Abstract
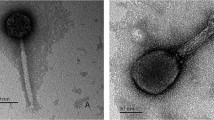
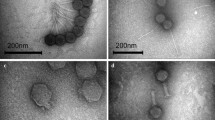
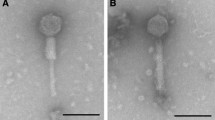
A new bacteriophage ϕK of microorganisms belonging to the genus Bordetella was isolated from cells of the earlier characterized strains 662-2 (1 and 2) obtained upon phage conversion of B. parapertussis 17 903 cells by B. pertussis bacteriophage ϕ134. Bacteriophage ϕK is identical to previously described Bordetella bacteriophages ϕT, ϕ134, and ϕ214 in morphology and some biological properties but has a permuted genome different from all other phages. DNA of bacteriophage ϕK is not integrated in the chromosome of B. parapertussis 17 903, similar to DNA of bacteriophages ϕT, ϕ134, and ϕ214 that are not integrated into B. pertussis and B. bronchiseptica chromosomes, but may be present in a small part of the bacterial population as linear plasmids. Sequences homologous to DNA of bacteriophage ϕK were detected in the chromosome of strain 662-2 (1 and 2) and in chromosomes of all tested strains B. pertussis and B. bronchiseptica. Prophage integration in chromosomes of microorganisms of the genus Bordetella may vary in different bacterial strains and species. An assumption about abortive lysogeny of B. parapertussis bacteria for ϕK phage and of B. pertussis and B. bronchiseptica for closely related phages ϕT, ϕ134, and ϕ214 has been advanced. The possibility of involvement of B. pertussis insertion sequences in the formation of the chromosomal structure in 662-2 convertants and in phage genomes is considered.
Similar content being viewed by others
References
Hendrix, R.W., Smith, C.M., Burns, R.N., et al., Evolutionary Relationships among Diverse Bacteriophages and Prophages: All the World’s a Phage, Proc. Natl. Acad. Sci. USA, 1999, vol. 96, pp. 2192–2197.
Brusson, H., Canchaya, C., and Hardt, W.-D., Phage and Evolution of Bacteria Pathogens: From Genomic Rearrangements to Lysogenic Conversion, Microbiol. Mol. Biol. Rev., 2004, vol. 68, pp. 560–602.
Canchaya, C., Proux, C., Fournous, G., et al., Prophage Genomics, Microbiol. Mol. Biol. Rev., 2003, vol. 67, pp. 238–276.
Krylov, V.N., Role of Horizontal Gene Transfer by Bacteriophages in the Origin of Pathogenic Bacteria, Rus. J. Genet., 2003, vol. 39, pp. 483–504.
Wagner, P.L. and Waldor, M.K., Bacteriophage Control of Bacterial Virulence, Infect. Immunol., 2002, vol. 70, pp. 3985–3993.
Lapaeva, I.A., Mebel’, S.M., Pereverzev, N.A., and Sinyashina, L.N., Bacteriophage of Bordetella pertussis, Zh. Mikrobiol. Epidemiol. Immunobiol., 1980, no. 5, pp. 85–90.
Pereverzev, N.A. and Sinyashina, L.N, Structural Organization of a Bacteriophage Isolated from Bordetella pertussis, Zh. Mikrobiol. Epidemiol. Immunobiol., 1981, no. 5, pp. 54–57.
Sinyashina, L.N., Lapaeva, I.A., and Mebel’, S.M., Transparent Solid Medium for Studying the Lytic Spectrum of Bacteriophages Active on Bacteria of the Genus Bordetella, Zh. Mikrobiol. Epidemiol. Immunobiol., 1982, no. 6, pp. 53–55.
Sinyashina, L.N., Lapaeva, I.A., and Mebel’, S.M., Characterization of Main Biological Properties of Bordetella Bacteriophages, Zh. Mikrobiol. Epidemiol. Immunobiol., 1982, vol. 8, pp. 66–69.
Lapaeva, I.A., Mebel’, S.M., Sinyashina, L.N., and Shakhvatova, O.Yu., Toxigenic Conversion by Pertussoid Phages in Bordetella parapertussis, Zh. Mikrobiol. Epidemiol. Immunobiol., 1982, no. 9, pp. 60–64.
Gol’tsmaier, T.A., Karataev, G.I., and Rozinov, M.N., Specifics of Lysogeny and Conversion in Bacteria of the Genus Bordetella, Zh. Mikrobiol. Epidemiol. Immunobiol., 1987, no. 5, pp. 9–13.
