Abstract—
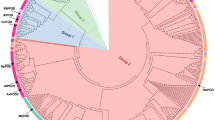
Comparative phylogenetic analysis of NAD+-dependent formate dehydrogenases (NAD+-FDH) genes, which have been detected in all available genomes of methylotrophs of the genera Ancylobacter, Starkeya, and Angulomicrobium, as well as in other members of the family Xanthobacteraceae (Xanthobacter, Aquabacter, Azorhizobium), was carried out. The position of Xanthobacteraceae on the tree constructed based on comparison of NAD+−FDH amino acid sequences was found to correlate with the 16S rRNA gene-based phylogeny. The sequences of the NAD+−FDH proteins of the genera Ancylobacter, Starkeya, and Angulomi-crobium exhibited 87.8–98.3% identity, indicating that this protein is very conservative within this group of methylotrophs. For the first time, analysis of the NAD+−FDH functional genes is recommended as a supplementary criterion for interspecies differentiation between methylotrophic bacteria of the genus Ancylobacter.

Similar content being viewed by others
REFERENCES
Alekseeva, A.A., Savin, S.S., and Tishkov, V.I., NAD+-dependent formate dehydrogenase from plants, Acta Naturae, 2011, vol. 3, no. 4 (11), pp. 38–54.
Chistoserdova, L., Modularity of methylotrophy, revisited, Environ. Microbiol., 2011, vol. 13, pp. 2603–2622.
Chun, J., Oren, A., Ventosa, A., Christensen, H., Arahal, D.R., da Costa, M.S., Rooney, A.P., Yi, H., Xu, X.-W., De Meyer, S., and Trujillo, M.E., Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes, Int. J. Syst. Evol. Microbiol., 2018, vol. 68, pp. 461–466.
Egorov, A.M., Avilova, T.V., Dikov, M.M., Popov, V.O., Rodionov, Y.V., and Berezin, I.V., NAD-dependent formate dehydrogenase from methylotrophic bacterium, strain 1: purification and characterization, Eur. J. Biochem., 1979, vol. 99, pp. 569–576.
Filippova, E.V., Polyakov, K.M., Tikhonova, T.V., Boiko, K.M., Tishkov, V.I., and Popov, V.O., Crystal structures of complexes of NAD+-dependent formate dehydrogenase from methylotrophic bacterium Pseudomonas sp. 101 with formate, Crystallography Rep., 2006, vol. 51, no. 4, pp. 627–631.
Hatrongjit, R. and Packdibamrung, K., A novel NADP+-dependent formate dehydrogenase from Burkholderia stabilis 15516: screening, purification and characterization, Enzyme Microb. Technol., 2010, vol. 46, no. 7, pp. 557–561.
Keltjens, J.T., Pol, A., Reimann, J., and Op den Camp, H.J., PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference, Appl. Microbiol. Biotechnol., 2014, vol. 98, pp. 6163–6183.
McDonald, I.R. and Murrell, J.C., The methanol dehydrogenase structural gene mxaF and its use as a functional gene probe for methanotrophs and methylotrophs, Appl. Environ. Microbiol., 1997, vol. 63, pp. 3218–3224.
Meier-Kolthoff, J.P., Auch, A.F., Klenk, H.-P., and Göker, M., Genome sequence-based species delimitation with confidence intervals and improved distance functions, BMC Bioinformatics, 2013, vol. 14, pp. 1–14.
Nanba, H., Takaoka, Y., and Hasegawa, J., Purification and characterization of formate dehydrogenase from Ancylobacter aquaticus strain KNK607M, and cloning of the gene, Biosci., Biotechnol. Biochem., 2003, vol. 67, no. 4, pp. 720–728.
Ramachandran, A. and Walsch, D.A., Investigation of XoxF methanol dehydrogenases reveals new methylotrophic bacteria in pelagic marine and freshwater ecosystems, FEMS Microbiol. Ecol., 2015, vol. 91, p. fiv105.
Richter, M. and Rosselló-Móra, R., Shifting the genomic gold standard for the prokaryotic species definition, Proc. Natl. Acad. Sci. U. S. A., 2009, vol. 106, pp. 19126–19131.
Shabalin, I.G., Serov, A.E., Skirgello, O.E., Timofeev, V.I., Samygina, V.R., Popov, V.O., Tishkov, V.I., and Kuranova, I.P., Recombinant formate dehydrogenase from Arabidopsis thaliana: preparation, crystal growth in microgravity, and preliminary X-ray diffraction study, Crystallography Rep., 2010, vol. 55, no. 5, pp. 806–810.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S., MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods, Mol. Biol. Evol., 2011, vol. 28, pp. 2731–2739.
Taubert, M., Grob, C., Howat, A.M., Burns, O.J., Dixon, J.L., Chen, Y., and Murrell, J.C., XoxF encoding an alternative methanol dehydrogenase is widespread in coastal marine environments, Environ. Microbiol., 2015, vol. 17, pp. 3937–3948.
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G., The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools, Nucleic Acids Res., 1997, vol. 25, pp. 4876–4882.
Trotsenko, Y.A., Doronina, N.V., and Torgonskaya, M.L., Aerobnye metilibakterii (Aerobic Methylobacteria), Pushchino: ONTI PSC RAS, 2010.
Funding
The work was supported by the Russian Federation Ministry of Science and Higher Education, project no. 075-15-2021-1051.
Author information
Authors and Affiliations
Contributions
All authors analyzed and discussed the data and participated in the preparation of the manuscript.
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of inte-rest.
This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Chemodurova, A.A., Reshetnikov, A.S., Agafonova, N.V. et al. Genes of NAD+-Dependent Formate Dehydrogenases in Taxonomy of Aerobic Methylotrophic Bacteria of the Genus Ancylobacter. Microbiology 91, 834–838 (2022). https://doi.org/10.1134/S0026261722601932
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261722601932