Abstract
Colorectal cancer (CRC) is a major contributor to global morbidity and mortality, necessitating more effective therapeutic approaches. T cells, prominent in the tumor microenvironment, exert a crucial role in modulating immunotherapeutic responses and clinical outcomes in CRC. This study introduces a pioneering method for characterizing the CRC immune microenvironment using single-cell sequencing data. Unlike previous approaches, which focused on individual T-cell signature genes, we utilized overall infiltration levels of colorectal cancer signature T-cells. Through weighted gene co-expression network analysis, Lasso regression, and StepCox analysis, we developed a prognostic risk model, TRGS (T-cell related genes signatures), based on six T cell-related genes. Multivariate Cox analysis identified TRGS as an independent prognostic factor for CRC, showcasing its superior predictive efficacy compared to existing immune-related prognostic models. Immunoreactivity analysis revealed higher Immunophenoscore and lower Tumor Immune Dysfunction and Exclusion scores in the low-risk group, indicating potential responsiveness to immune checkpoint inhibitor therapy. Additionally, patients in the low-risk group demonstrated heightened sensitivity to 5-fluorouracil-based chemotherapy regimens. In summary, TRGS emerges as a standalone prognostic biomarker for CRC, offering insights to optimize patient responses to immunotherapy and chemotherapy, thereby laying the groundwork for personalized tumor management strategies.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) stands as the third most prevalent malignancy globally, ranking second in mortality1,2, thereby posing substantial therapeutic challenges and an economic burden. Despite advancements in early CRC screening leading to improved 5 years survival rates, a subset of patients still presents at stage IV, resulting in a dismal prognosis3. Immune checkpoint inhibitors (ICIs) have demonstrated remarkable therapeutic efficacy in diverse solid tumors, establishing them as the primary treatment for microsatellite instability-high (MSI-H) metastatic CRC4. However, given that this patient subset constitutes less than 5% of all CRC cases5, there is a critical need to identify more efficient biomarkers to guide the application of ICIs in colorectal cancer.
The tumor microenvironment (TME) plays an important role in tumors, mainly in terms of tumor progression, angiogenesis, tumor metastasis, and influencing treatment outcome6,7,8,9. TME encompasses diverse cellular components such as immune cells, stromal cells, and non-cellular components including extracellular matrix molecules (ECM) and soluble signaling molecules10. Among these, T cells, including CD4+ and CD8+ T cells, represent the primary immune cells exerting anti-tumor activity11. After antigen-presenting cells present tumor cell antigens to T cells, activating them, the activated T cells infiltrate the tumor vicinity to exert cytotoxic effects on tumor cells12. However, tumor cells can evade immune surveillance through various mechanisms, such as HLA-I loss13, expression of immune checkpoint molecules14, or inhibitory effects from other cells in the microenvironment15,16. Although more studies are focusing on a specific single T cell, few studies have been reported on overall T cells infiltration. A comprehensive understanding of the alterations in T cells within the tumor microenvironment is crucial for devising future personalized therapeutic strategies.
This study utilized bioinformatics methodologies to integrate single-cell RNA sequencing (scRNA-seq) data and transcriptome sequencing data from CRC samples. We performed unsupervised consensus clustering of CRC patients based on the overall infiltration levels of characteristic T cells in the tumor microenvironment instead of focusing on a single T cell’s signature genes. Through this approach, a novel T-cell-related prognostic risk model, termed TRGS, was developed and subsequently validated. This model holds significant promise for elucidating the intricate role of T cells within the tumor immune microenvironment of CRC. Notably, when compared with existing prognostic models, TRGS demonstrates superior efficacy in prognostic prediction. Moreover, TRGS exhibits the capacity to predict treatment outcomes, encompassing responses to both immunotherapy and chemotherapy among CRC patients. Additionally, it may identify novel therapeutic targets to improve the effectiveness of ICIs.
