Abstract
Understanding the mechanism of rabies virus (RABV) infection is vital for prevention and therapy of virulent rabies. However, the infection mechanism remains largely uncharacterized due to the limited methods and viral models. Herein, we utilized a powerful single-virus tracking technique to dynamically and globally visualize the infection process of the live attenuated rabies vaccine strain-SRV9 in living Vero cells. Firstly, it was found that the actin-enriched filopodia is in favor of virus reaching to the cell body. Furthermore, by carrying out drug perturbation experiments, we confirmed that RABV internalization into Vero cells proceeds via classical dynamin-dependent clathrin-mediated endocytosis with requirement for intact actin, but caveolae-dependent endocytosis is not involved. Then, our real-time imaging results unambiguously uncover the characteristics of viral internalization and cellular transport dynamics. In addition, our results directly and quantitatively reveal that the intracellular motility of internalized RABV particles is largely microtubule-dependent. Collectively, our work is crucial for understanding the initial steps of RABV infection and elucidating the mechanisms of post-infection. Significantly, the results provide profound insight into development of novel and effective antiviral targets.
Similar content being viewed by others
Introduction
The rabies is an acute zoonotic disease with highly fatality, causing in excess of 50 000 human deaths annually, the majority of which occur in Asia1. As the culprit of the rabies, rabies virus (RABV) belonging to the genus Lyssavirus of the Rhabdoviridae family, is highly neurotropic2,3. RABV is characterized as an enveloped, nonsegmented, negative single stranded RNA virus encoding five proteins, N, P, M, G and L4. This nucleocapsid of rabies virus is surrounded in “bullet” by a lipid bilayer containing the transmembrane glycoprotein G, which is exposed on the virus surface as a trimeric spike5,6. The RABV G plays a critical role in the infectious cycle7,8,9. The human health has been severely threatened by rabies virus for a long time. To design antiviral drug, it is urgent to deeply investigate the cellular mechanisms underlying the viral infection.
The viral infection was determined as a complex process, which involves in the virus-cell attachment, the viral entry, the membrane fusion and the intracellular transport from the cytosol entry site to gene replication site10,11,12,13,14,15. Inherently, the viral infection process is a very intriguing virus-cell interaction, in which viruses try their best to take advantage of host cell machineries for efficient infection. The first step of viral infection is viral attachment to cell surface. Remarkably, after attachment, viruses have been frequently observed to exploit dynamic cell surface protrusions-filopodia to gain access into the entry sites at the cell body16,17,18,19. Most animal viruses make use of receptor-mediated endocytosis to enter their host cells. The endocytic pathway includes various types, of which clathrin-mediated endocytosis (CME) and caveolae-dependent endocytosis (CavME) are leading entry pathways20. Subsequently, to overcome the barrier from the complex cellular environment, the internalized viruses have adapted to utilize the cellular transport systems to reach a specific site for genome release21. In this process, many viruses have been reported to exploit the microtubules as transport tracks, such as adenovirus, herpes simplex virus, HIV and influenza virus10,15,22,23.
Understanding the mechanism of virus infection is vital for therapy of viral diseases. It has been established that rabies virus enter the host cell via receptor-mediated endocytosis and subsequent low pH-dependent fusion24,25 and the endocytic pathway is proven to be clathrin-mediated26. However, these previous experimental methods were mainly based on electron microscopy (EM), which cannot disclose the dynamic process of viral infection with the disadvantage to only acquire static images. The appearance of real-time fluorescence imaging techniques solves this problem. In recent years, it has been applied to the investigation of RABV infection mechanism including the entry pathway and intracellular transport27,28. Moreover, live cell imaging technique has indicated that RABV exploits axonal transport mechanisms during CNS invasion by interacting with p75NTR29. However, the life cycle of rabies virus is very sophisticated and many questions about RABV infection mechanism remain unknown30. Therefore, it is imperative to develop a novel viral model to combine with the excellent method to further investigate the infectious mechanism.
In this study, the live attenuated rabies vaccine strain-SRV9 was used as the novel viral model to investigate the infection mechanism of RABV. Compared to recombinant RABV, apart from the advantage of production convenience, the live and integral rabies strain with all the characteristics of RABV morphologically and compositionally, is a good alternative to further investigate the infection mechanism of authentic RABV. Significantly, we employed real-time tracking technique to monitor individual SRV9 particles in living cells, which allows us to intuitively visualize dynamic interactions between viruses and host cells. The combination of the novel model with the excellent method facilitates us to further characterize the authentic RABV infection mechanism. Our results globally and directly revealed the infection process of RABV exploited by a single virus including virus-cell attachment, internalization and intracellular trafficking, which not only support the previous results from recombinant viruses, but also provide some novel findings.
