Abstract
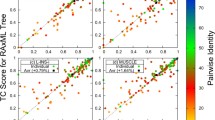
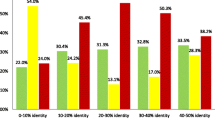
The accuracy of global Smith-Waterman alignments and Pareto-optimal alignments depending on the degree of sequence similarity (percent of coincidence, %id, and the number of removed fragments NGap) has been examined. An algorithm for constructing a set of three to six alignments has been developed of which the best alignment on the average exceeds in accuracy the best alignment that can be constructed using the Smith-Waterman algorithm. For weakly homologous sequences (%id 15, NGap 20), the increase in accuracy is on the average about 8%, with the average accuracy of the global Smith-Waterman alignments being about 38% (the accuracy was estimated on model test sets).
Similar content being viewed by others
References
Mathematical Methods for DNA Sequences, Ed. by M. S. Waterman (CRC Press, Boca Raton, FL, 1989).
G. H. Gonnet, M. A. Cohen, and S. A. Benner, Science 256(5062), 1443 (1992).
M. Zvelebil and J. O. Baum, Understanding Bioinformatics (Garland Science, London, 2007).
S. R. Sunyaev, G. A. Bogopolsky, N. V. Oleynikova, et al., Proteins 54, 569 (2004).
G. Vogt, T. Etzold, and P. Argos, J. Mol. Biol. 249, 816 (1995).
V. Polyanovsky, M. A. Roytberg, and V. G. Tumanyan, Comput. Biol. 15(4), 379 (2008).
I. I. Litvinov, M. Yu. Lobanov, A. A. Mironov, et al., Mol. Biol. 40(3), 533 (2006).
A. Wallqvist, Y. Fukunishi, L. R. Murphy, et al., Bioinformatics 16, 988 (2000).
M. S. Waterman, Proc. Natl. Acad. Sci. USA 80,10 3123 (1983).
T. M. Byers and M. S. Waterman, Oper Res. 32, 1381 (1984).
M. Vingron, Curr. Opinion Struct. Biol. 6(3), 346 (1996).
W. M. Fitch and T. F. Smith, Proc. Natl. Acad. Sci. USA 80, 1382 (1983).
M. S. Waterman, M. Eggert, and E. Lander, Proc. Natl. Acad. Sci. USA 89, 6090 (1992).
D. Fernandez-Baca and S. Srinivasam, Operat. Res. Letters 10, 87 (1991).
D. Gusfield, K. Balasubramian, and K. Naor, in Proc. 3rd Ann. ACM-SIAM Discrete Algorithms (1992), pp. 432–439.
M. A. Roytberg, Preprint (ONTI NTsBI, Pushchino, 1994) [in Russian].
M. A. Roytberg, M. N. Simeonenkov, and O. Yu. Tabolina, Biofizika 44(4). 581 (1998).
I. M. Gukov, T. V. Astahova, M. A. Roytberg, et al., Proc. Moscow Conf. Computat. Molecular Biol. (MCCMB’05, Moscow, 2005), p. 136.
V. Pareto, Manual of Political Economy (A. M. Kelley, New York, 1972).
R. C. Edgar, Nucl. Acids Res. 32(5), 1792 (2004).
S. A. Benner, M. A. Cohen, and G. H. Gonnet, J. Mol. Biol. 229, 1065 (1993).
M. Dayhoff, R. Schwartz, and B. Orcutt, Ed. M. Dayhoff, in Atlas of Protein Sequence and Structure (National Biomedical Research Foundation, Washington, 1978), pp. 345–352.
V. O. Polyanovsky, M. A. Roytberg, and V. G. Tumanyan, Biofizika 53(4), 533 (2008).
G. Vogt, T. Etzold, and P. Argos, J. Mol. Biol. 249, 816 (1995).
F. S. Domingues, P. Lackner, A. Andreeva, and M. J. Sippl, J. Mol. Biol. 297, 1003 (2000).
T. F. Smith and M. S. Waterman, J. Mol. Biol. 147, 195 (1981).
Author information
Authors and Affiliations
Additional information
Original Russian Text © V.V. Yakovlev, M.A. Roytberg, 2010, published in Biofizika, 2010, Vol. 55, No. 6, pp. 965–975.
Rights and permissions
About this article
Cite this article
Yakovlev, V.V., Roytberg, M.A. Increasing the accuracy of global alignment of amino acid sequences by constructing a set of alignment candidates. BIOPHYSICS 55, 891–900 (2010). https://doi.org/10.1134/S0006350910060011
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006350910060011