Abstract
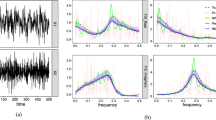
Because the number of parameters required by a Volterra series grows rapidly with both the length of its memory and the order of its nonlinearity, methods for identifying these models from measurements of input/output data are limited to low-order systems with relatively short memories. To deal with these computational and storage requirements one can either make extensive use of the structure of the Volterra series estimation problem to eliminate redundant storage and computations (e.g., the fast orthogonal algorithm), or apply a basis expansion, such as a Laguerre expansion, which seeks to reduce the number of model parameters, and hence, the size of the estimation problem. The use of an appropriate expansion basis can also decrease the noise sensitivity of the estimates. In this paper, we show how fast orthogonalization techniques can be combined with an expansion onto an arbitrary basis. We further demonstrate that the orthogonalization and expansion may be performed independently of each other. Thus, the results from a single application of the fast orthogonal algorithm can be used to generate multiple basis expansions. Simulations, using a simple nonlinear model of peripheral auditory processing, show the equivalence between the kernels estimated using a direct basis expansion, and those computed using the fast, implicit basis expansion technique which we propose. Running times for this new algorithm are compared to those for existing techniques. © 2000 Biomedical Engineering Society.
PAC00: 8710+e, 8719Dd, 4364Bt, 0545-a
Similar content being viewed by others
REFERENCES
Advanced Methods of Physiological System Modeling, edited by V. Z. Marmarelis. New York: Plenum Press, 1994, Vol. III.
Amorocho, J., and A.Brandstetter. Determination of nonlinear functional response functions in rainfall runoff processes. Water Resour. Res.7:1087–1101, 1971.
Barahona, M., and C.-S. Poon. Detection ofnonlinear dynamics in short, noisy time series. Nature (London)381:215–217, 1996.
Boyd, S., and L. O. Chua. Fading memory and the problem of approximating nonlinear operators withVolterra series. IEEE Trans. Circuits Syst.CAS-32(11):1150–1161, 1985.
Golub, G., andC. Van Loan. Matrix Computations. Baltimore: Johns Hopkins University Press, 1989.
Korenberg, M. J. Functional expansions, parallel cascades, and nonlinear difference equations. In:Advanced Methods of Physiological System Modelling, edited by V. Z. Marmarelis. Los Angeles: Biomedical Simulations Resource, 1987, Vol. 1, pp. 221–240.
Korenberg, M. J.Identifying nonlinear difference equation and functional expansion representations: The fast orthogonal algorithm. Ann. Biomed. Eng.16:123–142, 1988.
Korenberg, M. J. A robust orthogonal algorithmfor system identification and time series analysis. Biol. Cybern.60:267–276, 1989.
Korenberg, M. J. Parallel cascade identification and kernel estimation for nonlinear systems. Ann.Biomed. Eng.19:429–455, 1991.
Korenberg, M. J., and I. W. Hunter. Theidentification of nonlinear biological systems: Wiener kernel approaches. Ann. Biomed. Eng.18:629–654, 1990.
Korenberg, M. J., and I. W. Hunter. The identification of nonlinear biologicalsystems: Volterra kernel approaches. Ann. Biomed. Eng.24:250–268, 1996.
Lee, Y. W.,and M. Schetzen. Measurement of the Wiener kernels of a non-linear system by cross correlation. Int. J. Control2:237–254, 1965.
Ljung, L. System Identification: Theory for the user.Englewood Cliffs, NJ: Prentice Hall, 1987.
Marmarelis, V. Z. Practicable identification ofnonstationary nonlinear systems. IEE Proc.-D: Control Theory Appl.128:211–214, 1981.
Marmarelis, V. Z. Identification of nonlinear biological systems using Laguerre expansions of kernels.Ann. Biomed. Eng.21:573–589, 1993.
Marmarelis, V. Z. Modeling methodology fornonlinear physiological systems. Ann. Biomed. Eng.25:239–251, 1997.
Ogura, H.Estimation of Wiener kernels of a nonlinear system and fast algorithm using digital Laguerre filters. Proc NIBB Conf.15:14–62, 1986.
Sakai, H. M. White-noise analysis in neurophysiology.Physiol. Rev.72:491–505, 1992.
Watanabe, A., and L. Stark. Kernel method fornonlinear analysis: identification of a biological control system. Math. Biosci.27:99–108, 1975.
Westwick, D. T., and R. E. Kearney. Generalized eigenvector algorithm for nonlinear systemidentification with non-white inputs. Ann. Biomed. Eng.25:802–814, 1997.
Westwick, D. T., and R. E. Kearney. Identification of physiological systems: A robust method for non-parametric impulse response estimation. Med. Biol. Eng. Comput.35:83–90, 1997.
Westwick, D.T., and R. E. Kearney. Nonparametric identi-fication of nonlinear biomedical systems, part I: Theory. Crit. Rev. Biomed. Eng.26:153–226, 1998.
Westwick, D. T., B. Suki, and K. R. Lutchen. Sensitivity analysis of kernel estimates: Implications in nonlinear physiological system identification. Ann. Biomed. Eng.26:488–501, 1998.
Wiener, N. Nonlinear Problems in RandomTheory. New York: Wiley, 1958.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Westwick, D.T., Lutchen, K.R. Fast, Robust Identification of Nonlinear Physiological Systems Using an Implicit Basis Expansion. Annals of Biomedical Engineering 28, 1116–1125 (2000). https://doi.org/10.1114/1.1310217
Issue Date:
DOI: https://doi.org/10.1114/1.1310217