Abstract
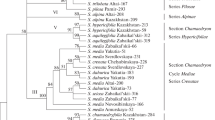
Comparative analysis of a group of closely related Drosophila species (D. virilis, D. lummei, D. novamexicana, D. americana texana, D. flavomontana, D. montana, D. borealis, D. lacicola, D. littoralis, D. kanekoi, and D. ezoana) was conducted based on an incomplete sequence of gene Ras1. The pattern of the relationships among the species corresponded to that expected from analysis of morphological and cytogenetic characters. Statistical data favoring neutrality of the substitutions examined in the Ras1 gene are presented. This character of the gene Ras1 evolution confers more reliability to reconstruction of phylogenetic relationships among closely related species. The resultant tree for main phylads of the group is as follows: D. virilis (D. lummei, D. montana, D. ezoana).
Similar content being viewed by others
References
Rasnitsyn, A.P., Classic and Nonclassic Systematics: Another View, Zh. Obshch. Biol., 2006, vol. 67, no. 5, pp. 385–388.
Spicer, G.S., Molecular Evolution and Phylogeny of the Drosophila virilis Species Group As Inferred by Two-Dimensional Electrophoresis, J. Mol. Evol., 1991, vol. 33, no. 4, pp. 379–394.
Spicer, G.S. and Bell, C.D., Molecular Phylogeny of the Drosophila virilis Species Group (Diptera: Drosophilidae) Inferred from Mitochondrial 12S and 16S Ribosomal RNA Genes, Ann. Entomol. Soc. Am., 2002, vol. 95, no. 2, pp. 156–161.
Civetta, A. and Singh, R.S., High Divergence of Reproductive Tract Proteins and Their Association with Postzygotic Reproductive Isolation in Drosophila melanogaster and Drosophila virilis Group Species, J. Mol. Evol., 1995, vol. 41, no. 6, pp. 1085–1095.
Caletka, B.C. and McAllister, B.F., A Genealogical View of Chromosomal Evolution and Species Delimitation in the Drosophila virilis Species Subgroup, Mol. Phylogenet. Evol., 2004, vol. 33, no. 3, pp. 664–670.
Kulikov, A.M., Mel’nikov, A.I., Gornostaev, N.G., et al., Morphometric Analysis of Male Genitalia in Sibling Species of Drosophila virilis Sturt., Russ. J. Genet., 2004, vol. 40, no. 2, pp. 125–138.
Sorokina, S.Yu., Mugue, N.S., Andrianov, B.V., and Mitrofanov, V.G., Variation of the 3′-Terminal Fragment of 16S rRNA Gene in Closely Related Species of Drosophila virilis Group, Russ. J. Genet., 2005, vol. 41, no. 8, pp. 853–858.
Patterson, J.T. and Stone, W.S., Evolution in the Genus Drosophila, New York: Macmillan, 1952.
Throckmorton, L.H, The virilis Species Group, in The Genetics and Biology of Drosophila, Ashburner, M. and Novitsky, E., Eds., London: Academic, 1982, pp. 227–296.
Tominaga, H. and Narise, S., Sequence Evolution of the Gpdh Gene in the Drosophila virilis Species Group, Genetics, 1995, vol. 96, no. 3, pp. 293–302.
Cohen, M.J., Evolution of 5S Ribosomal RNA Genes in the Chromosomes of the virilis Group of Drosophila, Chromosoma, 1976, vol. 55, no. 4, pp. 359–371.
Nurminsky, D.I., Moriyama, E.N., Lozovskaya, E.R., and Hartl, D.L., Molecular Phylogeny and Genome Evolution in the Drosophila virilis Species Group: Duplications of the Alcohol Dehydrogenase Gene, Mol. Biol. Evol., 1996, vol. 13, no. 1, pp. 132–149.
Gasperini, R. and Gibson, G., Absence of Protein Polymorphism in the Ras Genes of Drosophila melanogaster, J. Mol. Evol., 1999, vol. 49, no. 5, pp. 583–590.
Gloor, G.B., Preston, C.R., Johnson-Schlitz, D.M., et al., Type I Repressors of P Element Mobility, Genetics, 1993, vol. 135, pp. 81–95.
Welsh, J. and McClelland, M., Fingerprinting Genomes Using PCR with Arbitrary Primers, Nucleic Acids Res., 1990, vol. 18, no. 24, pp. 7213–7218.
Kumar, S., Tamura, K., and Nei, M., MEGA3: Integrated Software for Molecular Evolutionary Genetics Analysis and Sequence Alignment, Briefings Bioinf., 2004, no. 5, pp. 150–163.
