Abstract
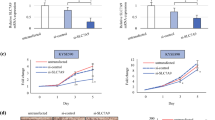
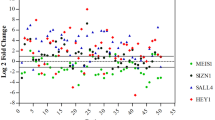
The investigation of molecular mechanisms contributing to cancer progression is the burning problem of current research. Considerable attention has been focused on the study of gene expression in cancer cells. Sphingomyelin synthase 1 gene (SGMS1) is one of the genes, the expression of which can be altered in cancer. SMS1 enzyme encoded by this gene catalyzes synthesis of sphingomyelin and diacylglycerol from phosphatidylcholine and ceramide. SMS1 may maintain the balance between cell death and survival by regulating the formation of the proaptotic mediator ceramide and anti-apoptotic mediator diacylglycerol. In addition, changes in the sphingomyelin level and sphingomyelin synthase activity have been observed in cancers of many tissues. However, the peculiarities of SGMS1 gene transcription have been insufficiently explored. In this work, the expression of transcripts of SGMS1 has been investigated by the method of real-time PCR in matched pairs of samples of human lung and esophagus cancer and adjacent tissues without pathology. A significant decrease in SMS1 transcript expression has been found in the samples of human lung cancer. At the same time, in the samples of human esophagus cancer and the adjacent tissues, the expression of SMS1 transcripts varies insignificantly, i.e., it is increased in seven and decreased in five of the fifteen samples. The obtained results indicate that SGMS1 gene is expressed differently in cancers of different genesis.
Similar content being viewed by others
References
Ferlay J., Steliarova-Foucher E., Lortet-Tieulent J., Rosso S., Coebergh J.W., Comber H., Forman D., Bray F. 2013. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries in 2012. Eur. J. Cancer. 49, 1374–1403.
Demidyuk I.V., Shubin A.V., Gasanov E.V., Kurinov A.M., Demkin V.V., Vinogradova T.V., Zinovyeva M.V., Sass A.V., Zborovskaya I.B., Kostrov S.V. 2013. Alterations in gene expression of proprotein convertases in human lung cancer have a limited number of scenarios. PLoS ONE. 8, e55752.
Zinovyeva M.V., Monastyrskaya G.S., Kopantzev E.P., Vinogradova T.V., Kostina M.B., Sass A.V., Filyukova O.B., Uspenskaya N.Y., Sukhikh G.T., Sverdlov E.D. 2010. Identification of some human genes oppositely regulated during esophageal squamous cell carcinoma formation and human embryonic esophagus development. Dis. Esophagus. 23, 260–270.
Kopantzev E.P., Monastyrskaya G.S., Vinogradova T.V., Zinovyeva M.V., Kostina M.B., Filyukova O.B., Tonevitsky A.G., Sukhikh G.T., Sverdlov E.D. 2008. Differences in gene expression levels between early and later stages of human lung development are opposite to those between normal lung tissue and non-small lung cell carcinoma. Lung Cancer. 62, 23–34.
Korobko I.V., Zinov’eva M.V., Kopantsev E.P., Allakhverdiev A.K., Zborovskaya I.B., Sverdlov E.D. 2007. Expression of C-MET and HGF in non-small cell lung carcinomas. Mol. Genet. Microbiol. Virol. (Moscow). 22, 59–63.
Dergunova L.V., Raevskaya N.M., Voloshenyuk E.L., Limborskaya S.A. 2007. Characteristics of expression of the genes of EGR1, neurotrophins, and their receptors in normal human lung tissues and lung cancers. Mol. Genet. Microbiol. Virol. (Moscow). 22, 68–74.
Vladychenskaya I.P., Dergunova L.V., Limborska S.A. 2002. In vitro and in silico analysis of the predicted human MOB gene encoding a phylogenetically conserved transmembrane protein. Biomol. Eng. 18, 263–268.
Vladychenskaya I.P., Dergunova L.V., Dmitrieva V.G., Limborska S.A. 2004. Human gene MOB: Structure specification and aspects of transcriptional activity. Gene. 338, 257–265.
Rozhkova A.V., Dmitrieva V.G., Zhapparova O.N., Sudarkina O.Y., Nadezhdina E.S., Limborska S.A., Dergunova L.V. 2011. Human sphingomyelin synthase 1 gene (SMS1): Organization, multiple mRNA splice variants and expression in adult tissues. Gene. 481, 65–75.
