., Kelso, J., et al., Excavating Neandertal and Denisovan DNA from the genomes of Melanesian individuals, Science, 2016, vol. 352, no. 6282, pp. 235—239. https://doi.org/10.1126/science.aad9416
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated by N. Maleeva
Rights and permissions
About this article
Cite this article
Uspenskaya, N.Y., Akopov, S.B., Snezhkov, E.V. et al. The Rate of Human Germline Mutations—Variable Factor of Evolution and Diseases. Russ J Genet 55, 523–534 (2019). https://doi.org/10.1134/S1022795419050144
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795419050144