Abstract
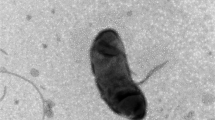
An obligately anaerobic bacterium, strain Bkl1T, was isolated from an enrichment culture of iron-reducing bacteria (IRB) obtained from a sample of the bottom sediments of the cold freshwater Lake Baikal (Russia). Cells of the strain were Gram-stain-negative, motile, spore-forming straight rods (0.6–0.7 × 2.0–7.0 µm) with a fermentative metabolism. Strain Bkl1T grew in the temperature range from 7 to 38°C (optimum 20°C) and at pH 7.0–9.5 (optimum pH 7.6). The novel isolate was capable to reduce ferric citrate (FC), anthraquinone-2,6-disulfonate (AQDS) and Cr(VI) in the presence of lactate as carbon source. Ferrihydrite was not reduced in the absence of AQDS. Based on its 16S rRNA gene sequence, strain Bkl1T was affiliated to the family Sporomusaceae and, more specifically, to the genus Pelosinus. The strain was most closely related to Pelosinus fermentans DSM 17108T (99.2%) and P. propionicus DSM 13327T (99.1%). Genome relatedness indexes revealed that the average nucleotide identity (ANI) and digital DNA-DNA hybridization (dDDH) values between strain Bkl1T and its closest phylogenomic relative (P. fermentans DSM 17108T) were 93.3 and 54.2%, respectively. The G + C content of the genome of strain Bkl1T was 39.1 mol % and its size was 5.32 Mb with 4939 protein-coding genes. The predominant fatty acids in cell walls were С15:1, С17:1, and С16:1. Based on the phylogenetic analyses and phenotypic differences between the novel isolate and type strains of the genus Pelosinus, strain Bkl1T (=VKM B-3511Т = JCM 39258T) is proposed to represent a novel species Pelosinus baikalensis sp. nov.



Similar content being viewed by others
REFERENCES
Andrews, S., FastQC: a quality control tool for high throughput sequence data, 2010. https://github.com/s-andrews/FastQC.
Arkin, A., Cottingham, R., Henry, C., Harris, N., et al., KBase: The United States Department of Energy Systems Biology Knowledgebase, Nature Biotechnology, 2018, vol. 36, pp. 566−569. https://doi.org/10.1038/nbt.4163
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A.A., Dvorkin, M., et al., SPAdes: a new genome assembly algorithm and its applications to single cell sequencing, J. Comput. Biol., 2012, vol. 19, pp. 455–477. https://doi.org/10.1089/cmb.2012.0021
Benson, D.A., Boguski, M.S., Lipman, D.J., Ostell, J., Ouellette, B.F., GenBank, Nucleic Acids Res., 1998, vol. 26, pp. 1−7. https://doi.org/10.1093/nar/26.1.1
Boga, H.I., Ji, R., Ludwig, W., Brune, A., Sporotalea propionica gen. nov, sp. nov., a hydrogen-oxidizing, oxigen-reducing, propionigenic firmicute from the intestinal tract of a soil-feeding termite, Arch. Microbiol., 2007, vol. 187, pp. 15−27. https://doi.org/10.1007/s00203-006-0168-7
Bolger, A.M., Lohse, M., Usadel, B., Trimmomatic: a flexible trimmer for Illumina sequence data, Bioinformatics, 2014, vol. 30, pp. 2114–2120. https://doi.org/10.1093/bioinformatics/btu170.8