Results and Discussion
Biological characterization of purified SRV9
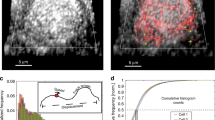
The SRV9 belongs to living attenuated rabies vaccine strain with good safety and immunogenicity, resulting from the alteration of Arg→Ser in the 333 position of glycoprotein of rabies viruses31. The amino acid mutation does not alter the features of rabies viruses in morphology. The electron microscopic analysis showed that the SRV9 particles are in bullet shape and measured on average 180nm in length and 80nm in width (Fig. 1A). The SRV9 is morphologically consistent with the wild type (WT) RABV. To carry out single particle fluorescence imaging, we covalently labeled the glycoprotein of SRV9 with fluorescent dye molecules Cy5-NHS. The Cy5 is an amino-active dye that spontaneously binds with the glycoproteins and its surface density was sufficiently high so that its fluorescence allowed the labeled SRV9 to be clearly detected. The confocal laser scanning microscopy (CLSM) was used to image the Cy5-labeled SRV9 particles adsorbed onto glass coverslips. As shown in Fig. 1B, some scattered individual SRV9 particles were observed, which indicates that the purified SRV9 particles were suitable for single particle tracking experiment. Next, we co-incubated the cells with labeled SRV9 particles for 30 min at 37 °C. The confocal image revealed that numerous virus particles had entered the cells and distributed throughout the interior of the cell, including the cell periphery and perinuclear positions (Fig. 1C). The result indicated that the labeled SRV9 still enable to enter the cell.
Biological characterization of purified SRV9.
(A) Electron micrograph of density gradient-purified SRV9 negatively stained with PTA. (B) Confocal image of Cy5-stained SRV9 particles immobilized on glass coverslips. (C) The fluorescence image of Vero cells that internalized Cy5-labled SRV9 particles. The nucleus region was circled by white dashed line.
To study whether the dye-labeling has effect on the infectivity of viruses, we carried out the immunofluorescence assay and Western blotting analysis (Figure S1 and Figure S2 in the Supplementary Information). The results suggested that the dye-labeling has no significant effect on the infectivity of viruses. These experimental results suggest that we have successfully purified relatively homogeneous populations of SRV9 particles. The fluorescence labeled SRV9 particles are well appropriate for single-particle tracking and imaging. In this work, we will choose the SRV9 as a new model to dissect rabies virus infection mechanisms.
SRV9 are associated with filopodia prior to viral entry
Generally, virus-cell interaction starts with the attachment of the virus particle to the cell surface. After the attachment, the virus reaches to endocytic zones at the cell body to complete the internalization. It has been reported that the member of Rhabdoviruses VSV was directly transported into clathrin-coated pits by filopodia after the cell attachment18. The filopodia are actin-enriched cell surface protrusions and considered as the feeler of the cell to probe the extracellular environment. However, whether filopodia actively contribute to SRV9 entry is still unclear. Herein, we applied live-cell fluorescence imaging to investigate the process of viral entry.
To better perform the fluorescence imaging experiment, a labeling strategy was taken, in which the Vero cells and SRV9 were labeled with the lipophilic membrane dye DiO and Cy5-NHS, respectively. In this process, the filopodia as cell surface protrusion were also stained by DiO.
To observe the association of SRV9 with the filopodia, we implemented the following contrasting experiment. Visualization of the Vero cells in unexposed SRV9 revealed few filopodia (Fig. 2A; mock). Notably, after cells exposured to SRV9 for short periods of time, a strong induction of filopodia was observed (Fig. 2A; +SRV9). Simultaneously, we quantitatively analyzed the effect of the SRV9 exposure on the number of filopodia per cell and the average length of filopodia in more than 100 cells. (Fig. 2B,C). The results indicated SRV9 addition has activated more filopodia production and elongation, which may induce increased virus uptake from the surrounding environment.
The correlation of the viruses with filopodia.
(A) The confocal images of Vero cells (green) that unexposed or exposed to SRV9. (B) The number of filopodia per cell with or without SRV9 exposure was quantified. Error bars represent standard error of the mean (SEM). (C) The average length of filopodia with or without SRV9 exposure was quantified. Error bars represent SEM. In D and E, SRV9 (red) were added to Vero cells (green) and immediately monitored over time by confocal fluorescence microscopy. (D) Arrowheads mark the movement of single virus particles via retrograde transport from the filopodium periphery toward the cell body. (E) An example of single virus particles reaching the cell body through filopodial retraction where the filopodial tip is marked by arrowheads. Scale bars: 10 μm.
Next, we further investigated whether filopodia structures play a role in viral entry. Time-lapse confocal microscopy was performed on live Vero cells infected by SRV9. We observed retrograde viral transport of an individual viral particle (i.e., “surfing”) along filopodia until reaching the cell body (Fig. 2D, see Movie S1 in the Supplementary Information). In addition to the viral active transport, another mode of viral particle getting to the cell body via filopodia is also observed, which is termed as filopodial retraction. As observed (Fig. 2E, see Movie S2 in the Supplementary Information), the individual viral particle attached to the filopodia was transferred to cell body as the filopodia retracted. Meanwhile, another filopodia served as the “helper” filopodia in the homing process of the viral particle towards the cell body. The similar phenomenon has been found in other studies32.
Our results indicate that SRV9 are closely related to the filopodia in viral entry. This is in accordance with the results of previous studies19,33. Meanwhile, the coincidence with the behavior of VSV suggests that the Rhabdoviridae viruses possibly share the same mechanism in the process of viral reaching to the cell body. Movement along filopodia may represent an efficient infectious pathway, by which viruses can rapidly and directly reach the endocytic zones for entry into cells. However, the activation mechanism of filopodia needs further investigation.