Gu, X. and Zhang, J.J., A Simple Method for Estimating the Parameter of Substitution Rate Variation among Sites, Mol. Biol. Evol., 1997, no. 11, pp. 1106–1113.
Clement, M., Posada, D., and Crandall, K.A., TCS: A Computer Program to Estimate Gene Genealogies, Mol. Ecol., 2000, vol. 9, no. 10, pp. 1657–1660.
Takezaki, N., Rzhetsky, A., and Nei, M., Phylogenetic Tests of the Molecular Clock and Linearized Trees, Mol. Biol. Evol., 1995, vol. 12, no. 5, pp. 823–833.
UCSC Genome Bioinformatics—http://genomewiki.ucsc.edu.
Ohta, T. and Kimura, M., On the Constancy of the Evolutionary Rate of Cistrons, J. Mol. Evol., 1971, vol. 1, no. 1, pp. 18–25.
Valencia, A., Chardin, P., Wittinghofer, A., and Sander, C., The Ras Protein Family: Evolutionary Tree and Role of Conserved Amino Acids, Biochemistry, 1991, vol. 30, no. 19, pp. 4637–4648.
Neuman-Silberberg, F.S., Schejter, E., Hoffmann, F.M., and Shilo, B.Z., The Drosophila Ras Oncogenes: Structure and Nucleotide Sequence, Cell, 1984, vol. 37, no. 3, pp. 1027–1033.
Brock, H.W., Sequence and Genomic Structure of Ras Homologues Dmras85D and Dmras64B of Drosophila melanogaster, Gene, 1987, vol. 51, nos. 2–3, pp. 129–137.
Riley, R.M., Jin, W., and Gibson, G., Contrasting Selection Pressures on Components of the Ras-Mediated Signal Transduction Pathway in Drosophila, Mol. Ecol., 2003, vol. 12, no. 5, pp. 1315–1323.
Bergman, C.M., Pfeiffer, B.D., Limas, D.E., et al., Assessing the Impact of Comparative Genomic Sequence Data on the Functional Annotation of the Drosophila Genome, Genome Biol., 2002, vol. 3, no. 12, research 0086.1-0086.20.
Drton, M., Eriksson, N., and Leung, G., Ultra-Conserved Elements in Vertebrate and Fly Genomes, Algebraic Statistics for Computational Biology, Patcher, L. and Sturmfels, B., Eds., Cambridge: Cambridge Univ. Press, 2005, pp. 387–402.
Grantham, R., Gautier, G., Gouy, M., et al., Codon Catalog Usage and the Genome Hypothesis, Nucleic Acids Res., 1980, vol. 8, no. 1, pp. r49–r62.
Filipski, J., Evolution of DNA Sequence: Contributions of Mutation and Selection to the Origin of Chromosomal Compartments, Advances in Mutagenesis Research, Obe, G., Ed., Berlin: Springer Verlag, 1990, vol. 2, pp. 1–54.
Forsdyke, D.R., A Stem-Loop “Kissing” Model for the Initiation of Recombination and the Origin of Introns, Mol. Biol. Evol., 1995, vol. 12, no. 5, pp. 949–958.
Johnston, L.A. and Gallant, P., Control of Growth and Organ Size in Drosophila, BioEssays, 2002, vol. 24, no. 1, pp. 54–64.
Caldwell, P.E., Walkiewicz, M., and Stern, M., Ras Activity in the Drosophila Prothoracic Gland Regulates Body Size and Developmental Rate via Ecdysone Release, Curr. Biol., 2005, vol. 15, no. 20, pp. 1785–1795.
Wang, B.C., Park, J., Watabe, H.A., et al., Molecular Phylogeny of the Drosophila virilis Section (Diptera: Drosophilidae) Based on Mitochondrial and Nuclear Sequences, Mol. Phylogenet. Evol., 2006, vol. 40, no. 2, pp. 484–500.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © A.I. Chekunova, A.M. Kulikov, S.S. Mikhailovskii, O.E. Lazebny, I.V. Lazebnaya, V.G. Mitrofanov, 2008, published in Genetika, 2008, Vol. 44, No. 3, pp. 336–345.
Rights and permissions
About this article
Cite this article
Chekunova, A.I., Kulikov, A.M., Mikhailovskii, S.S. et al. The relationships among the species of the Drosophila virilis group inferred from the gene Ras1 sequences. Russ J Genet 44, 286–294 (2008). https://doi.org/10.1134/S1022795408030071
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795408030071