Abstract
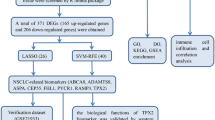
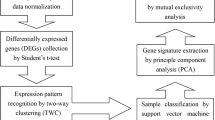
Chronic obstructive pulmonary disease (COPD) is a complex human disease with a high mortality rate. So far, the studies of COPD have not been well organized despite the well-documented role of cigarette smoking in the genesis of COPD. In the recent years, microarray analyses have helped to identify some potential disease related genes. However, the low reproducibility of many published gene signatures has been criticized. It therefore suggested that incorporation of network or pathway information into prognostic biomarker discovery might improve the prediction performance. In this analysis, we combined protein-protein interactions (PPI) information with the support vector machine (SVM) method to identify potential COPD-related genes that would allow one to distinguish accurately severe emphysema from non-/mildly emphysematous lung tissue. We identified 8 COPD-related feature genes. When compared with another SVM method which did not use the prior PPI information, the prediction accuracy was significantly enhanced (AUC was increased from 0.513 to 0.909). On the base of results obtained one can suppose that incorporating network of prior knowledge into gene selection methods significantly improves classification accuracy. Consequently, the gene expression profiles from human emphysematous lung tissue may provide insight into the pathogenesis, and a good classification prediction algorithm based on prior biological knowledge can further strengthen this performance.
Similar content being viewed by others
References
Sethi J.M., Rochester C.L. 2000. Smoking and chronic obstructive pulmonary disease. Clin. Chest. Med. 21, 67–86.
Snider G.L. 1989. Chronic obstructive pulmonary disease: Risk factors, pathophysiology and pathogenesis. Annu. Rev. Med. 40, 411–429.
Acquaah-Mensah G.K., Malhotra D., Vulimiri M., McDermott J.E., Biswal S. 2012. Suppressed expression of T-box transcription factors is involved in senescence in chronic obstructive pulmonary disease. PLoS Comput. Biol. 8, e1002597.
Salvi S.S., Barnes P.J. 2009. Chronic obstructive pulmonary disease in nonsmokers. Lancet. 374, 733–743.
Buist A.S., McBurnie M.A., Vollmer W.M., Gillespie S., Burney P., Mannino D.M., Menezes A.M., Sullivan S.D., Lee T.A., Weiss K.B. 2007. International variation in the prevalence of COPD (the BOLD study): A population-based prevalence study. Lancet. 370, 741–750.
Vibhuti A., Arif E., Deepak D., Singh B., Qadar Pasha M.A. 2007. Genetic polymorphisms of GSTP1 and mEPHX correlate with oxidative stress markers and lung function in COPD. Biochem. Biophys. Res. Commun. 359, 136–142.
Palmer L.J., Celedón J.C., Chapman H.A., Speizer F.E., Weiss S.T., Silverman E.K. 2003. Genome-wide linkage analysis of bronchodilator responsiveness and postbronchodilator spirometric phenotypes in chronic obstructive pulmonary disease. Hum. Mol. Genet. 12, 1199–1210.
Silverman E.K., Palmer L.J., Mosley J.D., Barth M., Senter J.M., Brown A., Drazen J.M., Kwiatkowski D.J., Chapman H.A., Campbell E.J., et al. 2002. Genomewide linkage analysis of quantitative spirometric phenotypes in severe early-onset chronic obstructive pulmonary disease. Am. J. Hum. Genet. 70, 1229–1239.
Furey T.S., Cristianini N., Duffy N., Bednarski D.W., Schummer M., Haussler D. 2000. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics. 16, 906–914.
Aoshiba K., Nagai A. 2009. Senescence hypothesis for the pathogenetic mechanism of chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 6, 596–601.
Cun Y., Fröhlich H. 2012. Prognostic gene signatures for patient stratification in breast cancer: Accuracy, stability and interpretability of gene selection approaches using prior knowledge on protein-protein interactions. BMC Bioinform. 13(69), doi 10.1186/1471-2105-11131169
Rapaport F., Zinovyev A., Dutreix M., Barillot E., Vert J.P. 2007. Classification of microarray data using gene networks. BMC Bioinform. 8, 35.
Dudoit S., Fridlyand J., Speed T.P. 2002. Comparison of discrimination methods for the classication of tumors using gene expression data. J. Am. Stat. Assoc. 97, 77–87.
Spira A., Beane J., Pinto-Plata V., Kadar A., Liu G., Shah V., Celli B., Brody J.S. 2004. Gene expression profiling of human lung tissue from smokers with severe emphysema. Am. J. Respir. Cell Mol. Biol. 31, 601–610.
Zhang S. 2007. A comprehensive evaluation of SAM, the SAM R-package and a simple modification to improve its performance. BMC Bioinform. 8, 230.
Reiner A., Yekutieli D., Benjamini Y. 2003. Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics. 19, 368–375.