Bowen De León, K., Utturkar, S.M., Camilleri, L.B., Elias, D.A., Arkin, A.P., Fields, M.W., Brown, S.D., Wall, J.D., Complete genome sequence of Pelosinus fermentans JBW45, a member of a remarkably competitive group of Negativicutes in the Firmicutes phylum, Genome Announcements, 2015, vol. 3, p. е01090-15.https://doi.org/10.1128/genomeA.01090-15
Brown, S.D., Podar, M., Klingeman, D.M., Johnson, C.M., Yang, Z.K., Utturkar, S.M., Land, M.L., Mosher, J.J., Hurt, R.A., Phelps, T.J., Palumbo, A.V., Arkin, A.P., Hazen, T.C., and Elias, D.A., Draft genome sequences for two metal-reducing Pelosinus fermentans strains isolated from a Cr(VI)-contaminated site and for type strain R7, J. Bacteriol., 2012, vol. 194, p. 5147−5148. https://doi.org/10.1128/JB.01174-12
Cervantes, F.J., van der Velde, S., Lettinda, G., and Field, J.A., Competition between methanogenesis and quinone respiration for ecologically important substrates in anaerobic consortia, FEMS Microbiol. Ecol., 2000, vol. 34, pp. 161–171. https://doi.org/10.1111/j.1574-6941.2000.tb00766.x
Chun, J., Oren, A., Ventosa, A., Christensen, H., Arahal, D.R., da Costa, M.S., Rooney, A.P., Yi, H., Xue-Wei Xu, De Meyer, S., and Trujillo, M.E., Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes, Int. J. Syst. Evol. Microbiol., 2018, vol. 68, pp. 461−466. https://doi.org/10.1099/ijsem.0.002516
Cline, J.D., Spectrophotometric determination of hydrogen sulphide in natural water, Limnol Oceanogr., 1969, vol. 14, pp. 444–458. https://doi.org/10.4319/lo.1969.14.3.0454
Griess-Romijn-van Eck, Physiological and chemical tests for drinking water, NEN 1056, IY–2, 1966, Nederlandse Normalisatie Instituut Rijswijk.
Han, R., Geller, J.T., Yang, L., Brodie, E.L., Chakrabor-ty, R., Larsen, J.T., and Beller, H.R., Physiological and transcriptional studies of Cr(VI) reduction under aerobic and denitrifying conditions by an aquifer-derived pseudomonad, Environ. Sci. Technol., 2010, vol. 44, pp. 7491−7497. https://doi.org/10.1021/es101152r
Hungate, R.E., A roll tube method for cultivation of strict anaerobes, in Methods in Microbiolology, Norris, R. and Ribbons, R.W., Eds., NY: Academic, 1969, vol. 13, pp. 117−132.
Kappler, A., Benz, M., Schink, B., and Brune, A., Electron shuttling via humic acids in microbial iron(III) reduction in a freshwater sediment, FEMS Microbiol. Ecol., 2004, vol. 47, p. 85092. https://doi.org/10.1016/S0168-6496(03)00245-9
Kumar, S., Stecher, G., and Tamura, K., MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets, Mol. Biol. Evol., 2016, vol. 33, pp. 1870−1874. https://doi.org/10.1093/molbev/msw054
Lovley, D.R., Giovannoni, S.J., White, D.C., Champine, J.E., Phillips, E.J.P., Gorby, Y.A., and Goodwin, S., Geobacter metallireducens gen. nov., sp. nov., a microorganism capable of coupling the complete oxidation of organic compounds to the reduction of iron and other metals, Arch. Microbiol., 1993, vol. 159, pp. 336–344. https://doi.org/10.1007/BF00290916
Marmur, J., A procedure for isolation of deoxyribonucleic acid from microorganism, J. Mol. Biol., 1961, vol. 3, pp. 208−218.
