Abstract
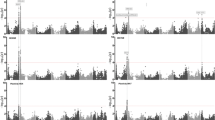
The collection of seeds and living plants at the N.I. Vavilov All Russian Institute of Plant Genetic Resources (VIR) contains historic chickpea landraces sampled from major historic centers of chickpea cultivation and secondary diversification. In this work, we analyze the results obtained from phenotype characterization of 407 historic chickpea landraces grown in the trial fields of Kuban experimental station in 2017. GWAS analysis identified three single nucleotide polymorphisms on chromosomes 2, 7 and 8 that were highly associated with the days to flowering phenotype trait. These single nucleotide polymorphisms and the regions of genome near to these single nucleotide polymorphisms were identified in previous studies. Estimation of the Genotype–Environment interaction for phenotypic traits coincident with plant seeds led to identification of the best genotypes in the environments. These genotypes have an alternative homozygote at loci Ca7:30930779 which shows similar phenotypic traits that were described during phenotype characterization of seeds grown in the trial fields of the Kuban experimental station in 2016 and with the period of flowering typical of phenotype characterization of the plants in 2017. These results can accelerate the search for the seed samples that are the most promising for cultivation of chickpea landraces.




Similar content being viewed by others
REFERENCES
R. J. Redden and J. D. Berger, In Chickpea Breeding and Management, Ed. by S. S. Yadav, R. Redden, W. Chen, and B. Sharma (CABI, Wallingford, UK, 2007), pp. 1–13.
A. Sokolkova, S. V. Bulyntsev, P.L. Chang, et al., Int. J. Mol. Sci. 21, 3952 (2020).
A.B. Sokolkova, P. L. Chang, N. Carrasquila-Garcia, et al., Biophysics (Moscow) 65 (2), 237 (2020).
E. J. von Wettberg, P. L. Chang, A. Greenspan, et al., Nat. Commun. 9, Art. 649 (2018). https://doi.org/10.1038/s41467-018-02867-z
A. McKenna, M. Hanna, E. Banks, et al., Genome Res. 20, 1297 (2010).
P. Danecek, A. Auton, G. Abecasis, et al., Bioinformatics 27, 2156 (2011).
R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria. 2018). https://www.R-project. org/ (accessed on June 15, 2020).
C. Lippert, J. Listgarten, Y. Liu, et al., Nat. Methods 8, 833 (2011).
J. D. Storey, Ann. Stat. 31, 2013 (2003).
T. Olivoto and A. D. Lucio, Methods Ecol. Evol. 11, 783 (2020). https://doi.org/10.1111/2041-210X.13384
R. Chollet, J. Vidal, M. H. O’Leary, Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 273 (1996).
R. K. Varshney, M. Thudi, M. Roorkiwal, et al., Nat Genet. 51, 857 (2019). https://doi.org/10.1038/s41588-019-0401-3
E. Plekhanova, M. A. Vishnyakova, S. Bulyntsev, et al., Sci. Rep. 7, 4816 (2017).
Funding
Phenotyping at the Kuban Experimental Station in 2017, GWAS- and GGE biplot-analyzes were supported by the Russian Science Foundation grant No. 16-16-00007 (for ABS, SVN, MGS). The work was also supported by a cooperation agreement from the United States Agency for International Development within the framework of the Feed the Future program AID-OAA-A-14-00008 (for DC, IF, EHV), Zumburge Foundation (for EHV), Grant IOS-1339346 from the Plant Genome Research Program of the National Science Foundation of the United States (for DC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest.
COMPLIANCE WITH ETHICAL STANDARDS
The study was performed without the use of animals or people as subjects.
Additional information
Translated by P. Kuchina
Rights and permissions
About this article
Cite this article
Sokolkova, A.B., Bulyntsev, S.V., Chang, P.L. et al. A Genomic Analysis of Historic Chickpea Landraces. BIOPHYSICS 66, 32–39 (2021). https://doi.org/10.1134/S0006350921010061
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006350921010061