Abstract
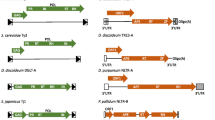
Non-Long Terminal Repeats (non-LTR or LINE) retrotransposons belong to the class of mobile genetic elements that are transposed into the host genome by reverse transcription of the RNA intermediate. Most of non-LTR retrotransposons contain two open reading frames (ORFs). The ORF1 codes for a gag-like protein, while the ORF2 codes for a reverse transcriptase (RT). We cloned two constructs based on Jockey-like non-LTR retrotransposon from genome Chironomus thummi (NLR1Cth). The retroposition assay performed in Chinese hamster ovary (CHO) cells demonstrated genome integrations of both constructs. The finding that the insect mobile element NLR1Cth is functional in mammalian cells demonstrates that this element possess universal enzymatic machinery allowing for active propagation in the genome of distant taxa. This suggests that the NLR1Cth transposon system may represent a useful tool for genetic analysis and manipulation in vertebrate cells.
Similar content being viewed by others
References
Berezikov, E., Bucheton, A., and Busseau, I., A Search for Reverse Transcriptase-Coding Sequences Reveals New Non-LTR Retrotransposons in the Genome of Drosophila melanogaster, Genome Biol., 2000, vol. 1, no. 6. P. research0012.1–12.15.
International Human Genome Sequencing Consortium: Initial Sequencing and Analysis of the Human Genome, Nature, 2001, vol. 409, pp. 860–921.
Moran, J.V. and Gilbert, N., Mammalian LINE-1 Retrotransposons and Related Elements, in: Mobile DNA II, Craig N., et al. Eds., Washington, DC: ASM Press, 2002, pp. 836–869.
Brouha, B., Schustak, J., Badge, R.M., et al., Hot L1s Account for the Bulk of Retrotransposition in the Human Population, Proc. Natl. Acad. Sci. USA, 2003, vol. 100, pp. 5280–5285.
Goodier, J.L., Ostertag, E.M., Du, K., and Kazazian, H.H., A Novel Active L1 Retrotransposon Subfamily in the Mouse, Genome Res., 2001, vol. 11, pp. 1677–1685.
Ostertag, E.M., DeBerardinis, R.J., Goodier, J.L., et al., A Mouse Model of Human L1 Retrotransposition, Nat. Genet., 2002, vol. 32, pp. 655–660.
Prak, E.T., Dodson, A.W., Farkash, E.A., and Kazazian, H.H., Jr., Tracking an Embryonic L1 Retrotrans-position Event, Proc. Natl. Acad. Sci. USA, 2003, vol. 100, pp. 1832–1837.
Muotri, A.R., Chu, V.T., Marchetto, M.C., et al., Somatic Mosaicism in Neuronal Precursor Cells Mediated by L1 Retrotransposition, Nature, 2005, vol. 435, pp. 903–910.
Kazazian, H.H., Wong, C., Youssoufian, H., et al., A Resulting from de novo Insertion of L1 Sequences Represents a Novel Mechanism for Mutation in Man, Nature, 1988, vol. 332, no. 6160, pp. 164–166.
Schwahn, U., Lenzner, S., Dong, J., et al., Positional Cloning of the Gene for X-Linked Retinitis Pigmentosa 2, Nat. Genet., 1998, vol. 19, pp. 327–332.
Ostertag, E.M., Prak, E.T., DeBerardinis, R.J., et al., Determination of L1 Retrotransposition Kinetics in Cultured Cells, Nucleic Acids Res., 2000, vol. 28, pp. 1418–1423.
Miki, Y., Nishisho, I., Horii, A., et al., Disruption of the APC Gene by a Retrotransposal Insertion of L1 Sequence in a Colon Cancer, Cancer Res., 1992, vol. 52, pp. 643–645.
Moran, J.V., Holmes, S.E., Naas, T.P., et al., High Frequency Retrotransposition in Cultured Mammalian Cells, Cell, 1996, vol. 87, no. 5, pp. 917–927.
Shi, X., Seluanov, A., and Gorbunova, V., Cell Divisions Are Required for L1 Retrotransposition, Mol. Cell. Biol., 2007, vol. 27, pp. 1264–1270.
Malik, S., Burke, W.D., and Eickbush, T.H., The Age and Evolution of Non-LTR Retrotransposable Element, Mol. Biol. Evol., 1999, vol. 16, no. 6, pp. 793–805.
Novikova, O. and Blinov, A., Origin, Evolution and Distribution of Non-LTR Retrotransposons among Eukaryotes, Russ. J. Genet., 2009, vol. 45, no. 2, pp. 129–138.
Luan, D.D., Korman, M.H., Jakubczak, J.L., and Eickbush, T.H., Reverse Transcription of R2Bm RNA is Primed by a Nick at the Chromosomal Target Site: A Mechanism for Non-LTR Retrotransposition, Cell, 1993, vol. 72, no. 4, pp. 595–605.