Methods
Data download and processing
For the TCGA-CRC cohort, we retrieved normalized RNA sequencing data (expressed as transcripts per million, TPM) for 549 colorectal cancer (CRC) patients from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). We specifically included patients with a survival time of ≥ 30 days who did not undergo neoadjuvant therapy. The normalized matrix files for the GSE39582 and GSE38832 cohorts from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/) were utilized as independent external cohorts for the validation of our risk model. Similarly, normalized matrix files for the GSE28702, GSE19860, and GSE45404 cohorts from the GEO database served as independent external cohorts for predicting chemotherapy outcomes. Additionally, single-cell sequencing data were obtained from the GSE132465 dataset, including 23 primary colorectal cancer and 10 matched normal mucosa samples.
scRNA-Seq data analysis
The “Seurat” R package (v4.4.0)17 was used to perform unsupervised clustering of single cells using the read count matrix as input. We annotated the cells according to the original annotation18. Cell populations originally annotated as T-cells were extracted and normalized using the SCTransform method19 with resolution choice 1.2 for re-clustering. To find marker genes in various clusters, the “Find All Markers” tool was used with |Log2FC| and min. pct cutoff values set to 0.25 and 0.25, respectively.
Identification of immune class by consensus clustering
The “ConsensusClusterPlus” R package20 was used to perform the unsupervised consensus clustering analysis. We used 80% of the items for subsampling and divided each subsample into groups by the k-means algorithm, repeating this clustering process 1000 times. Then, the cumulative distribution function (CDF) curve and consensus matrix are used to identify the optimal k-value. Based on the optimal k-value, patients were clustered into sub-clusters.
Tumor immune microenvironment evaluation
The MCP-counter algorithm21 based on the “MCPcounter” R package was used to assess the 26 kinds of T cell clusters identified from the scRNA-seq analysis. The ESTIMATE algorithm22 tool was used to calculate stromal, immune, and estimated scores. Single sample gene set enrichment analysis (ssGSEA)23 based on the “GSVA” R package was used to quantify 28 kinds of immune cell infiltration scores of the TCGA cohort.
Weighted correlation network analysis (WGCNA)
Coexpression protein-coding RNA networks of TCGA-CRC were generated using the WGCNA package24. An appropriate soft threshold β was calculated to meet the criteria for the scale-free network. Then, the adjacency was transformed into a topological overlap matrix (TOM), and genes with similar expression patterns were divided into the same module. Finally, genes from modules that had high correlation coefficients with T-cell relative immune status were identified for subsequent analysis.
Screening and functional annotation of T cell-related genes
DEGs were screened by the limma R package. Adj. P < 0.05 and |log2FC|> 0.6 were set as the selection conditions for screening the downregulated and upregulated DEGs. T cell-related genes were obtained by overlapping DEGs and WGCNA. Enrichment analysis in Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG)25,26,27 were based on the “Clusterprofiler” R package28, where “adjusted p < 0.05” was considered significant.
Construction and validation of T cell‑related genes prognosis signature (TRGS)
The TRGS was developed based on the TCGA-CRC cohort. First, univariate Cox regression analysis was utilized to screen T cell-related genes that were associated with OS using the “survival” R package (P < 0.05). Next, we performed machine learning algorithms including the least absolute shrinkage and selection operator (LASSO) regression analysis and stepwise Cox algorithm to improve the accuracy and reliability of prognostic signature using the “glmnet” and “StepReg” R packages, respectively. The prognostic signature was established as follows: TRGS = \(\sum\nolimits_{{{\text{i}} = 1}}^{{\text{n}}} {({\text{Coef}} \times {\text{Ni}})}\), where Coef referred to the coefficient of gene i and Ni represented the expression value of gene i.
To further validate the predictive power of the prognostic signature, we performed Kaplan–Meier survival analysis in the TCGA-CRC, GSE39582, and GSE38832 cohorts, where the cutoff value was set to the median TRGS. Time-dependent receiver operating characteristic curves (ROC) were plotted using the “timeROC” R package to predict OS at 1, 3, and 5 years in the training dataset (TCGA-CRC); 1, 2, and 3 years in GSE38832 cohort, and 3, 7, 10 years in GSE39582 cohort. Finally, we further analyzed the relationship between the prognostic signature and clinical characteristics (including age, gender, and stage) to clarify the independent prognostic value of TRGS using multivariate Cox regression analyses.