Entry of SRV9 into Vero cells is via clathrin-mediated and actin-dependent pathway rather than caveola-dependent
Although the entry mechanism of RABV has been reported, the reconstructed RABV were used in previous studies8,27. Here, we employed live SRV9 as the surrogate of wild-type RABV to comprehensively evaluate the endocytosis pathway. Several disruption drugs that specifically hinder different entry pathways were utilized. In the inhibition experiments, cells were pretreated with various inhibitors prior to SRV9 infection. The inhibitors were maintained throughout the experiment process. As positive control, the Vero cells are infected by SRV9 without any drug pretreatment. As shown in Fig. 3A, a large number of virions were seen inside the untreated control cell.
SRV9 Entry depends on clathrin-mediated endocytosis rather than caveola-dependent.
The Vero cells were labeled by DiO (green). In the control group, the Vero cells were incubated with Cy5-labeled SRV9 for 30 min at 37 °C and washed by PBS immediately before imaging. In treated group, the Vero cells were pretreated by low temperature, as well as several inhibitors including sucrose (200 mM), CPZ (10μg/ml), CB (20 μM), MβCD (10 mM), nystatin (100 μM) and filipin (10 μg/ml) for 30 min prior to SRV9 infection, respectively. More than 100 cells were examined by CLSM in each treatment. (A) Three-dimensional confocal images of Vero cells infected with SRV9 under different treatment conditions. Successive z-stacks spaced by 200 nm were recorded to construct the 3D image. The XY image plane is located about 4 μm above the bottom of the cell. The XZ image plane that used for recording in all experiments is approximately 1;μm thick as indicated by the white dashed lines. Scale bars: 10 μm. (B) Quantitation of viral entry rate. The viral entry rate was quantified as the average fluorescence intensity of virus-infected cell relative to that for control cells. The viral entry rate of control cells was factitiously set as 100%. Error bars represent SEM. (C) SRV9 particles colocalize with clathrin. Vero cells were transfected with pEGFP-LCa prior to incubation with Cy5-labeled SRV9. The arrows indicate an individual SRV9 particle (red) colocalized with clathrin (green). Scale bars are 10 μm.
First, to verify whether or not the SRV9 enters the cell via endocytosis pathway, we incubated Vero cells with SRV9 at 4 °C for 30 min. Compared to the control cells, the low temperature pretreatment made the SRV9 internalization drastically inhibited (>98%) (Fig. 3A,B), suggesting the endocytosis pathway was exploited. Then, the role of clathrin-mediated pathway in SRV9 entry was assessed by using sucrose and CPZ (Chlorpromazine). Hypertonic sucrose is generally considered as an inhibitor of CME by causing dissociation of clathrin vesicles from the plasma membrane34, while CPZ inhibition on CME is attributable to blocking the assembly of clathrin-coated pits35. Compared to the control, pretreatment of Vero cells with sucrose (200 mM) or CPZ (10 μg/mL) strongly inhibited SRV9 internalization, about 99% and 95%, respectively (Fig. 3A,B). Dynamin is an essential cellular GTPase involved in clathrin-dependent and lipid raft/caveola-dependent endocytic pathways36. To validate the role of dynamin in SRV9 entry, we treated the Vero cells using dynasore, which rapidly and specifically inhibits the GTPase activity of dynamin37. SRV9 particles were obviously reduced in infected dynasore-treated cells (Figures S3 in the Supplementary Information). Furthermore, to further determine if SRV9 particles were associated with clathrin, Vero cell were transfected with pEGFP-LCa and infected with SRV9. As shown in Fig. 3C, some virus particles were observed to colocalize with clathrin clusters (green) as indicated by white arrows. The proportion of colocalization is statistically analyzed at about 58.52% when virus infecting for 30min. Meanwhile, the dynamic process of single virus particle entry into cell through the clathrin is observed using real-time tracking technique (Movie S3 in Supplementary Information). Together, these results indicate that dynamin-dependent and clathrin-mediated endocytosis is involved SRV9 entry.
Recently, some studies have revealed that the rhabdoviruses VSV, IHNV and ABLV are internalized through clathrin-dependent pathways with the help of actin38,39,40. The contribution of actin to SRV9 entry was studied using the actin-depolymerizing drug CB (cytochalasin B). As expected, the entry of SRV9 was seriously inhibited by the pretreatment of cells with cytochalasin B (>97%) (Fig. 3A,B), suggesting that clathrin-mediated SRV9 internalization is associated with actin polymerization. Meanwhile, the effect of CB inhibition on viral entry also may be correlated with inhibition of actin-enriched filopodia.
As reported previously, some viruses have exploited more than one type of endocytic pathway to efficiently enter host cells41,42,43,44. Accordingly, we determined whether or not the other pathway is involved in SRV9 entry into host cells. MβCD (methyl-β-cyclodextrin) is generally used to extract cholesterol from the membranes. When the Vero cells were pretreated with MβCD, the uptake of SRV9 was apparently reduced by around 95% (Fig. 3A,B). Although the caveolae formation is strictly dependent on cholesterol45, the cholesterol depletion by MβCD also inhibits clathrin-coated pit (CCP) budding46. Consequently, the result merely shows that the uptake of SRV9 was dependent of membrane cholesterol. This finding is compatible with a clathrin- or caveolae- mediated entry pathway for SRV9.
To further test the contribution of CavME to SRV9, the effect of the cholesterol-binding drug nystatin or filipin was studied. Due to the caveolae invagination requiring for cholesterol, isolation with sterol-binding drugs nystatin or filipin will dramatically block caveola-dependent endocytosis. However, pretreatment of Vero cells with nystatin or filipin almost did not obviously decrease the uptake of SRV9 into Vero cells (25% and 2%, respectively) (Fig. 3A,B). These results suggest that SRV9 entry into Vero cells is caveolae-independent.
