Abstract
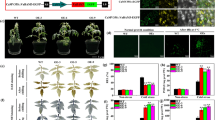
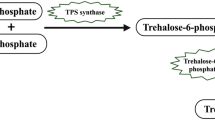
The faldh gene encodes the Brevibacillus brevis glutathione-dependent formaldehyde dehydrogenase (FALDH), an enzyme involved in formaldehyde metabolism. In the present work, we have investigated the physiological characteristics of transgenic faldh tobacco under formaldehyde stress. Overexpression of B. brevis FALDH confers tobacco tolerance to high HCHO concentrations. The transgenic tobacco lines had the higher biomass, produced the higher content of total proteins and soluble sugars, the lower levels of MDA, protein carbonyl (PC), and H2O2 as compared with the wild-type tobacco under HCHO stress. The contents of chlorophyll (Chl), including Chl a, Chl b, and the ratio of Chl a/b, and the content of anthocyanidin in transgenic plants under HCHO stress were also higher than that in wild-type tobacco. These results show that high HCHO tolerance and changes of physiological characteristics related to stress tolerance were due to the overexpressing of FALDH in tobacco.
Similar content being viewed by others
Abbreviations
- FALDH:
-
glutathione-dependent formaldehyde dehydrogenase
- C1:
-
one carbon
- Chl:
-
chlorophyll
- DNPH:
-
dinitrophenylhydrazine
- Km:
-
kanamycin
- PC:
-
protein carbonyl
- TBA:
-
thiobarbituric acid
- WT:
-
wild type
References
Blunden, G., Carpenter, B.G., Adrian-Romero, M., Yang, M.H., and Tyihak, E., Formaldehyde in the plant kingdom, Acta Biol. Hung., 1998, vol. 49, pp. 239–246.
Trezl, L., Hullan, L., Szarvas, T., Csiba, A., and Szende, B., Determination of endogenous formaldehyde in plants (fruits) bound to L-arginine and its relation to the folate cycle, photosynthesis and apoptosis, Acta Biol. Hung., 1998, vol. 49, pp. 253–263.
Giese, M., Bauer-Doranth, U., Langebartels, C., and Sandermann, H., Jr., Detoxification of formaldehyde by the spider plant (Chlorophytum comosum L.) and by soybean (Glycine max L.) cell-suspension cultures, Plant Physiol., 1994, vol. 104, pp. 1301–1309.
Schmitz, H., Hilgers, U., and Weidner, M., Assimilation and metabolism of formaldehyde by leaves appear unlikely to be of value for indoor air purification, New Phytol., 2000, vol. 147, pp. 307–315.
Achkor, H., Diaz, M., Fernandez, M.R., Biosca, J.A., Pares, X., and Martinez, M.C., Enhanced formaldehyde detoxification by overexpression of glutathionedependent formaldehyde dehydrogenase from Arabidopsis, Plant Physiol., 2003, vol. 132, pp. 2248–2255.
Li, R., Moore, M., Bonham-Smith, P.C., and King, J., Overexpression of formate dehydrogenase in Arabidopsis thaliana resulted in plants tolerant to high concentrations of formate, J. Plant Physiol., 2002, vol. 159, pp. 1069–1076.
Chen, L.M., Yurimoto, H., Li, K.Z., Orita, I., Akita, M., Kato, N., Sakai, Y., and Izui, K., Assimilation of formaldehyde in transgenic plants due to the introduction of the bacterial ribulose monophosphate pathway genes, BoiSci. Biotechnol. Biochem., 2010, vol. 74, pp. 627–635.
Song, Z.B., Orita, I., Yin, F., Yurimoto, H., Kato, N., Sakai, Y., Izui, K., Li, K.Z., and Chen, L.M., Overexpression of an HPS/PHI fusion enzyme from Mycobacterium gastri in chloroplasts of geranium enhances its ability to assimilate and phytoremediate formaldehyde, Biotechnol. Lett., 2010, vol. 32, pp. 1541–1548.
Mutters, R.G., Madore, M., and Bytnerowicz, M., Formaldehyde exposure affects growth and metabolism of common bean, Air Waste, 1993, vol. 43, pp. 113–116.
Nian, H., Meng, Q., Zhang, W., and Chen, L., Overexpression of the formaldehyde dehydrogenase gene from Brevibacillus brevis to enhance formaldehyde tolerance and detoxification of tobacco, Appl. Biochem. Biotechnol., 2013, vol. 169, pp. 170–180.
Bradford, M.M., A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding, Anal. Biochem., 1976, vol. 72, pp. 248–254.
Seifer, S., Dayton, S., Novic, B., and Muntwyler, E., Estimation of glycogen with anthrone reagent, Arch. Biochem., 1950, vol. 25, pp. 191–200.
Baccouch, S., Chaoui, A., and Ferjani, E.E., Nickelinduced oxidative damage and antioxidant responses in Zea mays shoots, Plant Physiol. Biochem., 1998, vol. 36, pp. 689–694.
Gurel, A., Coskun, O., Armutcu, F., Kanter, M., and Ozen, O.A., Vitamin E against oxidative damage caused by formaldehyde in frontal cortex and hippocampus: biochemical and histological studies, J. Chem. Neuroanat., 2005, vol. 29, pp. 173–178.
Gay, C.A. and Gebicki, J.M., Measurement of protein and lipid hydroperoxides in biological systems by the ferric-xylenol orange method, Anal. Biochem., 2003, vol. 315, pp. 29–35.
Strain, H.H., Cope, B.T., and Svec, W.A., Analytical procedures for the isolation, identification, estimation and investigation of the chlorophylls, Methods Enzymol., 1971, vol. 23, pp. 452–476.
Madhusudan, R. and Ravisankar, G.A., Gradients of anthocyanin in cell aggregates of Daucus carota in suspension cultures, Biotechnol. Lett., 1996, vol. 18, pp. 1253–1256.
Wang, S.S., Song, Z.B., Sun, Z., Zhang, J., Mei, Y., Nian, H.J., Li, K.Z., and Chen, L.M., The effects of formaldehyde stress on the physiological characteristics and gene expression associated with photosynthesis in Arabidopsis thaliana, Plant Mol. Biol. Rep., 2012, vol. 30, pp. 1291–1302.
Chen, Q., Zhang, X.D., Wang, S.S., Wang, Q.F., Wang, G.Q., Nian, H.J., Li, K.Z., Yu, Y.X., and Chen, L.M., Transcriptional and physiological changes of alfalfa in response to aluminum stress, J. Agric. Sci., 2011, vol. 149, pp. 737–751.
Nian, H., Wang, G., and Chen, L., Physiological and transcriptional analysis of the effects of aluminum stress on Cryptococcus humicola, World J. Microbiol. Biotechnol., 2012, vol. 28, pp. 2319–2329.
Author information
Authors and Affiliations
Corresponding author
Additional information
This text was submitted by the authors in English.
These authors contributed equally to this work.
Rights and permissions
About this article
Cite this article
Nian, H.J., Meng, Q.C., Cheng, Q. et al. The effects of overexpression of formaldehyde dehydrogenase gene from Brevibacillus brevis on the physiological characteristics of tobacco under formaldehyde stress. Russ J Plant Physiol 60, 764–769 (2013). https://doi.org/10.1134/S1021443713060083
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1021443713060083