Development and evaluation of the nomogram
We performed the univariate and multivariate Cox regression analyses for clinical parameters and TRGS. In the multivariate Cox model, variables with P < 0.05 were included in the construction of the nomogram by using the “rms” R package. The calibration analysis was used to assess the prognostic accuracy of the nomogram model. Decision curve analysis (DCA) curves were used to evaluate the net benefit of the nomogram.
Immune checkpoints inhibitors sensitivity prediction
We utilized Immunophenoscore (IPS) obtained from The Cancer Immunome Atlas (TCIA) website (tcia.at/home)29 to predict the response to ICIs. We also utilized the Tumor Immune Dysfunction and Exclusion (TIDE) algorithm through an online website (http://tide.dfci.harvard.edu)30 to predict patients’ ICIs responses. Higher TIDE prediction scores were generally associated with worse ICIs responses.
Anticancer drug sensitivity prediction
The half-maximal inhibitory concentration (IC50) of common chemotherapeutic and targeted therapeutic drugs was estimated by using the “pRRophetic” R package31. GSE28702, GSE19860, and GSE45404 were transcriptome sequencing data from patients who received adjuvant chemotherapy (ACT), which were utilized as external validation cohorts for predicting chemotherapy drug sensitivity.
Statistical analysis
All statistical analyses were performed using R software (version 4.3.1, R: The R Project for Statistical Computing (r-project.org)). Wilcoxon test was used to compare the differences between groups. The log-rank test was used to compare Kaplan–Meier survival curves. Univariate and multivariate Cox analyses were performed to establish independent prognostic factors. All P values were two-sided and less than 0.05 were considered statistically significant.
Results
ScRNA-Seq analysis of CRC samples
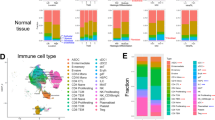
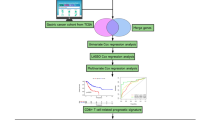
The flowchart outlining our study is presented in Supplementary Fig. 1. We obtained 10 × scRNA-seq data from the GSE132465 dataset, which comprised 33 CRC samples, including 23 primary colorectal cancer and 10 matched normal mucosa samples. The 23,115 T cells extracted from the original dataset were re-clustered. Based on the clustree results (Supplementary Fig. 2A), a resolution of 1.2 was chosen, leading to the identification of 26 distinct cell subgroups through hierarchical clustering utilizing UMAP analysis (Fig. 1A). Annotation of the 26 clusters of T cells was performed using literature reports32,33, CellMarker 2.034, and T cellmap35. We presented the characteristic genes for each cell cluster (Fig. 1B). Meanwhile, we have referenced the previous publication35 and utilized functional gene sets to validate the annotations' appropriateness (Fig. 1C,D). Differential expression genes (DEGs) of each cluster were provided in Supplementary Table S1 and shown in Supplementary Fig. 2B. Furthermore, we explored the differences between these cells in cancer and adjacent tissues (Fig. 1E). Subsequently, the characteristic genes of each cell cluster were employed to estimate the differences in cell populations of TCGA patients using the MCP-counter algorithm (Fig. 1F). This result illustrated that the T-cell infiltration levels predicted by the MCP-counter algorithm for TCGA are consistent with the infiltration levels in the single-cell database. This serves to substantiate the rationality of our choice of the MCP-counter algorithm.
Different T cell clustering in 10X scRNA-seq data of colorectal carcinoma. (A) Cluster annotation and cell type identification using UMAP. (B) Bubble plot displaying marker gene expression across identified clusters. (C, D) Heatmap depicting the expression of curated gene signatures within CD8+ T and CD4+ T cell clusters. (E) Distribution of defined T cell clusters between tumor and normal groups in GSE132465. (F) MCP-counter enrichment scores of defined T cell clusters between normal and tumor groups in the TCGA-CRC cohort.