Karataev, G.I., Moskvina, I.L., Ryabinina, O.P., et al., Isolation and Characterization of a Bacteriophage from Phase I Vaccine Strain Tohama I, Mol. Genet. Microbiol. Virusol., 1988, no. 4, pp. 22–25.
Mebel, S., Lapaeva, I.A., Sinyashina, L.N., and Rustenbach, S., Changes within the Genus Bordetella Influenced by Phages, Proc. 4th Int. Symp. on Pertussis, Geneva, 1984, vol. 61, pp. 281–288.
Holzmayer, T.A., Karataev, G.I., Rozinov, M.N., et al., Bacteriophages of Bordetella sp.: Features of Lysogeny and Conversion, Zbl. Bakt. Hyg., 1988, vol. 269, pp. 147–155.
Shelton, C.B., Crosslin, D.R., Casey, J.L., et al., Purification and Characterization of a Temperate Transducing Bacteriophage for Bordetella avium, J. Bacteriol., 2000, vol. 82, pp. 6130–6136.
Liu, M., Deora, R., Doulatov, S., et al., Reverse Transcriptase-Mediated Tropism Switching in Bordetella Bacteriophage, Science, 2002, vol. 295, pp. 2091–2094.
Liu, M., Gingery, M., Doulatov, S.R., et al., Genomic and Genetic Analysis of Bordetella Bacteriophages Encoding Reverse Transcriptase-Mediated Tropism-Switching Cassettes, J. Bacteriol., 2004, vol. 186, pp. 1503–1517.
Maniatis, T., Fritsch, E.F., and Sambrook, J.E., Molecular Cloning: A Laboratory Manual, Cold Spring Harbor, New York: Cold Spring Harbor Lab., 1982.
Davis, R.W., Simon, H., and Devidson, N., Electron Microscope Heteroduplex Method for Mapping Regions of Base Sequence Homology in Nucleic Acids, Methods Enzymol., 1971, vol. 21, pp. 413–428.
Streisinger, G., Edgar, R.S., and Denhardt, G.H., Chromosome Structure in Phage T4: I. Circularity of the Linkage Map, Proc. Natl. Acad. Sci. USA, 1964, vol. 51, pp. 775–779.
Tye, B.K., Chan, R.K., and Botstein, D., Packaging of an Oversize Transducing Genome by Salmonella Phage P22, J. Mol. Biol., 1974, vol. 85, pp. 485–500.
Tye, B.K., Huberman, J.A., and Botstein, D., Non-Random Circular Permutation of Phage P22 DNA, J. Mol. Biol., 1974, vol. 85, pp. 501–532.
Vander, Byl. C. and Kropinski, A.M., Sequence of the Genome of Salmonella Bacteriophage P22, J. Bacteriol., 2000, vol. 182, pp. 6472–6481.
Casjens, S., Molecular Organization of the Bacteriophage P22 Coat Protein Shell, J. Mol. Biol., 1979, vol. 131, pp. 1–14.
MacHattie, L.A., Ritchie, D.A., Thomas, C.A., and Richardson, C.C., Terminal Repetition in Permuted T2 Bacteriophage DNA Molecules, J. Mol. Biol., 1967, vol. 23, pp. 355–363.
Lee, C.S., Davis, R.W., and Davidson, N., A Physical Study by Electron Microscopy of the Terminally Repetitious, Circularly Permuted DNA from the Coliphage Particles of Escherichia coli 15, J. Mol. Biol., 1970, vol. 48, pp. 1–22.
Ikeda, H. and Tomizawa, J., Prophage P1 and Extrachromosomal Replication Unit, Cold Spring Harb Symp. Quant. Biol., 1968, vol. 33, pp. 791–798.
Pokrovskaya, M.S., Karataev, G.I., and Smirnov, G.B., Specifics of the Formation and Properties of pLS Plasmids Resulting from Deletions from the Bacteriophage λatt80 (Tn9) Genome, Mol. Genet. Microbiol. Virusol., 1986, no. 1, pp. 12–18.
Gerald, G., Leffers, Jr., and Gottesman, S., λ Xis Degradation in Vivo by Lon and FtsH, J. Bacteriol., 1998, vol. 180, pp. 1573–1577.
Svarchevsky, A.N. and Rybchin, V.N., Characteristics of Plasmid Properties of Bacteriophage N15, Mol. Gen. Microbiol. Virusol., 1984, vol. 10, pp. 34–39.
Ravin, N.V., Kuprianov, V.V., Gilcrease, E.B., and Casjens, S.R., Bidirectional Replication from an Internal ori Site of the Linear N15 Plasmid Prophage, Nucleic Acids Res., 2003, vol. 31, pp. 6552–6560.