Luan, D.D. and Eickbush, T.H., RNA Template Requirements for Target DNA-Primed Reverse Transcription by the R2 Retrotransposable Element Mol. Cell Biol., 1995, vol. 15, no. 7, pp. 3882–3890.
Christensen, S. and Eickbush, T.H., Footprint of the Retrotransposon R2Bm Protein on Its Target Site before and after Cleavage, J. Mol. Biol., 2004, vol. 336, no. 5, pp. 1035–1045.
Christensen, S.M., Ye, J., and Eickbush, T.H., RNA from the 5’ End of the R2 Retrotransposon Controls R2 Protein Binding to and Cleavage of Its DNA Target Site, Proc. Natl. Acad. Sci. USA, 2006, vol. 103, no. 47, pp. 1762–1837.
Christensen, S.M., Bibillo, A., and Eickbush, T.H., Role of the Bombyx mori R2 Element N-Terminal Domain in the Target-Primed Reverse Transcription (TPRT) Reaction, Nucleic Acids Res., 2005, vol. 33, no. 20, pp. 6461–6468.
Blinov, A.G., Sobanov, Y.V., Bogachev, S.S., et al., The Chironomus thummi Genome Contains a Non-LTR Retrotransposon, Mol. Gen. Genet., 1993, vol. 237, no. 3, pp. 412–420.
Gruhl, M.C., Scherbik, S.V., Aimanova, K.G., et al., Insect Globin Gene Polymorphisms: Intronic Minisatellites and a Retroposon Interrupting Exon 1 of Homologous Globin Genes in Chironomus (Diptera), Gene, 2000, vol. 251, no. 2, pp. 153–163.
Heidmann, O. and Heidmann, T., Retrotransposition of a Mouse IAP Sequence Tagged with an Indicator Gene, Cell, 1991, vol. 64, no. 1, pp. 159–170.
Kajikawa, M. and Okada, N., LINEs Mobilize SINEs in the eel through a Shared 3′ Sequence, Cell, 2002, vol. 111, no. 3, pp. 433–444.
Matsumoto, T., Takahashi, H., and Fujiwara, H., Targeted Nuclear Import of Open Reading Frame 1 Protein Is Required for in vivo Retrotransposition of a Telomere-Specific Non-Long Terminal Repeat Retrotransposon, SART1, Mol. Cell Biol., 2004, vol. 24, no. 1, pp. 105–122.
Freeman, J.D., Goodchild, N.L., and Mager, D.L., A Modified Indicator Gene for Selection of Retrotrans-position Events in Mammalian Cells, Biotechniques, 1994, vol. 17, no. 1, pp. 46, 48–49, 52.
Beresikov, E., Novikova, O., Makarevich, I., et al., Evolutionary Relationships and Distribution of Non-LTR Retrotransposons in Eukaryotes, Proc. 5th Int. Conf. Bioinformatics of Genome Regulation and Structure-BGRS, Kolchanov, N.A., Ed., Inst. Cytol. Genet. Siberian Branch Russ. Acad. Sci., 2004, vol. 2, pp. 181–184.
Dombroski, B.A., Feng, Q., Mathias, S.L., et al., An in vivo Assay for the Reverse Transcriptase of Human Retrotransposon L1 in Saccharomyces cerevisiae, Mol. Cell. Biol., 1994, vol. 14, pp. 4485–4492.
Hirochika, H., Okamoto, H., and Kakutani, T., Silencing of Retrotransposons in Arabidopsis and Reactivation by the ddm1 Mutation, Plant Cell, 2000, vol. 12, no. 3, pp. 357–369.
Walsh, C.P., Chaillet, J.R., and Bestor, T.H., Transcription of IAP Endogenous Retroviruses is Constrained by Cytosine Methylation, Nat. Genet., 1998, vol. 20, no. 2, pp. 116–117.
Yoder, J.A., Walsh, C.P., and Bestor, T.H., Cytosine Methylation and the Ecology of Intragenomic Parasites, Trends Genet., 1997, vol. 13, no. 8, pp. 335–340.
Yu, F., Zingler, N., Schumann, G., and Stratling, W.H., Methyl-CpG-Binding Protein 2 Represses LINE-1 Expression and Retrotransposition but not Alu Transcription, Nucleic Acids Res., 2001, vol. 29, no. 21, pp. 4493–4501.
Nolan, T., Braccini, L., Azzalin, G., et al., The Post-Transcriptional Gene Silencing Machinery Functions Independently of DNA Methylation to Repress a LINE-1 Retrotransposon in Neurospora crassa, Nucleic Acids Res., 2005, vol. 33, no. 5, pp. 1564–1573.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Genetika, 2011, Vol. 47, No. 6, pp. 774–782.
The article is published in the original.
Recipient of a DAAD fellowship.
Rights and permissions
About this article
Cite this article
Novikova, O., Papusheva, E., Ponimaskin, E. et al. A retroposition assay for the NLR1Cth from midge Chironomus thummi genome in the Chinese hamster ovary cells. Russ J Genet 47, 682–690 (2011). https://doi.org/10.1134/S1022795411060147
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795411060147