Abstract
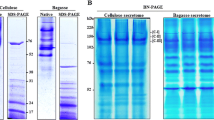
The cellulose-degrading fungus strain Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) Y-94, which was isolated in 1982 from soil in Japan, and strain S6-25, a derivative of Y-94 with higher cellulase productivity, are fungi that exhibit promise as alternatives to Trichoderma reesei for the industrial production of cellulase. MALDI-TOF MS analysis of the T. cellulolyticus Y-94 protein secretome demonstrated that strain Y-94 expressed and secreted a broad spectrum of proteins for the degradation of hemicellulose, starch, or pectin; these proteins include 2 types of α-L-arabinofuranosidases, 2 types of putative glucoamylases, Streptomyces laminarinase-like protein, mutanase, α-amylase, polysaccharide lyase family 6/pectate lyase 3, and β-N-acetylhexosaminidase. The cellulase system of strain S6-25 had the same cellobiohydrolases (Cel6A and Cel7A), β-glucosidase (Bgl3A), xylanase (Xyl10A), and endoglucanase (Cel5A) as strain CF-2612, another derivative of Y-94. However, instead of Cel7B, S6-25 cells expressed another endoglucanase (Cel5C). Two cellobiohydrolases, Cel7A and Cel6A, represented approximately 60% (wt/wt) of the proteins secreted by the strain and were the main enzymes of the cellulase complex. The increase in cellulase complex activity in the S6-25 strain compared with Y-94 may be explained, at least partly, by the presence of an additional copy of the cel7A gene in the cellular genome, which was confirmed by RT-qPCR analysis.



Similar content being viewed by others
REFERENCES
Sajith, S., Priji, P., Sreedevi, S., and Benjamin, S., Int. J. Food Sci. Nutr., 2016, vol. 6, pp. 1–13. https://doi.org/10.4172/2155-9600.1000461
Ikeda, Y., Hayashi, H., Okuda, N., and Park, E.Y., Biotechnol. Prog., 2007, vol. 23, pp. 333–338. https://doi.org/10.1021/bp060201s
Gusakov, A.V., Trends Biotechnol., 2011, vol. 29, pp. 419–425. https://doi.org/10.1016/j.tibtech.2011.04.004
Payne, C.M., Knott, B.C., Mayes, H.B., Hansson, H., Himmel, M.E., Sandgren, M., et al., Chem. Rev., 2015, vol. 115, pp. 1308−1448. https://doi.org/10.1021/cr500351c
Yamanobe, T., Mitsuishi, Y., and Takasaki, Y., Agric. Biol. Chem., 1987, vol. 51, pp 65–74. https://doi.org/10.1080/00021369.1987.10867998
Fujii, T., Fang, X., Inoue, H., Murakami, K., and Sawayama, S., Biotechnol. Biofuels, 2009, vol. 2, p. 24. https://doi.org/10.1186/1754-6834-2-24
Amore, A., Giacobbe, S., and Faraco, V., Curr. Genomics, 2013, vol. 14, pp. 230–249. https://doi.org/10.2174/1389202911314040002
JP Patent no. 4025848, 2007.
Fang, X., Yano, S., Inoue, H., and Sawayama, S., J. Biosci. Bioeng., 2009, vol. 107, pp. 256–261. https://doi.org/10.1016/j.jbiosc.2008.11.022
Inoue, H., Decker, S.R., Taylor II, L.E., Yano, S., and Sawayama, S., Biotechnol. Biofuels, 2014, vol. 7, no. 151, pp. 1–13. https://doi.org/10.1186/s13068-014-0151-5
Fujii, T., Hoshino, T., Inoue, H., and Yano, S., FEMS Microbiol. Lett., 2014, vol. 351, pp. 32–41. https://doi.org/10.1111/1574-6968.12352
JP Patent no. 6515810, 2019.
