Abstract
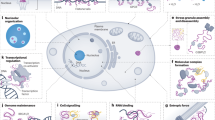
Until recently, the point of view that the unique tertiary structure is necessary for protein function has prevailed. However, recent data have demonstrated that many cell proteins do not possess such structure in isolation, although displaying a distinct function under physiological conditions. These proteins were named the naturally, or intrinsically, disordered proteins. The fraction of intrinsically disordered regions in such proteins may vary from several amino acid residues to a completely unordered sequence of several tens or even several hundreds of residues. The main distinction of these proteins from structured (globular) proteins is that they have no unique tertiary structure in isolation and acquire it only upon interaction with their partners. The conformation of these proteins in a complex is determined not only by their own amino acid sequence (as is typical of structured, or globular, proteins) but also by the interacting partner. This review discusses the structure-function relationships in structured and intrinsically disordered proteins. The intricateness of this problem and the possible ways to solve it are illustrated by the example of the EF1A elongation factor family.
Similar content being viewed by others
References
Dunker A.K., Lawson D.J., et al. 2001. Intrinsically disordered protein. J. Mol. Graphics Modelling. 19, 26–59.
Uversky V.N., Gillspie J.R., Fink A.L. 2000. Why are “natively unfolded” proteins unstructured under physiologic conditions? Proteins. 41, 415–427.
Jeffry C.J. 1999. Moonlighting proteins. Trends Biochem. Sci. 24, 8–11.
Wright P.E., Dyson H.J. 1999. Intrinsically unstructured proteins: Reassembling the protein structure-function paradigm. J. Mol. Biol. 293, 321–331.
Bogatyreva N.S., Finkelstein A.V., Galzitskaya O.V. 2006. Trend of amino acid compositions of different taxa. J. Bioinf. Comput. Biol. 4, 597–608.
Lemieux U.R., Spohr U. 1994. How Emil Fisher was led to the lock and key concept for enzyme specificity. Adv. Carbohydrate Chem. Biochem. 50, 1–20.
Mirsky A.E., Pauling L. 1936. On the structure of native, denaturated, and coagulated proteins. Proc. Natl. Acad. Sci. USA. 22, 439–447.
Edsall J.T. 1952. Some comments on proteins and protein structure. Proc. R. Soc. London B. 147, 97–113.
Anfinsen C.B. 1973. Principles that govern the folding of protein chains. Science. 181, 223–230.
Kendrew J., Dickerson R., Standberg B., Hart R., Davies D., Philips P., Shore V. 1960. Structure of myoglobin. Nature. 185, 422–428.
Perutz M., Rossmann M., Gullis A., Muirhead H., Will G., North A., 1960. Structure of haemoglobin. Nature. 185, 416–421.
Karush F. 1950. Heterogenity of the binding sites of bovine serum albumin. J. Am. Chem. Soc. 72, 2705–2713.
Koshland D.E., Jr. 1958. Application of a theory of enzyme specificity to protein synthesis. Proc. Natl. Acad. Sci. USA. 44, 98–104.
Koshland D.E., Jr. 1994. The key-lock theory and the induced fit theory. Angew. Chem. Int. Ed. Engl. 33, 2375–2378.
Blake C.C., Koenig D.F., et al. 1965. Structure of hen egg-white lysozyme: A three-dimensional Fourier synthesis at 2 Angstrom resolution. Nature. 206, 757–761.
McDonald R.C., Steitz T.A., Engelman D.M. 1979. Yeast hexokinase in solution exhibits a large conformational change upon binding glucose or glucose 6-phosphate. Biochemistry. 18, 338–342.
Kjeldgaard M., Nissen P., Thirup S., Nyborg J. 1993. The crystal structure of elongation factor EF-Tu from Thermus aquaticus in the GTP conformation. Structure. 1, 35–50.
Calmettes P., Roux B., Durand D., Desmadril M., Smith J.C. 1993. Configurational distribution of denaturated phosphoglycerate kinase. J. Mol. Biol. 231, 840–848.
Hinck A.P., Markus M.A., Huang S., Grzesiek S., Kustonovich I., Draper D.E., Torchia D.A. 1997. The RNA binding domain of ribosomal protein L11: Three-dimensional structure of the RNA-bound form of the protein and its interaction with 23S rRNA. J. Mol. Biol. 274, 101–113.
Kissinger C.R., Parge H.E., et al. 1995. Crystal structure of human calcineurin and the human FKBP12-FK506-cacineurin complex. Nature. 378, 641–644.
Wutrich K. 1998. The second millennium—in to the third millennium. Nature Struct. Biol. 5, 492–495.