Identification of T cell infiltration subgroups
Based on the assessment of T cell infiltration using the MCP-counter package, we conducted a consensus cluster analysis, initially dividing all CRC samples into k clusters (k = 2–9). The cumulative distribution function (CDF) curves of the consensus score matrix revealed that the optimal number of clusters was achieved when k = 2 (Fig. 2A,B). The two consensus clusters (C1 and C2) exhibited significant differences in the distribution of 26 T cell clusters, with C1 displaying substantially higher T cell infiltration abundance compared to C2 (Fig. 2C). Consequently, we categorized C1 as “immune-hot” tumors and C2 as “immune-cold” tumors. To ensure the robustness of the two consensus clusters and mitigate potential algorithmic biases, ssGSEA was employed to validate the disparity in immune cell infiltration between C1 and C2, revealing a higher overall immune cell infiltration abundance in C1 (Fig. 2D). Additionally, the ESTIMATE algorithm was utilized to illustrate that immune-hot tumors exhibited a higher immune score compared to immune-cold tumors (Fig. 2E).
Unsupervised consensus cluster analysis of CRC patients in the TCGA cohort based on MCP-counter enrichment score. (A) The consensus score matrix of all samples when k = 2. (B) The CDF curves of the consensus matrix for each k (indicated by colors). (C) The infiltration abundance of defined T cell clusters for two clusters. (D) The infiltration abundance of 28 immune cell subsets evaluated by ssGSEA for two clusters. (E) The distribution of immune score inferred by ESTIMATE algorithm between two clusters in the TCGA-CRC cohort. Statistic test: two-sided unpaired t-test. (*P < 0.05, ** P < 0.01, *** P < 0.001).
Identification of candidate T cell-related genes
To identify T cell-related genes associated with immune cell infiltration, we initially conducted weighted gene co-expression network analysis (WGCNA). In this procedure, we set the soft threshold β to 6, resulting in a scale-free topology fit index of 0.85 (Supplementary Fig. 3A). Subsequently, we utilized the “merged dynamics” algorithm to derive fourteen modules (Fig. 3A). The heatmap depicted the eigengene adjacency of these modules (Fig. 3B). Through assessment of the correlation coefficient and P value, we observed that the blue module exhibited the strongest correlation with the previously identified immune status (Fig. 3C), achieving a correlation coefficient of 0.78 between gene significance (GS) and module membership (MM) (Fig. 3D). Concurrently, we conducted differential expression analysis of the TCGA-CRC cohort using the “limma” package, identifying a total of 686 differentially expressed genes (DEGs). By intersecting these results with those from WGCNA, we extracted 369 overlapping genes deemed as T cell-related genes for subsequent analyses (Fig. 3E). GO and KEGG enrichment analyses were conducted to provide a comprehensive functional overview of the overlapping genes (Fig. 3F,G and Supplementary Table S2). The results revealed predominant enrichment in biological processes (BP) such as leukocyte-mediated immunity, leukocyte cell–cell adhesion, and regulation of T-cell activation. In terms of cellular composition (CC), enrichment was observed in areas including the external side of the plasma membrane, MHC protein complex, and MHC class II protein complex. Additionally, molecular function (MF) analysis highlighted immune receptor activity, chemokine receptor binding, and MHC protein complex binding as significantly enriched categories. Furthermore, KEGG pathway analysis indicated significant enrichment in pathways such as cytokine-cytokine receptor interaction, cell adhesion molecules, and chemokine signaling, all of which were closely associated with anti-cancer immunity.
Identification of T cell-related genes. (A) The dendrogram of modules. (B) The heatmap revealed the eigengene adjacency of modules. (C) Correlation analysis between module eigengenes and clinical traits. (D) The strong correlation between GS and MM in the blue module. (E) Venn diagram displaying overlapping genes between WGCNA and TRDEGs. (F, G) Go and KEGG function enrichment of the intersected genes.
T cell signature establishment and external validation
To evaluate the prognostic impact of the identified genes on CRC, we initially identified 12 genes associated with overall survival (OS) through univariate Cox analysis (Supplementary Fig. 3B). Subsequently, to prevent overfitting and exclude co-expressed T cell-related genes (Supplementary Fig. 3C), we employed lasso regression analysis to construct a predictive prognostic model comprising T cell-related genes. The optimal λ value was determined based on the minimum partial likelihood deviance achieved via the leave-one-out cross-validation (LOOCV) framework (Fig. 4A). Ten genes with nonzero Lasso coefficients were further subjected to stepwise Cox proportional hazards regression, resulting in the identification of a final set of 6 genes (Fig. 4B), and we assessed their impact on prognosis at the same time (Supplementary Fig. 3D–I).