Stoppel, R.D., Meyer, M., and Schlegel, H.G., The Nickel Resistance Determinant Cloned from the Enterobacterium Klebsiella oxytoca: Conjugational Transfer, Expression, Regulation and DNA Homologies to Various Nickel-Resistant Bacteria, Biometals, 1995, vol. 8, pp. 70–79.
Hertwig, S., Klein, I., Lurz, R., et al., PY54, a Linear Plasmid Prophage of Yersinia enterocolitica with Covalently Closed Ends, Mol. Microbiol., 2003, vol. 48, pp. 989–1003.
Barbour, A.G. and Garon, C.F., Linear Plasmids of the Borrelia burgdorferi Have Covalently Closed Ends, Science, 1987, vol. 237, pp. 409–411.
Fraser, C.M., Casjens, S., Huang, W.M., et al., Genomic Sequence of a Lyme Disease Spirochaete, Borrelia burgdorferi, Nature, 1997, vol. 390, pp. 580–586.
Casjens, S., Murphy, M., van Vugt, R., et al., Telomeres of the Linear Chromosomes of the Lyme Disease Spirochetes: Nucleotide Sequence and Possible Exchange with Linear Plasmid Telomeres, Mol. Microbiol., 1997, vol. 26, pp. 581–596.
Barbour, A.G., Plasmid Analysis of Borrelia burgdorferi, the Lyme Disease Agent, J. Clin. Microbiol., 1988, vol. 26, pp. 475–478.
Grimm, D., Eggers, C.H., Caimano, M.J., et al., Experimental Assessment of the Roles of Linear Plasmids lp25 and lp28-1 of Borrelia burgdorferi throughout the Infectious Cycle, Infect. Immunol., 2004, vol. 72, pp. 5938–5946.
Parkhill, J., Sebaihia, M., Preston, A., et al., Comparative Analysis of the Genome Sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica, Nat. Genet., 2003, vol. 35, pp. 32–40.
Kirillov, M.Yu., Nechaeva, E.V., and Karataev, G.I., A Mobile Genetic Element on a Bordetella pertussis Chromosome Stimulates Cointegrate Formation in Escherichia coli Strains, Rus. J. Genet., 1997, vol. 33, no. 12, pp. 1384–1390.
Nechaeva, E.V., Volgina, T.I., Kirillov, M.Yu., et al., Mobile Element RSBR1 of Bordetella pertussis, Mol. Genet. Microbiol. Virusol., 1999, no. 12, pp. 22–27.
Nechaeva, E.V., Sivov, I.G., and Karataev, G.I., Transposition and Inheritance of a Tn Element of Bordetella in Escherichia coli K12 Cells, Mol. Genet., Mikrobiol. Virusol., 2000, no. 1, pp. 20–23.
Kirillov, M.Y., Shumakov, Y., Nechaeva, E.V., et al., Repeated Sequences Isolated from Bordetella pertussis Induce DNA Rearrangements and Deletions at High Frequency, Gene, 1995, vol. 166, pp. 111–116.
Sivov, I.G., Bol’shakova, T.N., and Karataev, G.I., Integration and Intramolecular Transposition of the TnBP3 Bordetella pertussis Transposon in the Escherichia coli K-12 Cells Mutant for the Phosphoenolpyruvate-Dependent Phosphotransferase System Hpr Protein, Rus. J. Genet., 2001, vol. 37, no. 7, pp. 741–747.
Stibitz, S., IS481 and IS1002 of Bordetella pertussis Create a 6-Base Pair Duplication upon Insertion at Consensus Target Site, J. Bacteriol., 1998, vol. 180, pp. 4963–4966.
Sinyashina, L.N., Vorontsov, V.V., Semin, E.G., et al., bvg-Negative Regulation of Transposition of Repetitive Sequences in Bordetella pertussis Cells, Genetika, 2005, vol. 41, no. 12, pp. 1608–1616.
Author information
Authors and Affiliations
Additional information
Original Russian Text © L.N. Sinyashina, G.I. Karataev, 2006, published in Genetika, 2006, Vol. 42, No. 3, pp. 339–348.
Rights and permissions
About this article
Cite this article
Sinyashina, L.N., Karataev, G.I. Molecular evidence for the lysogenic state of microorganisms belonging to the genus Bordetella and characterization of Bordetella parapertussis temperate bacteriophage 662-2 . Russ J Genet 42, 259–267 (2006). https://doi.org/10.1134/S1022795406030057
Received:
Issue Date:
DOI: https://doi.org/10.1134/S1022795406030057