Laemmli, U.K., Nature, 1970, vol. 227, pp. 680–685. https://doi.org/10.1038/227680a0
Miller, G.L., Anal. Chem., 1959, vol. 31, no. 3, pp. 426–428. https://doi.org/10.1021/ac60147a030
Dumina, M.V., Zhgun, A.A., Novak, M.I., Domratcheva, A.G., Petukhov, D.V., Dzhavakhiya, V.V., et al., World J. Microbiol. Biotechnol., 2014, vol. 30, no. 11, pp. 2933–2941. https://doi.org/10.1007/s11274-014-1721-1
Kansarn, S., Matsushita, N., Kono, T., and Okada, G., J. Appl. Glycosci., 2000, vol. 47, pp. 177–186. https://doi.org/10.5458/jag.50.21
Kansarn, S., Nihira, T., Hashimoto, E., Suzuki, M., Kono, T., and Okada, G., J. Appl. Glycosci., 2000, vol. 47, pp. 293–302. https://doi.org/10.5458/jag.47.293
Mitsuishi, Y., Yamanobe, T., Yagisawa, M., and Takasaki, Y., Agric. Biol. Chem., 1987, vol. 51, pp. 3207–3213. https://doi.org/10.1271/bbb1961.51.3207
Nihira, T., Kansarn, S., Kono, T., and Okada, G., J. Appl. Glycosci., 2001, vol. 48, pp. 45–54. https://doi.org/10.5458/jag.48.45
Nihira, T., Kansarn, S., Kono, T., and Okada, G., J. Appl. Glycosci., 2003, vol. 50, pp. 21–25. https://doi.org/10.5458/jag.50.21
Schulz, B. and Boyle, C., Mycol. Res., 2005, vol. 109, pp. 661–686. https://doi.org/10.1017/s095375620500273x
Morales, G., Melero, J.A., Iglesias, J., and Paniagua, M., J. Adv. Chem. Eng., 2014, vol. 4, no. 1, pp. 1–3. https://doi.org/10.4172/2090-4568.1000e101
Robl, D., da Silva Delabona, P., Mergel, C., Rojas, J.D., dos Santos Costa, P., Pimentel, I.C., et al., BMC Biotechnol., 2013, vol. 94, pp. 13–94. https://doi.org/10.1186/1472-6750-13-94
Seethikal, D.S.D., Govind, S.R., and Jogaiah, S., Int. J. Pharm. Pharm. Sci., 2015, vol. 7, suppl. 1, pp. 205–211.
Baramee, S., Teeravivattanakit, T., Phitsuwan, P., Waeonukul, R., Pason, P., Tachaapaikoon, C., et al., Appl. Microbiol. Biotechnol., 2017, vol. 101, no. 3, pp. 1175–1188. https://doi.org/10.1007/s00253-016-7895-8
Liu, J., Liu, W.D., Zhao, X.L., Shen, W.J., Cao, H., and Cui, Z.L., Appl. Microbiol. Biotechnol., 2011, vol. 89, no. 4, pp. 1083–1092. https://doi.org/10.1007/s00253-010-2828-4
Kamal, S., Khan, S.U., Muhammad, N., Shoaib, M., Omar, M., Kaneza, P., et al., Biochem. Mol. Biol., 2018, vol. 3, no. 1, pp. 15–35. https://doi.org/10.11648/j.bmb.20180301.13
Li, C.X., Zhao, S., Zhang, T., Xian, L., Liao, L.S., Liu, J.L., and Feng, J.X., Sci. Rep., 2017, vol. 7, no. 490, pp. 1–10. https://doi.org/10.1038/s41598-017-00567-0
Portnoy, T., Margeot, A., Linke, R., Atanasova, L., Fekete, E., Sandor, E., et al., BMC Genomics, 2011, vol. 12, no. 269, pp. 1–12. https://doi.org/10.1186/1471-2164-12-269
Sun, J. and Glass, N. L., PLoS One, 2011, vol. 6, no. 9, art. e25654. https://doi.org/10.1371/journal.pone.0025654
Li, Z.H., Yao, G.S., Wu, R.M., Gao, L.W., Kan, Q.B., Liu, M., et al., PLoS Genet., 2015, vol. 11. no. 9, pp. 1–45. art. e1005509. https://doi.org/10.1371/journal.pgen.1005509
Liao, G.Y., Zhao, S., Zhang, T., Li, C.X., Liao, L.S., Zhang, F.F., et al., Biotechnol. Biofuels, 2018, vol. 11, no. 276, pp. 1–17. https://doi.org/10.1186/s13068-018-1276-8
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Supplementary Information
Rights and permissions
About this article
Cite this article
Ptitsyn, L.R., Yampolskaya, T.A., Kutukova, E.A. et al. Identification of Core Cellulolytic Enzymes from the Talaromyces cellulolyticus Strains Y-94 and S6-25. Appl Biochem Microbiol 57 (Suppl 1), S38–S45 (2021). https://doi.org/10.1134/S0003683821100100
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683821100100