The establishment and external validation of the T cell-related genes signature. (A) Lasso regression profiles of TRGS to prevent over-fitting and tenfold cross-validation of variable selection with Lasso. (B) Coefficients of 6 T cell-related genes finally obtained in stepwise Cox regression. (C, E, G) Kaplan–Meier curves illustrating the prognostic value of TRGS. (D, F, H) Time-dependent ROC curves demonstrating the prognostic signature's predictive performance. (I) Nomogram integrating age, stage, and TRGS for prognosis assessment. (J) The calibration curve of the constructed nomogram of 1-year, 3-years, and 5-years survival prediction. (K) Decision curve analysis evaluating the 5-years survival benefit in the TCGA cohort. (L) Univariate and multivariate COX regression analysis of TRGS and various clinical features in TCGA cohort.
A linear prediction model was then developed using the weighted regression coefficients of the 6 T cell-related genes, represented as TRGS = (0.325 × ADAM8 exp) + (0.282 × ZNF385A exp) + (0.201 × G0S2 exp) + (− 0.195 × TAP1 exp) + (− 0.216 × MMP1 exp) + (− 0.347 × CCL22 exp). Based on the median of TRGS, all patients were stratified into high- and low-risk groups. Patients in the high-risk group had a higher tumor stage (Table 1) and, notably, exhibited significantly poorer overall survival (OS) compared to those in the low-risk group (Fig. 4C,E,G). The discriminatory ability of TRGS was assessed using receiver operating characteristic (ROC) analysis, with 1-, 3-, and 5-years areas under the curve (AUCs) of 0.725, 0.710, and 0.700 in TCGA-CRC (Fig. 4D); 3-, 7-, and 10-years AUCs of 0.620, 0.639, and 0.684 in the GSE39582 cohort (Fig. 4F); and 1-, 2-, 3-years AUCs of 0.656, 0.696, and 0.616 in GSE38832 cohort (Fig. 4H). Furthermore, to evaluate the prognostic capabilities of TRGS, we obtained 5 previously published prognostic models developed using immune-related genes36,37,38,39,40. Subsequently, we conducted a comparative analysis to assess the predictive performance of TRGS in terms of prognosis. Remarkably, our findings revealed that TRGS exhibited superior performance, as evidenced by higher AUC values in time-dependent ROC analysis when compared to the 5 existing prognostic models in TCGA, GSE39582, and GSE38832 cohorts. (Supplementary Fig. 4A–O).
Finally, we developed a nomogram incorporating TRGS and clinicopathological features to assess the clinical significance of the predictive model (Fig. 4I). Calibration curves demonstrated good agreement between the predicted and actual probabilities of survival at 1, 3, and 5 years (Fig. 4J). Decision curve analysis (DCA) revealed that the nomogram model provided a greater net benefit in predicting 5-years OS (Fig. 4K). Meanwhile, multifactorial Cox regression analysis confirmed TRGS as an independent prognostic risk factor for CRC (Fig. 4L and Supplementary Fig. 4P).
TRGS evaluation for immunotherapeutic response prediction
To evaluate the predictive capacity of TRGS as a biomarker for ICIs response, we initially investigated the relationship between low- and high-risk groups and the immune-hot and immune-cold phenotypes as defined in Fig. 2. Patients classified in the immune-cold group belonged to the high-risk category, whereas those in the low-risk group exhibited an immune-hot phenotype (Fig. 5A). Subsequently, we examined the association between TRGS and immune checkpoint blocker (ICB)-related pathways, as well as the seven steps in the Cancer-Immunity cycle. Our findings revealed a negative correlation between TRGS and ICB-related pathways and steps in the Cancer-Immunity cycle, suggesting that low-risk patients may possess a richer tumor immune microenvironment, potentially leading to a higher response rate to ICIs (Fig. 5B,C). Furthermore, we analyzed the distribution of IPS and TIDE scores across different risk groups. Our results indicated that the low-risk group exhibited higher IPS scores (Fig. 5D–G), whereas the high-risk group demonstrated elevated TIDE scores (Fig. 5H), Exclusion scores (Fig. 5I), and Dysfunction scores (Fig. 5J). These results were consistent with previous findings, suggesting that low-risk patients may exhibit increased sensitivity to ICIs.