Taken together, our results intuitively disclosed that SRV9 was internalized into Vero cells by a clathrin-dependent and actin-dependent pathway similar with VSV, IHNV and ABLV. Besides, the CavME is proven to not contribute to SRV9 entry into Vero cells. Through overall analysis of live SRV9 entry pathway, our results reflect the entry pathway of WT rabies virus to the maximum. These data presented here further reveal that the rhabdoviruses, such as VSV, IHNV, ABLV, RABV, may exploit the same endocytosis mechanism38,40,42.
Single-virus tracking of SRV9 entry and its intracellular transport
To investigate the infection process of individual SRV9, we observed the internalization and the transport of SRV9 by tracking single virus in live cells using time-lapse confocal laser scanning microscope. The SRV9 and Vero cell membranes were labeled with Cy5-NHS and DiO, respectively. The cellular vesicles were also nonspecifically stained by the lipophilic dye DiO. Firstly, the viruses were added to Vero cells for 10 min at 37 °C, then tracked in real-time. Figure 4A represents the image of Vero cells in bright field. The border of the Vero cell corresponding to that in the fluorescence images is marked by blue line. It is easy to identify the sites of SRV9 entry through visualizing the fusion site of SRV9 and the Vero cell membrane. The time-lapse confocal microscope image showed that the red fluorescences (Cy5-labeled SRV9) were colocalized with the green fluorescences (DiO-labeled cell membrane) as circled by white dashed line in Fig. 4B. The colocalization indicated that the virus was attached to cell membrane. After attachment, the virus continued to interact with the cell membrane for a short time (Fig. 4B,C). The interaction promoted the cell membrane to invaginate to form vesicle, as shown in Fig. 4D, then the virus was wrapped in the vesicle. At this moment, the vesicle was still located at cell membranes. The preexisting vesicle (pointed by red arrow in Fig. 4B) bound to the new-forming vesicle, which provided tensile force to facilitate the vesicles to detach from the membrane. In Fig. 4E, the single virus had already escaped from the vesicle and entered the cell. Afterwards, the virus fused with the preexisting vesicle and moved towards the cytoplasm. The whole internalization process of SRV9 was recorded in Movie S4 (Supplementary Information).
The process of individual SRV9 entry into Vero cell.
(A) Bright-field image of a Vero cell as circled by blue line. (B–H) Selected frames reveal the process that SRV9 particle was internalized into the Vero cell. In this experiment, Vero cells labeled with DiO (green) were infected with Cy5-labeled SRV9 (red). The imaging was performed at 10 min after the infection. The white dashed circles show the site of SRV9 entry into the Vero cell. Scale bar: 10 μm.
Next, to further reveal the infection process, we utilized the real-time tracking to monitor the individual virus transport in live cells. As shown in Fig. 5, a single virus was imaged in the cell as pointed by white arrow and the cell nucleus was circled by the blue dashed line. The typical trajectory of a virus starting in the cell periphery, moving to the perinuclear region and finally locating there is shown in Fig. 5. The rapid movement process is recorded in Movie S5 in Supplementary Information. In short, the single-virus tracking is an excellent tool to follow the fate of individual virus particle in live cells47. By means of this technique, for the first time, we directly revealed characteristics of the internalization and transport behavior of single SRV9 particle in living cells, which is significant for understanding the initial stage of viral infection process.
The process of individual SRV9 transport in a Vero cell.
Vero cells were infected with SRV9 (red) and imaged 10 min after the infection. The nucleus region was circled by blue dashed line. The white arrow points a single virus inside the cell. The trajectory of a Cy5-labeled virus inside a cell is shown by yellow line. The movement of the virus was recorded by real-time confocal microscopy. The time interval of serial images was 10 s. Scale bar: 20 μm.
Rapid microtubule-dependent Motility of SRV9
Upon internalization, virus usually exploits cellular transport system to deliver their genomes to specific compartments for replication. To observe the movement of SRV9 in cells in detail, we infected DiO-labeled Vero cells with Cy5-labeled SRV9. Then, the time-lapse individual particle imaging was performed in living cells using confocal microscope. The imaging was carried out about 30 minutes after virus infection. At this moment, the great majority of particles had entered cells and exhibited high motility inside the cells.
Figure 6A and Movie S6 in Supplementary Information represent typical examples of SRV9 motility in Vero cells. The Matlab software allowed us to analyze the trajectory of single-virus particle inside the cell, with the ability to distinguish the virions inside or on the cell from outside the cell. The analytical results were displayed in Fig. 6C, showing various types of movements. Most of viruses particles were able to achieve relatively high transport speeds (marked by the green arrow in Fig. 6C), despite some particles showed comparatively low speed motion (marked by the yellow arrow in Fig. 6C). The SRV9 particles were able to achieve speeds nearly up to 0.2 μm s−1 (blue trajectory velocity profile in Fig. 6E). Some of these relatively rapid moving particles displayed highly directed movements. It is noted that the slowly moving particles were localized to vesicular structures nonspecifically stained by the lipophilic dye DiO. However, the relatively rapid moving particles didn’t colocalized with the vesicular structures, which means that these viruses were possibly fused with specific endocytic vesicles that need to be labeled by specific makers or were free inside the cytosol. It is worth noting that the actual speeds are virtually much higher than the speeds calculated from the two-dimensional projection. The reason accounting for the deviation is that the virus particles moved inside the cell typically in all three dimensions, but only a two- dimensional projection of three-dimensional motion was measured.
