Abstract—
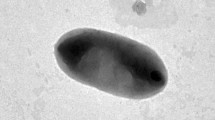
Two strains of ultrasmall gram-negative bacteria (USGNB), FM1 and FM2, were isolated from the skin of the smooth clawed frog Xenopus laevis. The cytological, physiological, biochemical, and genotypic characteristics of the isolates were studied. Based on the sequencing of their 16S rRNA genes and on their phenotypic properties, the isolates were assigned to the genus Chryseobacterium. The cells were extremely small, with cell volumes of ~0.06 and ~0.015 µm3 for developing cultures of strains FM1 and FM2, respectively. Since the USGNB cells were firmly attached to the skin surface and could not be removed by repeated washing with water, these bacteria may be classified as epibionts. Adhesive properties of the fimbria-like appendages revealed in strains FM1 and FM2 by electron microscopy could probably contribute to tight binding of USGNB cells to the skin. Localization of ultrasmall gram-negative bacteria on skin surface of the frogs may indicate their action as a protective bacterial filter; skin surface of Xenopus laevis is thus characterized for the first time as a specific habitat of ultrasmall Chryseobacterium strains. Isolation and characterization of two ultrasmall Chryseobacterium strains, FM1 and FM2, improves our understanding of diversity of the cellular structural and functional characteristics and of the ecological niches of this bacterial genus.








Similar content being viewed by others
REFERENCES
Antwis, R.E., Haworth, R.L., Engelmoer, D.J.-P., Ogilvy, V., Fidgett, A.L., and Preziosi, R.F., Ex situ diet influences the bacterial community associated with the skin of red-eyed tree frogs (Agalychnis callidryas), PLoS One, 2014, vol. 9. e85563.
Becker, M.H. and Harris, R.N., Cutaneous bacteria of the redback salamander prevent morbidity associated with a lethal disease, PLoS One, 2010, vol. 5. e10957.
Belden, L.K. and Harris, R.N., Infectious diseases in wildlife: the community ecology context, Front. Ecol. Environ., 2007, vol. 5, pp. 533–539.
Bell, S.C., Alford, R.A., Garland, S., Padilla, G., and Thomas, A.D., Screening bacterial metabolites for inhibitory effects against Batrachochytrium dendrobatidis using a spectrophotometric assay, Dis. Aquat. Organ., 2013, vol. 103, pp. 77–85.
Bernardet, J.-F., Nakagawa, Y., and Holmes, B., Subcommittee on the taxonomy of Flavobacterium and Cytophaga-like bacteria of the International Committee on Systematics of Prokaryotes. Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family, Int. J. Syst. Evol. Microbiol., 2002, vol. 52, pp. 1049–1070.
Bevins, C.L. and Zasloff, M., Peptides from frog skin, Annu. Rev. Biochem., 1990, vol. 59, pp. 395–414.
Boronina, L.G., Kukushkina, M.P., Krutova, K.V., and Blinova, S.M., Chryseobacterium (Flavobacterium) spp.: clinical significance, identification, antimicrobial susceptibility, Clin. Microbiol. Antimicrob. Chemother., 2003, vol. 5, pp. 243–250.
Brucker, R.M., Harris, R.N., Schwantes, C.R., Gallaher, T.N., Flaherty, D.C., Lam, B.A., and Minbiole, K.P., Amphibian chemical defense: antifungal metabolites of the microsymbiont Janthinobacterium lividum on the salamander Plethodon cinereus, J. Chem. Ecol., 2008, vol. 34, pp. 1422–1429.
Bulet, P., Stöcklin, R., and Menin, L., Anti-microbial peptides: from invertebrates to vertebrates, Immunol. Rev., 2004, vol. 198, pp. 169–184.
Cavicchioli, R. and Ostrowski, M., Ultramicrobacteria, in Encyclopedia of Life Sciences, Chichester: Wiley, 2003. https://doi.org/10.1038/npg.els.0000309
Clay, K., Defensive symbiosis: a microbial perspective, Funct. Ecol., 2014, vol. 28, pp. 293–298.
Conlon, J.M., Structural diversity and species distribution of host-defense peptides in frog skin secretions, Cell. Mol. Life Sci., 2011, vol. 68, pp. 2303–2315.
Conlon, J.M., Mechkarska, M., and King, J.D., Host-defense peptides in skin secretions of African clawed frogs (Xenopodinae, Pipidae), Gen. Comp. Endocrinol., 2012, vol. 176, pp. 513–518.
