Abstract
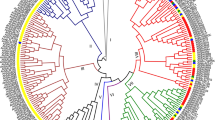
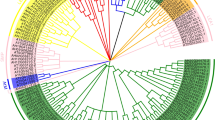
Peanut (Arachis hypogaea L.), as a source of oil and protein, is the second-most important grain legume cultivated in the world. Peanut is a relatively drought-tolerant crop; however, the molecular biology research of peanut is far behind other crops; the molecular mechanisms of its stress resistance are poorly understood. DELLA proteins, negative regulators of gibberellin signaling pathway in plants, promote survival of plants in adverse environments. In this study, four DELLA homologue genes were isolated using peanut transcriptome sequences. Molecular phylogenetic analysis revealed that these four AhDELLAs fall into three distinct groups: AhDELLA1 and AhDELLA2 clustered into two distinct groups, while AhDELLA3 and AhDELLA4 were in one group, which was separated from other DELLAs. qRT-PCR results showed that these four DELLA genes were expressed differentially in various peanut tissues. The AhDELLA1 and AhDELLA2 genes were expressed ubiquitously in different tissues. AhDELLA3 and AhDELLA4 showed much higher expression level in flowers and seeds as compared with other organs. The expression of four AhDELLA genes was temporally induced by PEG 6000 treatment. The AhDELLA3 and AhDELLA4 transcripts were significantly induced by NaCl treatment, while the expression of AhDELLA1 and AhDELLA2 did not change much under salt stress. The possible role of DELLA proteins in peanut development and responses to abiotic stresses is discussed.
Similar content being viewed by others
References
Yamaguchi, S., Gibberellin metabolism and its regulation, Annu. Rev. Plant Biol., 2008, vol. 59, pp. 225–251.
Harberd, N.P., Belfield, E., and Yasumura, Y., The angiosperm gibberellin-GID1-DELLA growth regulatory mechanism: How an “inhibitor of an inhibitor” enables flexible response to fluctuating environments, Plant Cell, 2009, vol. 21, pp. 1328–1339.
Sun, T.P., The molecular mechanism and evolution of the GA-GID1-DELLA signaling module in plants, Curr. Biol., 2011, vol. 21, pp. 338–345.
Ueguchi-Tanaka, M., Nakajima, M., Motoyuki, A., and Matsuoka, M., Gibberellin receptor and its role in gibberellin signaling in plants, Annu. Rev. Plant Biol., 2007, vol. 58, pp. 183–198.
Bolle, C., The role of GRAS proteins in plant signal transduction and development, Planta, 2004, vol. 218, pp. 683–692.
Boss, P.K. and Thomas, M.R., Association of dwarfism and floral induction with a grape ‘Green Revolution’ mutation, Nature, 2002, vol. 416, pp. 847–850.
Bassel, G.W., Zielinska, E., Mullen, R.T., and Bewley, J.D., Down-regulation of DELLA genes is not essential for germination of tomato, soybean, and Arabidopsis seeds, Plant Physiol., 2004, vol. 136, pp. 2782–2789.
Vandenbussche, F., Fierro, A.C., Wiedemann, G., Reski, R., and van der Straeten, D., Evolutionary conservation of plant gibberellin signalling pathway components, BMC Plant Biol., 2007, vol. 29, pp. 7–65.
Silverstone, A.L., Ciampaglio, C.N., and Sun, T.P., The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway, Plant Cell, 1998, vol. 10, pp. 155–169.
Dill, A., Jung, H.S., and Sun, T.P., The DELLA motif is essential for gibberellin-induced degradation of RGA, Proc. Natl. Acad. Sci. USA, 2001, vol. 98, pp. 14162–14167.
Nakajima, M., Shimada, A., Takashi, Y., Kim, Y.C., Park, S.H., Ueguchi-Tanaka, M., Suzuki, H., Katoh, E., Iuchi, S., Kobayashi, M., Maeda, T., Matsuoka, M., and Yamaguchi, I., Identification and characterization of Arabidopsis gibberellin receptors, Plant J., 2006, vol. 46, pp. 880–889.