Motility of SRV9 in the untreated, nocodazole-treated Vero cells.
(A,B) Confocal images of SRV9 (red) in live cells labeled with DiO dye (green) under untreated and nocodazole-treated conditions, respectively. (C, D) Representative trajectories corresponding to the untreated and nocodazole-treated images, respectively. Their instantaneous speeds as a function of time are shown in E and F. The green arrow and the yellow arrow in (C) indicate relatively high transport speed and low transport speed in a limited region, respectively. Scale bar: 10 μm.
As a fundamental component of cytoskeleton, microtubule is usually essential for the intracellular transport of many viruses. To validate whether SRV9 motility in cell was the microtubule-dependent, the effect of microtubule-disrupting drug was evaluated. In this experiment, cells were pretreated with nocodazole prior to the incubation with SRV9. By contrast with the control, the treatment of cells with nocodazole that can depolymerize microtubules resulted in great reduction of the viral motility. A large number of SRV9 trajectories inside the cell were analyzed (Fig. 6B), showing that the majority of SRV9 particles were almost static (Fig. 6D, Movie S7 in Supplementary Information). The trajectory velocity profile in Fig. 6F also revealed that the transport speed was extremely low. This result initially indicated that the microtubules may be involved in the intracellular transport of SRV9.
Next, we performed the colocalization experiment to analyze the involvement of microtubules in the intracellular motility of SRV9 (Fig. 7). The microtubules were recognized by antibody against tubulin. In the magnified image (inset), these yellow signals as indicated by white arrows represented the overlap of the microtubules (green) and Cy5-labeled SRV9 (red), indicating that some SRV9 colocalized with the microtubules. Our statistical result indicated that about 77.6% of SRV9 virions are colocalized with microtubules at 30 min for virus infection. In addition, SRV9 particles were observed to move along microtubules during infection (see Movie S8 in the Supplementary Information). The results suggested that microtubules may be crucial for virus transport during infection.
Colocalization of SRV9 particles with microtubules.
Immunofluorescence was performed at cells infected by Cy5-labele SRV9 (red) after 30 min using anti-tublin antibody (green). The white square region is magnified and shown in the inset. The white arrows represent some examples of colocalization of Cy5-labeled SRV9 with FITC-tagged microtubules. Scale bar: 10 μm.
Because of complexity and heterogeneity of the intracellular environment, the viruses can exploit more than one transport system simultaneously to efficiently infect their hosts48,49. Therefore, it is necessary to characterize and quantitatively analyze the viral motility by tracking a large number of virus particles in cells. The statistical analyses were implemented on tens of thousands of trajectories in many independent experiments. The Fig. 8A,B presented statistics histogram of speed in untreated cell and nocodazole-treated cell, respectively. The statistical analysis from the two group cells is shown in Fig. 8C, displaying that the percentage of trajectories with peak speeds >0.05 μm s−1 is 24.7% in untreated cells and 5.9% in nocodazole-treated cells. The 4.2-fold reduction in the speed of SRV9 motility occurred in microtubule-depolymerized cells. Therefore, we speculated that SRV9 motility was closely related to the microtubules in living cells. Recently, the microtubule-mediated behavior of virus motility has been proven as a complex event50. Future work will be required to determine whether this microtubule-associated motility way ultimately contributes to productive infection and if so what underlying mechanism is involved.
Due to high fatality of RABV, there have been extensive studies on RABV infection mechanism. It is a pity that the thorough understanding of the infection mechanism is limited by methods and models. Compared to previous studies, in this work, we utilized the real-time single-virus tracking method to study the infection mechanism of the live attenuated rabies vaccine strain-SRV9. The real-time tracking represents an excellent system that allowed us to gain a detailed observation of the infection process of single-virus particle, while the live attenuated rabies vaccine strain-SRV9 as a novel model enables to further reflect the infection characteristics of the wide-type RABV.
In summary, we combined the advanced imaging technique with a novel viral model to globally investigate the infection mechanism of RABV. The data presented in this study suggest that RABV internalization into host cells occurs through a dynamin-dependent CME pathway, rather than caveolae-dependent. Furthermore, the results showed that the actin cytoskeleton plays an indispensable role in efficient virus entry. The intact actin is necessary for the clathrin-mediated uptake of RABV, meanwhile, the actin-enriched filopodia are significant for viral attachment to the cell body. Our results revealed that the RABV may have the same endocytosis mechanism as other members of the Rhabdoviruses, such as VSV, IHNV and ABLV. Additionally, the real-time imaging results displayed the characteristics of viral internalization and cellular transport dynamics and revealed that the viral cellular transport is largely microtubule- dependent. Our work is not only significant for deeply understanding the mechanism of viral infection, but also paves the way for the development of novel and effective antiviral targets.
Methods
Cell and virus
Vero cells (from Shanghai Institute of Biological sciences) were maintained in a 5% CO2 environment at 37 °C in minimum Eagle medium (MEM, GIBCO) with 10% FBS (GIBCO), penicillin (100 internationalunits mL−1) and streptomycin (100 μg mL−1) and passaged every 2–3 days. SRV9 were amplified in Vero cells. The titer of virus is detected by sandwich ELISA. The cultures with high content of viruses were collected. One part were freezed at −70 °C as reserve and the other were prepared for purification. The cultures were concentrated by using Zn (AC)2, then purified in linear 10% ~ 50% sucrose gradients. Concentrated viruses stocks were stored in PBS containing 10 mM HEPES (pH 7.4) at −80 °C.