TRGS correlation with immune phenotype and ICIs response predictors in CRC. (A) Alluvial plot depicting the distribution of Immune hot and Immune cold phenotypes across different risk subtypes and survival statuses. (B, C) Correlation analysis of TRGS with immune checkpoint blockade (ICB) related pathways (B) and individual steps of the cancer-immune cycle (C). (D–J) Distribution of IPS, IPS_PD1_blocker, IPS_CTLA4_blocker, and IPS_CTLA4_PD1_blocker. (H–I) Distribution of TIDE, T cell exclusion score, and T cell dysfunction score. (*P < 0.05, ** P < 0.01, *** P < 0.001).
Predictive value of fluorouracil-based ACT benefits
We expanded our investigation to assess the predictive capability of TRGS in quantifying the benefits of fluorouracil-based adjuvant chemotherapy (ACT). Survival analysis among patients stratified into low- and high-risk groups, both receiving and not receiving ACT, revealed that individuals in the low-risk category who underwent therapy exhibited the most favorable survival outcomes. Conversely, among patients categorized as high-risk, no substantial disparity in survival was observed between those who received ACT and those who did not (Fig. 6A,B). This suggests that low TRGS may serve as a predictor for a more favorable response to ACT.
TRGS predicts the sensitivity of fluorouracil-based ACT. (A, B) Kaplan–Meier curves comparing patients who received ACT with those who did not in TCGA (A) and GSE39582 (B) cohorts. (C, D) The predicted IC50 values of chemotherapy (C) and targeted therapy (D) drugs in different risk groups in the TCGA cohort. (E–H) Distribution of TRGS between responders and non-responders of fluorouracil-based ACT in TCGA (E), GSE28702 (F), GSE19860 (G), and GSE45404 (H) cohorts. (*P < 0.05, ** P < 0.01, *** P < 0.001).
Subsequently, we calculated the IC50 values of common chemotherapy and targeted drugs for CRC to further explore the drug sensitivity between low- and high-risk groups. Comparison of the IC50 values between the two risk groups indicated that the low-risk group exhibited higher sensitivity to fluorouracil, gemcitabine, paclitaxel, and vincristine (Fig. 6C). However, patients in the high and low-risk groups showed no significant difference in the sensitivity to targeted therapy (Fig. 6D). These findings were validated in three datasets treated with fluorouracil-based ACT. Responders demonstrated a significantly lower TRGS score than non-responders in TCGA, GSE28702, GSE19860, and GSE45404 (Fig. 6E–H).
Discussion
Currently, conventional treatment strategies for colorectal cancer primarily involve surgery combined with adjuvant chemotherapy or targeted therapy, yet these approaches have not yielded significant efficacy41. Moreover, stage IV patients exhibit a high likelihood of chemotherapy resistance30, potentially leading to situations of overtreatment or undertreatment within the current staging-based treatment paradigm. Finding biomarkers to predict chemotherapy outcomes is of utmost urgency. Furthermore, immunotherapy, as a novel and effective treatment modality, remains underutilized in colorectal cancer, necessitating more effective biomarkers to guide the application of ICIs in colorectal cancer42. T cells, being the most abundant immune cells infiltrating the tumor microenvironment, play a crucial role in anti-tumor immunity. A comprehensive understanding of the T cell population's status within the colorectal cancer tumor microenvironment contributes to the identification of more effective immunotherapeutic strategies.
Our study utilized scRNA-seq data to identify a T cell signature in CRC, uncovering genes that indicate T cell presence. Through WGCNA, we classified CRC into immune subtypes based on T cell infiltration, highlighting genes linked to the immunogenic phenotype. Using COX regression, we selected key genes associated with prognosis and constructed a prognostic risk model with machine learning algorithms. Among these six genes, high expression of ADAM8, ZNF385A, and G0S2 was associated with poor prognosis, while high expression of TAP1, MMP1, and CCL22 was associated with favorable prognosis.