Fu, X., Richards, D.E., Fleck, B., Xie, D., Burton, N., and Harberd, N.P., The Arabidopsis mutant sleepy1gar2-1 protein promotes plant growth by increasing the affinity of the SCFSLY1 E3 ubiquitin ligase for DELLA protein substrates, Plant Cell, 2004, vol. 16, pp. 1406–1418.
Wen, C.K. and Chang, C., Arabidopsis RGL1 encodes a negative regulator of gibberellin responses, Plant Cell, 2002, vol. 14, pp. 87–100.
Dill, A. and Sun, T.P., Synergistic de-repression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana, Genetics, 2001, vol. 159, pp. 777–785.
Cheng, H., Qin, L., Lee, S., Fu, X., Richards, D.E., Cao, D., Luo, D., Harberd, N.P., and Peng, J., Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function, Development, 2004, vol. 131, pp. 1055–1064.
Tyler, L., Thomas, S.G., Hu, J., Dill, A., Alonso, J.M., Ecker, J.R., and Sun, T.P., DELLA proteins and gibberellin-regulated seed germination and floral development in Arabidopsis, Plant Physiol., 2004, vol. 135, pp. 1008–1019.
Lee, S., Cheng, H., King, K.E., Wang, W., He, Y., Hussain, A., Lo, J., Harberd, N.P., and Peng, J., Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibitions, Genes Dev., 2002, vol. 16, pp. 646–658.
Achard, P., Cheng, H., de Grauwe, L., Decat, J., Schoutteten, H., Moritz, T., van der Straeten, D., Peng, J., and Harberd, N.P., Integration of plant responses to environmentally activated phytohormonal signals, Science, 2006, vol. 311, pp. 91–94.
Achard, P., Renou, J.P., Berthomé, R., Harberd, N.P., and Genschik, P., Plant DELLAs restrain growth and promote survival of adversity by reducing the levels of reactive oxygen species, Curr. Biol., 2008, vol. 18, pp. 656–660.
Claeys, H., Skirycz, A., Maleux, K., and Inzé, D., DELLA signaling mediates stress-induced cell differentiation in Arabidopsis leaves through modulation of anaphase-promoting complex/cyclosome activity, Plant Physiol., 2012, vol. 159, pp. 739–747.
Chen, F., Li, Q., Sun, L., and He, Z., The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress, DNA Res., 2006, vol. 13, pp. 53–63.
Huang, C., Picimbon, J.F., Li, H.Q., Li, Z., Liu, Q., and Liu, W., An efficient method for total RNA extraction from peanut seeds, Russ. J. Plant Physiol., 2012, vol. 59, pp. 129–133.
Han, X., Chuanzhi, Z., Lei, H., Aiqin, L., Shuzhen, Z., Yuping, B., Jing, A., Yanxiu, Z., Shubo, W., and Xingjun, W., Transcriptome profiling of peanut gynophores revealed global reprogramming of gene expression during early pod development in darkness, BMC Genomics, 2013, vol. 14, p. 517.
Livak, K.J. and Schmittgen, T.D., Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method, Methods, 2001, vol. 25, pp. 402–408.
Gallego-Bartlomé, J., Minguet, E.G., Marín, J.A., Prat, S., Blázquez, M.A., and Alabadí, D., Transcriptional diversification and functional conservation between DELLA proteins in Arabidopsis, Mol. Biol. Evol., 2010, vol. 27, pp. 1247–1556.
Achard, P., Gong, F., Cheminant, S., Alioua, M., Hedden, P., and Genschik, P., The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism, Plant Cell, 2008, vol. 20, pp. 2117–2129.
Author information
Authors and Affiliations
Corresponding author
Additional information
This text was submitted by the authors in English.
These authors contributed equally to the work.
Rights and permissions
About this article
Cite this article
An, J., Hou, L., Li, C. et al. Cloning and expression analysis of four DELLA genes in peanut. Russ J Plant Physiol 62, 116–126 (2015). https://doi.org/10.1134/S1021443715010021
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1021443715010021