Segui B., Andrieu-Abadie N., Jaffrezou J.P., Benoist H., Levade T. 2006. Sphingolipids as modulators of cancer cell death: Potential therapeutic targets. Biochim. Biophys. Acta. 1758, 2104–2120.
Claus R.A., Dorer M.J., Bunck A.C., Deigner H.P. 2009. Inhibition of sphingomyelin hydrolysis: Targeting the lipid mediator ceramide as a key regulator of cellular fate. Curr. Med. Chem. 16, 1978–2000.
Hannun Y.A. 1994. The sphingomyelin cycle and the second messenger function of ceramide. J. Biol. Chem. 269, 3125–3128.
Kandyba A.G., Kobliakov V.A., Somova O.G., Dyatlovitskaya E.V. 2004. Change in contents of biologically active sphingolipids modulating cell growth and survival in hepatoma 27 compared to rat liver. Biochemistry (Moscow). 69, 497–500.
Narayan P., Dahiya R. 1991. Alterations in sphingomyelin and fatty acids in human benign prostatic hyperplasia and prostatic cancer. Biomed. Biochim. Acta. 50, 1099–1108.
Albi E., La Porta C.A., Cataldi S., Magni M.V. 2005. Nuclear sphingomyelin-synthase and protein kinase C delta in melanoma cells. Arch. Biochem. Biophys. 438, 156–161.
Merchant T.E., Meneses P., Gierke L.W., Den Otter W., Glonek T. 1991. 31P magnetic resonance phospholipid profiles of neoplastic human breast tissues. Br. J. Cancer. 63, 693–698.
Merchant T.E., De Graaf P.W., Minsky B.D., Obertop H., Glonek T. 1993. Esophageal cancer phospholipid characterization by 31P NMR. NMR Biomed. 6, 187–193.
Chomczynski P., Mackey K. 1995. Short technical reports. Modification of the TRI reagent procedure for isolation of RNA from polysaccharide- and proteoglycan-rich sources. Biotechniques. 19, 942–945.
Zhu Y.Y., Machleder E.M., Chenchik A., Li R., Siebert P.D. 2001. Reverse transcriptase template switching: A SMART approach for full-length cDNA library construction. Biotechniques. 30, 892–897.
Rubie C., Kempf K., Hans J., Su T., Tilton B., Georg T., Brittner B., Ludwig B., Schilling M. 2005. Housekeeping gene variability in normal and cancerous colorectal, pancreatic, esophageal, gastric and hepatic tissues. Mol. Cell Probes. 19, 101–109.
Krasnov G.S., Oparina N.Yu., Dmitriev A.A., Kudryavtseva A.V., Anedchenko E.A., Kondrat’eva T.T., Zabarovsky E.R., Senchenko V.N. 2011. RPN1, a new reference gene for quantitative data normalization in lung and kidney cancer. Mol. Biol. (Moscow). 45, 211–220.
Oparina N.Yu., Snezhkina A.V., Sadritdinova A.F., Veselovskii V.A., Dmitriev A.A., Senchenko V.N., Mel’nikova N.V., Speranskaya A.S., Darii M.V., Stepanov O.A., Barkhatov I.M., Kudryavtseva A.V. 2013. Differential expression of genes that encode glycolysis enzymes in kidney and lung cancer in humans. Russ. J. Genet. 49, 707–716.
Pfaffl M.W., Horgan G.W., Dempfle L. 2002. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 30, e36.
Morad S.A.F., Cabot M.C. 2013. Ceramide-orchestrated signalling in cancer cells. Nature Rev. Cancer. 13, 51–65.
Barceló-Coblijn G., Martin M.L., De Almeida R.F.M., Noguera-Salvà M.A., Marcilla-Etxenike A., Guardiola-Serrano F., Lüth A., Kleuser B., Halver J.E., Escribá P.V. 2011. Sphingomyelin and sphingomyelin synthase (SMS) in the malignant transformation of glioma cells and in 2-hydroxyoleic acid therapy. Proc. Natl. Acad. Sci. U. S. A. 108, 19569–19574.
van Blitterswijk W.J., Klarenbeek J.B., van der Luit A.H., Alderliesten M.C., van Lummel M., Verheij M. 2010. Fas/CD95 down-regulation in lymphoma cells through acquired alkyllysophospholipid resistance: Partial role of associated sphingomyelin deficiency. Biochem. J. 425, 225–234.
Meng A., Luberto C., Meier P., Bai A., Yang X., Hannun Y.A., Zhou D. 2004. Sphingomyelin synthase as a potential target for D609-induced apoptosis in U937 human monocytic leukemia cells. Exp. Cell Res. 292, 385–392.