Saviozzi S., Cordero F., Iacono M.L., Novello S., Giorgio S.V., Calogero R.A. 2006. Selection of suitable reference genes for accurate normalization of gene expression profile studies in non-small cell lung cancer. BMC Cancer. 6, 200.
Yang J., Feng X., Fu Z., Yuan C., Hong Y., Shi Y., Zhang M., Liu J., Li H., Lu K., et al. 2012. Ultrastructural observation and gene expression profiling of Schistosoma japonicum derived from two natural reservoir hosts, water buffalo and yellow cattle. PLoS ONE. 7, e47660.
Guyon I., Weston J., Barnhill S. 2002. Gene selection for cancer classification using support vector machines. Machine Learning. 46, 389–422.
Johannes M., Brase J.C., Fröhlich H., Gade S., Gehrmann M., Fälth M., Sültmann H., Beißbarth T. 2010. Integration of pathway knowledge into a reweighted recursive feature elimination approach for risk stratification of cancer patients. Bioinformatics. 26, 2136–2144.
Crammer K., Singer Y. 2001. On the algorithmic implementation of multiclass Kernel-based vector machines. J. Machine Learning Res. 2, 265–292.
Brin S., Page L. 1998. The anatomy of a large-scale hypertextual web search engine. Comput. Netw. ISDN Syst. 30, 107–117.
Prasad T.S.K., Goel R., Kandasamy K., Keerthikumar S., Kumar S., Mathivanan S., Telikicherla D., Raju R., Shafreen B., Venugopal A., et al. 2009. Human protein reference database-2009 update. Nucleic Acids Res. 37, D767–D772.
Boulesteix A.L. 2009. Stability and aggregation of ranked gene lists. Brief Bioinform. 10, 556–568.
Baldi P., Long A.D. 2001. A Bayesian framework for the analysis of microarray expression data: Regularized t-test and statistical inferences of gene changes. Bioinformatics. 17, 509–519.
Opgen-Rhein R., Strimmer K. 2007. Accurate ranking of differentially expressed genes by a distribution-free shrinkage approach. Stat. Appl. Genet. Mol. Biol. 6, 1–20.
John G.H., Langley P. 1995. Estimating continuous distributions in Bayesian classifiers. Proc. Eleventh Conf. on Uncertainty in Artificial Intelligence, Montreal, August 18–20, 1995. San Francisco, CA: Morgan Kaufmann, pp. 338–345.
Gutin G., Yeo A., Zverovich A. 2002. Traveling salesman should not be greedy: Domination analysis of greedy-type heuristics for the TSP. Discr. Appl. Math. 117, 81–86.
Furey T.S. 2000. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics. 16, 906–914.
Pang H., Lin A., Holford M., Enerson B.E., Lu B., Lawton M.P., Floyd E., Zhao H. 2006. Pathway analysis using random forests classification and regression. Bioinformatics. 22, 2028–2036.
Neil M., Fenton N., Tailor M. 2005. Using Bayesian networks to model expected and unexpected operational losses. Risk Analysis. 25, 1539–1548.
Lu P., Hankel I.L., Hostager B.S., Swartzendruber J.A., Friedman A.D., Brenton J.L., Rothman P.B., Colgan J.D. 2011. The developmental regulator protein Gon4l associates with protein YY1, co-repressor Sin3a, and histone deacetylase 1 and mediates transcriptional repression. J. Biol. Chem. 286, 18311–18319.
Natanek S.A., Riddoch-Contreras J., Marsh G.S., Hopkinson N.S., Man W.D.C., Moxham J., Polkey M.I., Kemp P.R. 2011. Yin Yang 1 expression and localisation in quadriceps muscle in COPD. Arch. Bronconeumol. 47, 296–302.
Steiling K., Kadar A.Y., Bergerat A., Flanigon J., Sridhar S., Shah V., Ahmad Q.R., Brody J.S., Lenburg M.E., Steffen M., et al. 2009. Comparison of proteomic and transcriptomic profiles in the bronchial airway epithelium of current and never smokers. PLoS ONE. 4, e5043.
Anelli T., Massimo A., Alexandre M., Thomas S., Fabio T., Angela B., Roberto S. 2002. ERp44, a novel endoplasmic reticulum folding assistant of the thioredoxin family. EMBO J. 21, 835–844.
Asyali M.H., Colak D., Demirkaya O., Inan M.S. 2006. Gene expression profile classification: A review. Curr. Bioinform. 1, 55–73.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Molekulyarnaya Biologiya, 2014, Vol. 48, No. 2, pp. 333–343.
The article is published in the original.
Rights and permissions
About this article
Cite this article
Hua, L., Zhou, P. Combining protein-protein interactions information with support vector machine to identify chronic obstructive pulmonary disease related genes. Mol Biol 48, 287–296 (2014). https://doi.org/10.1134/S0026893314020101
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893314020101