Lee L., Stollar E., et al. 2001. Expression of the Oct-1 transcription factor and characterization of its interactions with the Bob1 coactivator. Biochemistry. 40, 6580–6588.
Tanford C., Kawahara K., Lapanije S. 1967. Proteins as a random coils: Intrinsic viscosity and sedimentation coefficients in concentrated guanidine hydrochloride. J. Am. Chem. Soc. 89, 729–734.
House-Pompeo K., Xu Y., Joh D., Speziale M. 1996. Conformational changes in the fibronectin binding MSCRAMMs are induced by ligand binding. J. Biol. Chem. 271, 1379–1384.
Dyson H.J., Wright P.E. 2003. Intrinsically unstructured proteins and their function. Mol. Cell Biol. 6, 197–205.
Demarest S.J., Martinez-Yamout M., et al. 2002. Mutual synergetic folding in recruitment of CBP/p300 by p160 nuclear receptor coactivators. Nature. 415, 549–553.
Tompa P. 2002. Intrinsically unstructed proteins. Trends Biochem. Sci. 27, 527–533.
Vucetic S., Brown J.C., Dunker A.K., Obradovic Z. 2003. Flavors of protein disorder. Proteins. 52, 573–584.
Oldfield C.J., Cheng Y., Cortese M.S., Brown C.J., Uverskiy V.N., Dunker A.K. 2005. Comparing and combining of mostly disordered proteins. Biochemistry. 44, 1989–2000.
Galzitskaya O.V., Garbuzynskiy A.S., Lobanov M.Yu. 2006. Prediction of natively unfolded regions of the protein chain. Mol. Biol. 40, 341–348.
Frankel A.D., Kim P.S. 1991. Molecular structure of transcription factors: Implication for gene regulation. Cell. 65, 717–719.
Negrutskii B.S., Deutsher M.P. 1991. Channeling of aminoacyl-tRNA for protein synthesis in vivo. Proc. Natl. Acad. Sci. USA. 88, 4991–4995.
Dunker A.K., Brown C.J., Lawson J.D., Lakoucheva L.M., Obradovic Z. 2002. Intrinsic disorder in cell-signalling and cancer-associated proteins. Biochemistry. 41, 6573–6582.
Rechsteiner M., Rogers S.W. 1996. PEST sequence and regulation for proteolysis. Trends Biochem. Sci. 21, 267–271.
Schuster S.C., Khan S., 1994. The bacterial flagellar motor. Ann. Rev. Biophys. Biomol. Struct. 23, 509–539.
Daughdrill G.W., Chadsey M.S., Karlinsey J.E., Hughes K.T., Dahlquist F.W. 1997. The C-terminal half of the anti-sigma factor, FlgM, becomes structured when bound to its target, sigma. Nature Struct. Biol. 4, 285–291.
Kostyukova A.S., Pyatibratov M.G., Filimonov V.V., Fedorov O.V. 1988. Flagellin parts acquiring a regular structure during polymerization are disposed on the molecule ends. FEBS Lett. 241, 141–144.
Piaxco K.W., Gros M. 1997. The importance of being unfolded. Nature. 386, 657–658.
Lakoucheva L.M., Brown C.J., Lawson D.J., Obradovic Z., Dunker A.K. 2002. Intrinsic disorder in cell-signaling and cancer-associated proteins. J. Mol. Biol. 323, 573–584.
Shiemaker B.A., Portman J.J., Wolynes G. 2000. Speeding molecular recognition by using the foldoing funell: The fly-casting mechanism. Proc. Natl. Acad. Sci. USA. 97, 8868–8873.
Shulz G.E. 1979. Nucleotide binding proteins. In: Molecular Mechanism of Biological Recognition. Ed. Balaban N. N.Y.: Elsevier/North Holland Biomedical Press, 74–94.
Kriwacki R.W., Hengst. L., Tennant L., Reed S.I., Wright P.E. 1996. Structural studies of p21 Waf1/Cip1/Sdi1 in the free and Cdk2-bound state: Conformatopnal disorder mediates binding diversity. Proc. Natl. Acad. Sci. USA, 93, 11504–11509.
Gunasekaran K., Tsai Ch.-J., Kumar S., Zanuy D., Nussinov R. 2003. Extended disordered proteins: Targeting function with less scaffold. Trends Biochem. Sci. 28, 81–85.
Subramanian A.R. 1983. Structure and functions of ribosomal protein S1. Prog. Nucl. Acids Res. Mol. Biol. 28, 101–142.
Laughrea M., Moore P.B. 1977. Physical properties of ribosomal protein S1 and its interaction with the 30S ribosomal subunit of Escherichia coli. J. Mol. Biol. 112, 399–421.