Fluorescence Labeling of SRV9
Amine-reactive Cy5 (OPE TECH) was solubilized in DMSO at the concentration of 1mg/mL. To label SRV9, Cy5 was incubated with purified viruses at a final concentration of 1 μg/mL in a carbonate buffer (pH 9.3) at room temperature for 2 h with gentle vortexing. To remove the unbound Cy5, the virus mixture was centrifuged at 17000 g, finally eluted in PBS (pH 7.4) and stored at −80 °C. The labeled viral aggregates were immediately removed with 0.2 μm pore size filters before experiments and examined under electron microscopy or fluorescence microscope.
Drug Treatment
To perform fluorescence imaging, Vero cells were cultured in MEM with 10% FBS for 2 or 3 days to achieve 75% confluence in 35-mm glass-bottomed culture dishes (Shengyou Biotechnology Co., Ltd). The cells were pretreated with various chemical inhibitors purchased from Sigma. The pretreatment of cells with nocodazole (60 μm) or cytochalasin B (20 μm) was for 50 min, respectively; while cells were preincubated with sucrose (450 mM), chlorpromazine (CPZ, 10 μg/mL), methyl-β-cyclodextrin (MβCD 10 mM), Filipin III (10 μg/mL), Nystatin (100 μm), respectively, for 30 min. The sucrose and CPZ were used to inhibit clathrin-mediated endocytosis; filipin III and nystatin were used to inhibit caveola-mediated endocytosis; MβCD was used to inhibit cholesterol content-associated endocytic pathway. The drugs were maintained in the cell culture throughout the experiments. After drug treatment, the cells were incubated with the viruses at 37 °C in a CO2 incubator for 30 min before experiments. Cells were washed with PBS for 3 times, then incubated in serum-free and phenol red-free MEM as prepared for experiments.
Plasmids and Transfection
Two plasmids that respectively encodes GFP -α- tubulin, eGFP - LCa (clathrin light chain molecules) were used to transfect the Vero cells in this study. After cells were plated for 24 h, the transfection was performed by using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. The cells were subsequently cultured for 24 ~ 48 h prior to imaging acquisition.
Immunofluorescence microscopy
Cells were fixed with 4% paraformaldehyde, permeabilized with 0.2% Triton X-100 for 15 min and blocked with bovine serum albumin (BSA) (Sigma). Cells were then washed three times with PBS and incubated with primary antibodies against tubulin (1:100) diluted in 0.2% BSA at room temperature for 2 h. After washing, FITC-conjugated goat anti-mouse IgG was added and left at room temperature for 1h. Then, the cells were incubated with Hoechst 33342 at room temperature for 5 min after washing. Cells were imaged using confocal microscopy.
Immobilization of virions on glass coverslips
The purified and Cy5-labeled virions were immobilized as described as follows. In brief, the virions were loaded onto glass coverslips and adsorbed for 2 h at room temperature in the dark. After adsorbed onto the glass surface, the virions were directly analyzed by confocal laser scanning microscopy.
Electron microscopy
The purified virus particles were deposited onto carbon-coated copper grids and stained with 2% phosphotungstic acid (PTA) in H2O (pH 7.5). Then, the virus particles were imaged using a Tecnai G2 Spirit BioTWIN transmission electron microscope (FEI, Hillsboro, OR).
Fluorescence Microscopy and Image Analysis
The live cell fluorescence imaging was carried out using a Leica TCS SP2 confocal Microscope. Cy5-labeled SRV9 particles were excited with a 633 nm helium–neon laser, while the DiO-labeled membrane, GFP-tagged tubulin and eGFP-LCa were all excited with a 488 nm Ar–Kr laser. The fluorescence images were obtained with a NA = 1.40 100×oilimmersion objective. Experiments were conducted at 37 °C unless otherwise mentioned. Data were collected using a photomultiplier tube (PMT) and processed with Leica TCS software. For real-time tracking, we utilized the series scanning imaging of Leica TCS to obtain a sequence of digital images at discrete time points. The serial images are converted to a movie using Image-Pro Plus 6.0. Imaging analysis referring to the motility of viruses is carried out using custom-written MATLAB based on an algorithm described previously47. The fluorescence intensity is analyzed using MATLAB.
Additional Information
How to cite this article: Xu, H. et al. Real-time Imaging of Rabies Virus Entry into Living Vero cells. Sci. Rep. 5, 11753; doi: 10.1038/srep11753 (2015).
References
Sudarshan, M. K. et al. Assessing the burden of human rabies in ndia: results of a national multi-center epidemiological survey. Int. J. Infect. Dis. 11, 29–35 (2007).
Johnson, N., Vos, A., Freuling, C., Tordo, N., Fooks, A. R. & Mueller, T. Human rabies due to lyssavirus infection of bat origin. Vet. Microbiol. 142, 151–159 (2010).
Iwasaki, Y. & Clark, H. F. Cell to cell transmission of virus in central nervous-system .2. experimental rabies in mouse. Lab Invest. 33, 391–399 (1975).