Moe, W.M., Stebbing, R.E., Rao, J.U., Bowman, K.S., Nobre, M.F., da Costa, M.S., and Rainey, F.A., Pelosinus defluvii sp. nov., isolated from chlorinated solvent-contaminated groundwater, emended description of the genus Pelosinus and transfer of Sporotalea propionica to Pelosinus propionicus comb. nov., Int. J. Syst. Evol. Microbiol., 2012, vol. 62, pp. 1369−1376. https://doi.org/10.1099/ijs0.033753-0
Parks, D.H., Imelfort, M., Skennerton, C.T., Hugenholtz, P., and Tyson, G., CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes, Genome Res., 2015, vol. 25, pp. 1043–1055. https://doi.org/10.1101/gr.186072.114
Price, M.N., Dehal, P.S., and Arkin, A.P., FastTree 2— Approximately Maximum-Likelihood Trees for Large Alignments, PLoS One, 2010, vol. 5, p. е9490. https://doi.org/10.1371/journal.pone.0009490
Ray, A.E., Connon, S.A., Neal, A.L., Fujita, Y., Cummings, D.E., Ingram, J.C., and Magnuson, T.S., Metal transformation by a novel Pelosinus isolate from a subsurface environment, Front. Microbiol., 2018, vol. 9, p. 1689. https://doi.org/10.3389/fmicd.2018.01689
Reynolds, E.S., The use of lead citrate at high pH as an electron opaque stain in electron microscopy, J. Cell. Biol., 1963, vol. 17, pp. 208−212. https://doi.org/10.1083/jcb.17.1.208
Richter, M., Rosselló-Móra, R., Glöckner, F.O., and Peplies, J., JSpeciesWS: a web server for prokaryotic species circumscription based onpairwise genome comparison, Bioinformatics, 2015, vol. 5, pp. 929–931. https://doi.org/10.1093/bioinformatics/btv681
Shelobolina, E.S., Nevin, K.P., Blakeney-Hayward, J.D., Johnsen, C.V., Plaia, T.W., Kraden, P., Woodard, T., Holmes, D.E., Gaw VanPraagh, C., and Lovley, D.R., Geobacter pickeringii sp. nov., Geobacter argillaceus sp. nov. and Pelosinus fermentans gen. nov., sp. nov., isolated from subsurface kaolin lenses, Int. J. Syst. Evol. Microbiol., 2007, vol. 57, pp. 126−135. https://doi.org/10.1099/ijs.0.64221-0
Slobodkina, G.B., Merkel, A.Y., Novikov, A.A., Bonch-Osmolovskaya, E.A., and Slobodkin, A.I., (Pelomicrobium methylotrophicum gen. nov., sp. nov. a moderately thermophilic, facultatively anaerobic, lithoautotrophic and methylotrophic bacterium isolated from a terrestrial mud volcano, Extremophiles, 2020, vol. 24, pp. 177−185. https://doi.org/10.1007/s00792-019-01145-0
Smibert, R. and Krieg, N., Phenotypic characterization, in: Methods for General and Molecular Bacteriology, Gerhardt, P., Murray, R., Wood, W., and Krieg, N., Eds., Washington: Amer. Soc. Microbiol., 1994, pp. 607–654.
Talavera, G. and Castresana, J., Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments, Systematic Biology, 2007, vol. 56, pp. 564−577. https://doi.org/10.1080/10635150701472164
Wolin, E.A., Wolin, M.J., and Wolfe, R.S. Formation of methane by bacterial extracts, J. Biol. Chem., 1963, vol. 238, pp. 2882−2886
Zemskaya, T.I., Sitnikova, T.Y., Kiyashko, S.I., Kalmychkov, G.V., Pogodaeva, T.V., Mekhanikova, I.V., Naumo-va, T.V., Shubenkova, O.V., Chernitsina, S.M., Kotsar, O.V., Chernyaev, E.S., and Khlystov, O.M., Faunal communities at sites of gas- and oil-bearing fluids in Lake Baikal, Geo-Mar. Lett., 2012, vol. 32, pp. 437–451.
ACKNOWLEDGMENTS
The authors are grateful to T.I. Zemskaya (Laboratory of Carbohydrate Microbiology, Limnological Institute, Siberian Branch, Russian Academy of Sciences) and to O.P. Dagurova (Institute of General and Experimental Biology, Siberian Branch, Russian Academy of Sciences) for providing Lake Baikal bottom sediments.
Funding
This work was supported by the Russian Science Foundation (project no. 22-24-00518).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
DATA AVAILABILITY
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain Bkl1T is MW805760. The whole genome sequence of strain Bkl1T is available at the NCBI (www.ncbi.nlm.nih.gov.) under accession number NZ_JAJHJB000000000.
Rights and permissions
About this article
Cite this article
Zakharyuk, A.G., Kopitsyn, D.S., Suzina, N.E. et al. Pelosinus baikalensis sp. nov., an Iron-Reducing Bacterium Isolated from a Cold Freshwater Lake. Microbiology 92, 137–145 (2023). https://doi.org/10.1134/S0026261722602913
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261722602913