Itoh M., Kitano T., Watanabe M., Kondo T., Yabu T., Taguchi Y., Iwai K., Tashima M., Uchiyama T., Okazaki T. 2003. Possible role of ceramide as an indicator of chemoresistance: Decrease of the ceramide content via activation of glucosylceramide synthase and sphingomyelin synthase in chemoresistant leukemia. Clin. Cancer Res. 9, 415–423.
Abrams S.I. 2005. Positive and negative consequences of Fas/Fas ligand interactions in the antitumor response. Front. Biosci. 10, 809–821.
Listopad J.J., Kammertoens T., Anders K., Silkenstedt B., Willimsky G., Schmidt K., Kuehl A.A., Loddenkemper C., Blankenstein T. 2013. Fas expression by tumor stroma is required for cancer eradication. Proc. Natl. Acad. Sci. U. S. A. 110, 2276–2281.
Hueber A.O., Bernard A.M., Herincs Z., Couzinet A., He H.T. 2002. An essential role for membrane rafts in the initiation of Fas/CD95-triggered cell death in mouse thymocytes. EMBO Rep. 3, 190–196.
Miyaji M., Jin Z.X., Yamaoka S., et al. Amakawa R., Fukuhara S., Sato S.B., Kobayashi T., Domae N., Mimori T., Bloom E.T., Okazaki T., Umehara H. 2005. Role of membrane sphingomyelin and ceramide in platform formation for Fas-mediated apoptosis. J. Exp. Med. 202, 249–259.
Li Z., Hailemariam T.K., Zhou H., Li Y., Duckworth D.C., Peake D.A., Zhang Y., Kuo M.S., Cao G., Jiang X.C. 2007. Inhibition of sphingomyelin synthase (SMS) affects intracellular sphingomyelin accumulation and plasma membrane lipid organization. Biochim. Biophys. Acta. 1771, 1186–1194.
Roberts N.J., Zhou S., Diaz L.A., Jr., Holdhoff M. 2011. Systemic use of tumor necrosis factor alpha as an anticancer agent. Oncotarget. 2, 739–751.
Wang S., El-Deiry W.S. 2003. TRAIL and apoptosis induction by TNF-family death receptors. Oncogene. 22, 8628–8633.
Hoshi H., Sawada T., Uchida M., Iijima H., Kimura K., Hirakawa K., Wanibuchi H. 2013. MUC5AC protects pancreatic cancer cells from TRAIL-induced death pathways. Int. J. Oncol. 42, 887–893.
Walczak H. 2013. Death receptor-ligand systems in cancer, cell death, and inflammation. Cold Spring Harbor Perspect. Biol. 5, a008698.
Song J.H., Tse M.C., Bellail A., Phuphanich S., Khuri F., Kneteman N.M., Hao C. 2007. Lipid rafts and nonrafts mediate tumor necrosis factor related apoptosis-inducing ligand induced apoptotic and nonapoptotic signals in non small cell lung carcinoma cells. Cancer Res. 67, 6946–6955.
Ouyang W., Yang C., Zhang S., Liu Y., Yang B., Zhang J., Zhou F., Zhou Y., Xie C. 2013. Absence of death receptor translocation into lipid rafts in acquired TRAIL-resistant NSCLC cells. Int. J. Oncol. 42, 699–711.
Ding T., Li Z., Hailemariam T., Mukherjee S., Maxfield F.R., Wu M.P., Jiang X.C. 2008. SMS overexpression and knockdown: Impact on cellular sphingomyelin and diacylglycerol metabolism, and cell apoptosis. J. Lipid Res. 49, 376–385.
Igney F.H., Krammer P.H. 2002. Immune escape of tumors: Apoptosis resistance and tumor counterattack. J. Leukoc. Biol. 71, 907–920.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © A.V. Rozhkova, M.V. Zinovyeva, A.V. Sass, I.B. Zborovskaya, S.A. Limborska, L.V. Dergunova, 2014, published in Molekulyarnaya Biologiya, 2014, Vol. 48, No. 3, pp. 395–402.
Rights and permissions
About this article
Cite this article
Rozhkova, A.V., Zinovyeva, M.V., Sass, A.V. et al. Expression of sphingomyelin synthase 1 (SGMS1) gene varies in human lung and esophagus cancer. Mol Biol 48, 340–346 (2014). https://doi.org/10.1134/S0026893314030170
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893314030170