Tordo, N., Poch, O., Ermine, A., Keith, G. & Rougeon, F. Completion of the rabies virus genome sequence determination - highly conserved domains among the l (polymerase) proteins of unsegmented negative-strand rna viruses. Virology 165, 565–576 (1988).
Gaudin, Y., Ruigrok, R. W. H., Tuffereau, C., Knossow, M. & Flamand, A. Rabies virus glycoprotein is a trimer. Virology 187, 627–632 (1992).
Whitt, M. A., Buonocore, L., Prehaud, C. & Rose, J. K. Membrane-fusion activity, oligomerization and assembly of the rabies virus glycoprotein. Virology 185, 681–688 (1991).
Gastka, M., Horvath, J. & Lentz, T. L. Rabies virus binding to the nicotinic acetylcholine receptor alpha subunit demonstrated by virus overlay protein binding assay. J. Gen Virol. 77, 2437–2440 (1996).
Gaudin, Y. et al. Rabies virus-induced membrane fusion. Mol. Membr. Biol. 16, 21–31 (1999).
Mazarakis, N. D. et al. Rabies virus glycoprotein pseudotyping of lentiviral vectors enables retrograde axonal transport and access to the nervous system after peripheral delivery. Hum. Mol. Genet. 10, 2109–2121 (2001).
Lakadamyali, M., Rust, M. J., Babcock, H. P. & Zhuang, X. W. Visualizing infection of individual influenza viruses. Proc. Natl. Acad. Sci. USA 100, 9280–9285 (2003).
Hao, X. et al. Single-Particle Tracking of Hepatitis B Virus-like Vesicle Entry into Cells. small 7, 1212–1218 (2011).
Helenius, A., Kartenbeck, J., Simons, K. & Fries, E. Entry of semliki forest virus into bhk-21-cells. J. Cell Biol. 84, 404–420 (1980).
Zaitseva, E., Yang, S. T., Melikov, K., Pourmal, S. & Chernomordik, L. V. Virus Ensures Its Fusion in Late Endosomes Using Compartment-Specific Lipids PLoS Pathog. 6, e1001131 (2010).
Stoneham, C. A., Hollinshead, M. & Hajitou, A. Clathrin-mediated Endocytosis and Subsequent Endo-Lysosomal Trafficking of Adeno-associated Virus/Phage. J. Biol. Chem. 287, 35849–35859 (2012).
Dohner, K. et al. Function of dynein and dynactin in herpes simplex virus capsid transport. Mol. Biol. Cell. 13, 2795–2809 (2002).
Burckhardt, C. J. et al. Drifting Motions of the Adenovirus Receptor CAR and Immobile Integrins Initiate Virus Uncoating and Membrane Lytic Protein Exposure. Cell. Host. Microbe. 10, 105–117 (2011).
Coller, K. E. et al. RNA Interference and Single Particle Tracking Analysis of Hepatitis C Virus Endocytosis. PLoS Pathog. 5, e1000702 (2009).
Lehmann, M. J., Sherer, N. M., Marks, C. B., Pypaert, M. & Mothes, W. Actin-and myosin-driven movement of viruses along filopodia precedes their entry into cells. J. Cell Biol. 170, 317–325 (2005).
Schelhaas, M. et al. Human papillomavirus type 16 entry:Retrograde cell surface transport along actin-rich protrusions. PLoS Pathog. 4, e1000148 (2008).
Smith, J. L., Campos, S. K. & Ozbun, M. A. Human papillomavirus type 31 uses a caveolin 1-and dynamin 2-mediated entry pathway for infection of human keratinocytes. J. Virol. 81, 9922–9931 (2007).
Greber, U. F. & Way, M. A superhighway to virus infection. Cell 124, 741–754 (2006).
Suomalainen, M. et al. Microtubule-dependent plus- and minus end-directed motilities are competing processes for nuclear targeting of adenovirus. J. Cell Biol. 144, 657–672 (1999).
McDonald, D. et al. Visualization of the intracellular behavior of HIV in living cells. J. Cell Biol. 159, 441–452 (2002).
Gaudin, Y. Rabies virus-induced membrane fusion pathway. J. Cell Biol. 150, 601–611 (2000).
Roche, S. & Gaudin, Y. Evidence that rabies virus forms different kinds of fusion machines with different pH thresholds for fusion. J. Virol. 78, 8746–8752 (2004).
Lewis, P. & Lentz, T. L. Rabies virus entry into cultured rat hippo campalneurons. J. Neurocytol. 27, 559–573 (1998).
Piccinotti, S., Kirchhausen, T. & Whelan, S. P. J. Uptake of Rabies Virus into Epithelial Cells by Clathrin-Mediated Endocytosis Depends upon Actin. J. Virol. 87, 11637–11647 (2013).
Klingen, Y., Conzelmann, K.-K. & Finke, S. Double-labeled rabies virus: live tracking of envelopedvirus transport J. Virol. 82, 237–245 (2008).
Gluska, S. et al. Rabies Virus Hijacks and Accelerates the p75NTR Retrograde Axonal Transport Machinery. PLoS Pathog. 10, e1004348 (2014).
Albertini, A. A. V., Baquero, E., Ferlin, A. & Gaudin, Y. Molecular and Cellular Aspects of Rhabdovirus Entry. Viruses-Basel 4, 117–139 (2012).