Berestovskaya N.V., Vasiliev V.D., Volkov A.A., Chetverin A.B. 1988. Electron microscopy study of Qβ replicase. FEBS Lett. 228, 263–267.
Wower I.K., Zwieb C.W., Guven S.A., Wower J. 2000. Binding and crosslinking of tmRNA to ribosomal protein S1, on and off the Escherichia coli ribosome. EMBO J. 19, 6612–6621.
Budkevich T.V., Timchenko A.A., Tiktopulo E.I., Negrutskii B.S., Shalak V.F., Petrushenko Z.M., Aksenov V.L., Willumeit R., Kohlbrecher J., Serdyuk I.N., El’skaya A.V. 2002. Extended conformation of mamalian translation elongation factor 1A in solution. Biochemistry. 41, 15342–15349.
Kjeldgard M., Nyborg J. 1992. Refined structure of elongation factor EF-Tu from Escherichia coli. J. Mol. Biol. 223, 721–742.
Song H., Parsons M.R., Rowsell S., Leonard G., Phillips S.E.V. 1999. Crystal structure of intact elongation factor EF-Tu from Escherichia coli in GDP conformation at 2.05 Å resolution. J. Mol. Biol. 285, 1245–1256.
Reshetnikova L.S., Reiser C.O., Schrimer N.K., Bertchold H., Storm R., Heilgenfeld R., Sprinzl M. 1991. Crystal of intact elongation factor Tu from Thermus thermophilus diffracting to high resolution. J. Mol. Biol. 221, 375–377.
Kjeldgard M., Nissen P., Thirup S., Nyborg J. 1993. The crystal structure of elongation factor EF-Tu from Thermus aquaticus in the GTP conformation. Structure. 1, 35–50.
Serdyuk I.N., Pavlov M.Yu., Rublevskaya I.N., Zaccai G., Libermann R. 1994. The triple isotopic substitution method in small angle neutron scattering: Application to the study of the ternary complex EF-Tu*GTP*aminoacyl-tRNA. Biophys. Chem. 53, 123–130.
Vitagliano L., Masullo M., Sica F., Zagari A., Bocchini V. 2001 The crystal structure of Sulfolobus solfataricus elongation factor 1α in the complex with GDP reveals novel features in nucleotide binding and exchange. EMBO J. 20, 5305–5311.
Bertold H., Reshetnikova L., Reiser C.O.A., Schirmer N.K., Sprinzl M., Hilgenfeld R. 1993. Crystal structure of active elongation factor Tu reveals major domain rearrangements. Nature. 365, 126–132.
Andersen G.R., Pedersen L., Valente L., Chatterjee I., Kinzy T.G., Kjeldgaard M., Nyborg J. 2000. Structural basis for nucleotide exchange and competition with tRNA in the yeast elongation factor complex eEF1A:eEF1Bα. Mol. Cell. 6, 1261–1266.
Ptitsyn O.B. 1995. Molten globule and protein folding. Adv. Prot. Chem. 47, 83–229.
Receveur-Brechot V., Bourhis J., Uversky V.N., Canard B., Longhi S. 2006. Assessing protein disorder and induced folding. Proteins. 62, 24–45.
Wuttke D., Foster M.P., Case D.A., Gottesfeld J.M., Wright P. 1997. Solution structure of the first three zinc fingers of TFIIIA bound to the cognate DNA sequence: Determinants of affinity and sequence specificity. J. Mol. Biol. 273, 183–206.
Bruschweiler R., Liao X., Wright P.E. 1995. Long-range motional restrictions in a multidomain zinc-finger protein from anisotropic tumbling. Science. 268, 886–889.
Weikl T., Abelmann K., Buchner J. 1999. An unstructured C-terminal region of the Hsp90 co-chaperone p23 is important for its chaperone function. J. Mol. Biol. 293, 685–691.
Botuyan M.V., Mer G., Yi G.S., Koth C.M., Case D.A., Edwards A.M., Chazin W.J., Arrowsmith C.H. 2001. Solution structure and dynamics of yeast elongation C in complex with a von Hippel-Lindau peptide. J. Mol. Biol. 312, 177–186.
Author information
Authors and Affiliations
Additional information
Original Russian Text © I.N. Serdyuk, 2007, published in Molekulyarnaya Biologiya, 2007, Vol. 41, No. 2, pp. 297–313.
Rights and permissions
About this article
Cite this article
Serdyuk, I.N. Structured proteins and proteins with intrinsic disorder. Mol Biol 41, 262–277 (2007). https://doi.org/10.1134/S0026893307020082
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1134/S0026893307020082