Duda, V.I., Ultramicrobacteria, in Encyclopedia of Life Sciences, Chichester: Wiley, 2011, pp. 1–23. https://doi.org/10.1002/9780470015902.a0000309.pub2
Euzéby, J.P., List of prokaryotic names with standing in nomenclature–genus Chryseobacterium, 2009, рр. 1–19. www.bacterio.net.
Hector, J.S. and Johnson, A.R., Determination of genome size of Pseudomonas aeruginosa by PFGE: analysis of restriction fragments, Nucleic Acids Res., 1990, vol. 18, pp. 3171–3174.
Kämpfer, P., Lodders, N., Vaneechoutte, M., and Wauters, G., Transfer of Sejongia antarctica, Sejongia jeonii, and Sejongia marina to the genus Chryseobacterium as Chryseobacterium antarcticum comb. nov., Chryseobacterium jeonii comb. nov. and Chryseobacterium marinum comb. nov., Int. J. Syst. Evol. Microbiol., 2009, vol. 59, pp. 2238–2240.
King, J.D., Mechkarska, M., Coquet, L., Leprince, J., Jouenne, T., Vaudry, H., Takada, K., and Conlon, J.M., Host-defense peptides from skin secretions of the tetraploid frogs Xenopus petersii and Xenopus pygmaeus, and the octoploid frog Xenopus lenduensis (Pipidae), Peptides, 2012, vol. 33, pp. 35–43.
Marmur, J., A procedure for the isolation of deoxiribonucleic acid from microorganisms, J. Mol. Biol., 1961, vol. 3, pp. 208–218. http://dx.doi.org/https://doi.org/10.1016/S0022-2836(61)80047-8.
Reynolds, E.S., The use of lead citrate at high pH as an electron-opaque stain in electron microscopy, J. Cell. Biol., 1963, vol. 17, pp. 208–212.
Shaw, S.D., Berger, L., Bell, S., Dodd, S., James, T.Y., Skerratt, L.F., Bishop, P.J., and Speare, R., Baseline cutaneous bacteria of free-living New Zealand native frogs (Leiopelma archeyi and Leiopelma hochstetteri) and implications for their role in defense against the amphibian chytrid (Batrachochytrium dendrobatidis), J. Wildl. Dis., 2014, vol. 50, pp. 723–732.
Suzina, N.E., Duda, V.I., Esikova, T.Z., Shorokhova, A.P., Gafarov, A.B., Oleinikov, R.R., Abashina, T.N., Akimov, V.N., Polivtseva, V.N., and Boronin, A.M., Novel ultramicrobacteria, strains NF4 and NF5, of the genus Chryseobacterium: facultative epibionts of Bacillus subtilis, Microbiology (Moscow), 2011, vol. 80, pp. 535–548.
Suzina, N.E., Esikova, T.Z., Oleinikov, R.R., Gafarov, A.B., Shorokhova, A.P., Polivtseva, V.N., Ross, D.V., Abashina, T.N., Duda, V.I., and Boronin, A.M., Comparative characteristics of free-living ultramicroscopical bacteria obtained from natural biotopes, Appl. Biochem. Microbiol., 2015, vol. 51, pp. 159–168.
Vandamme, P., Bernardet, J.-F., Segers, P., Kersters, K., and Holmes, B., New perspectives in the classification of the Flavobacteria: description of Chryseobacterium gen. nov., Bergeyella gen. nov., and Empedobacter nom. rev., Int. J. Syst. Bacteriol., 1994, vol. 44, pp. 827–831.
Zairi, A., Tangy, F., Bouassida, K., and Hani, K., Dermaseptins and magainins: antimicrobial peptides from frogs’ skin-new sources for a promising spermicides microbicides ‒ a mini review, J. Biomed. Biotechnol., 2009, vol. 2009, article ID. 452567.
Zasloff, M., Magainins, a class of antimicrobial peptides from Xenopus skin: isolation, characterization of two active forms, and partial cDNA sequence of a precursor, Proc. Natl. Acad. Sci. U. S. A., 1987, vol. 84, pp. 5449–5453.
ACKNOWLEDGMENTS
Electron microscopy was carried out at the UNIQEM Collection Common Use Center.
The work was supported by the Presidium of the Russian Academy of Sciences Program of Basic Research no. 32 (Nanostructures: Physica, Chemistry, Biology, Basic Technologies), subprogram 3 (Nanobiotechnologies).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Translated by P. Sigalevich
Rights and permissions
About this article
Cite this article
Ross, D.V., Suzina, N.E., Gafarov, A.B. et al. Characterization of Ultrasmall Chryseobacterium Strains FM1 and FM2 Isolated from Xenopus laevis Skin. Microbiology 88, 172–182 (2019). https://doi.org/10.1134/S0026261719020103
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261719020103