Yuan, H. J., Hu, R.l., Zhang, S.f., Zhang, M.l. & Tu, C. c. Characteristics of the glycoprotein cDNA and antigenicity of Rabies virus vaccine strain SRV9 . Virologica Sinica 18, 63–67 (2003).
Rehman, Z., Sjollema, K. A., Kuipers, J., Hoekstra, D. & Zuhorn, I. S. Nonviral gene delivery vectors use syndecan-dependent transport mechanisms in filopodia to reach the cell surface. ACS Nano 6, 7521–7532 (2012).
Burckhardt, C. J. & Greber, U. F. Virus Movements on the Plasma Membrane Support Infection and Transmission between Cells. PLoS Pathog. 5, e1000621 (2009).
Martinez, M. G., Cordo, S. M. & Candurra, N. A. Characterization of Junin arenavirus cell entry. J. Gen. Virol. 88, 1776–1784 (2007).
Wang, L. H. & Anderson, R. G. W. Mis-assembly of clathrin lattices on endosomes reveals a regulatory switch for coated pit formation. J. Cell Biol. 123, 1107–1117 (1993).
Doherty, G. J. & McMahon, H. T. Mechanisms of endocytosis. Annu. Rev. Biochem. 78, 857–902 (2009).
Macia, E. et al. Dynasore, a cell-permeable inhibitor of dynamin. Dev. Cell. 10, 839–850 (2006).
Liu, H., Liu, Y., Liu, S., Pang, D. W. & Xiao, G. Clathrin-mediated endocytosis in living host cells visualized through quantum dot labeling of infectious hematopoietic necrosis virus. J. Virol. 85, 6252–6262 (2011).
Sun, X., Yau, V. K., Briggs, B. J. & Whittaker, G. R. Role of clathrin-mediated endocytosis during vesicular stomatitis virus entry into host cells. Virology 338, 53–60 (2005).
Weir, D. L., Laing, E. D., Smith, I. L., Wang, L. F. & Broder, C. C. Host cell virus entry mediated by Australian bat lyssavirus G envelope glycoprotein occurs through a clathrin-mediated endocytic pathway that requires actin and Rab5. Virol. J 11, 1–10 (2014).
Aleksandrowicz, P. et al. Ebola Virus Enters Host Cells by Macropinocytosis and Clathrin-Mediated Endocytosis. J. Infect. Dis. 204, S957–S967 (2011).
Saeed, M. F., Kolokoltsov, A. A., Albrecht, T. & Davey, R. A. Cellular Entry of Ebola Virus Involves Uptake by a Macropinocytosis-Like Mechanism and Subsequent Trafficking through Early and Late Endosomes. PLoS Pathog 6, e1001110 (2010).
Sieczkarski, S. B. & Whittaker, G. R. Influenza virus can enter and infect cells in the absence of clathrin-mediated endocytosis. J. Virol. 76, 10455–10464 (2002).
Schulz, W. L., Haj, A. K. & Schiff, L. A. Reovirus Uses Multiple Endocytic Pathways for Cell Entry. J. Virol. 86, 12665–12675 (2012).
Razani, B., Woodman, S. E. & Lisanti, M. P. Caveolae: From cell biology to animal physiology. Pharmacol. Rev. 54, 431–467 (2002).
Subtil, A. et al. Acute cholesterol depletion inhibits clathrin-coated pit budding. Proc. Natl. Acad. Sci. USA 96, 6775–6780 (1999).
Brandenburg, B. & Zhuang, X. W. Virus trafficking - learning from single-virus tracking Nat. Rev. Microbiol. 5, 197–208 (2007).
Arhel, N. G. et al. Quantitative four-dimensional tracking of cytoplasmic and nuclear HIV-1 complexes. Nat. Methods. 3, 817–824 (2006).
Sodeik, B. Trends. Mechanisms of viral transport in the cytoplasm. MicrobioL. 8, 465–472 (2000).
Liu, S. L. et al. Globally Visualizing the Microtubule-Dependent Transport Behaviors of Influenza Virus in Live Cells Anal. Chem. 86, 3902–3908 (2014).
Acknowledgements
This work was supported by MOST (Grant no. 2011CB933600 to HW). NSFC (Grant no. 21373200 to HW, 21203177 to XH, 31272579 to MZ and 31330082 to QQ).
Author information
Authors and Affiliations
Contributions
H.J.X., X.H. and Z.Y.W. performed the experiments. S.W.W. performed image analysis. M.J.C. and J.G.J. coordinated the study. H.D.W. designed an initiate the paper. H.J.X., H.D.W., M.L.Z. and Q.W.Q. wrote the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Xu, H., Hao, X., Wang, S. et al. Real-time Imaging of Rabies Virus Entry into Living Vero cells. Sci Rep 5, 11753 (2015). https://doi.org/10.1038/srep11753
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11753
- Springer Nature Limited
This article is cited by
-
Capturing the start point of the virus–cell interaction with high-speed 3D single-virus tracking
Nature Methods (2022)
-
Lipid droplets as endogenous intracellular microlenses
Light: Science & Applications (2021)
-
Early events during human coronavirus OC43 entry to the cell
Scientific Reports (2018)
-
The spread and evolution of rabies virus: conquering new frontiers
Nature Reviews Microbiology (2018)
-
Single-Particle Discrimination of Retroviruses from Extracellular Vesicles by Nanoscale Flow Cytometry
Scientific Reports (2017)