Interestingly, these six genes have previously been found to be associated with immune cell infiltration in earlier studies, consistent with our findings. Studies have reported that downregulating ADAM8 in colorectal cancer cells enhances the cytotoxic effect of tumor-infiltrating lymphocytes43. ZNF385A has been identified as a gene associated with lymphatic vessel formation in CRC, influencing immune cell infiltration, and is also associated with an immunosuppressive phenotype in HCC44,45. G0S2 is linked to evasion from cytotoxic T lymphocytes and knocking down G0S2 significantly inhibits proliferation and migration of CRC cell lines46. TAP1, a transporter associated with antigen processing protein, plays a crucial role in antigen presentation47, and its downregulation in colorectal cancer is associated with low infiltration of various lymphocytes48. MMP1, one of the markers of epithelial-mesenchymal transition, is associated with metastasis and angiogenesis in colorectal cancer49,50. In refractory HCC treated with Transcatheter arterial chemoembolization (TACE), a prognostic model incorporating MMP1 is related to immune cell infiltration51. Additionally, MMP1 may be one of the targets to enhance the effectiveness of immunotherapy in PDAC52. CCL22 is a chemokine that not only recruits monocytes but also regulates the infiltration of Th1 and Treg cells in colorectal cancer53,54.
Subsequently, we developed a T cell-related gene signature (TRGS) using these six genes and stratified patients into high- and low-risk groups. The TRGS proved to be a strong independent prognostic factor, with the low-risk group showing significantly improved overall survival. Furthermore, the TRGS outperformed existing immune-related models in predictive accuracy. The low-risk group showed a higher IPS, suggesting a better response to ICIs, while the high-risk group exhibited signs of immune evasion. These findings indicate that ICIs may be more efficacious for patients in the low-risk category.
Previous studies have shown that 5-FU treatment can induce a reduction in tumor-infiltrating myeloid-derived suppressor cells (MDSCs) and promote T cell-mediated anti-tumor immunity55. Additionally, in metastatic colorectal cancer patients receiving FOLFOX combined with bevacizumab, the accumulation of Th17 cells was associated with poor prognosis56. Similarly, in lung cancer, the combination treatment of oxaliplatin and cyclophosphamide led to an increased ratio of CD8 T cells to Treg cells within the tumor, resulting in significant inhibition of tumor growth57. These findings collectively indicate the existence of interactions between T cells and chemotherapy within the tumor microenvironment. Therefore, our study compared the sensitivity of high- and low-risk groups to common chemotherapy drugs, revealing that the low-risk group had a higher sensitivity to these agents. Validation using external datasets supported our findings, showing that patients responsive to fluorouracil-based chemotherapy had lower risk scores.
While our study has been successfully validated in external cohorts, several limitations warrant consideration. Firstly, our study relied on publicly available databases, and both our training cohort (TCGA) and validation cohorts (GEO) were retrospective. Hence, prospective studies are warranted to validate our conclusions rigorously. Secondly, there is still a shortage of sequencing databases containing data from CRC patients treated with ICIs, as well as a deficiency of large clinical cohorts of patients undergoing ICIs treatment. Therefore, our results still need validation in a large clinical cohort in the future. Thirdly, further research is needed to identify hub genes and elucidate the potential regulatory mechanisms to facilitate the discovery of targets that enhance the efficacy of ICIs and chemotherapy.
Conclusion
In summary, we pioneered a novel approach to categorize the colorectal cancer immune microenvironment utilizing single-cell data. Rather than focusing on individual T-cell signature genes, we employed the infiltration levels of the comprehensive colorectal cancer signature T-cells. This innovative strategy led to the construction of a TRGS comprising ADAM8, ZNF385A, G0S2, TAP1, MMP1, and CCL22. Our findings underscore the superiority of TRGS in prognostic prediction compared to existing immune-related prognostic models. Moreover, TRGS exhibits predictive capabilities not only for ICIs treatment but also for chemotherapy efficacy, presenting novel perspectives for the stratified diagnosis and treatment of colorectal cancer patients.
Data availability
The raw data utilized in our study were obtained from the TCGA dataset (https://portal.gdc.cancer.gov/) and the GEO dataset (https://www.ncbi.nlm.nih.gov/). Specifically, TCGA accession numbers for our study are TCGA-COAD and TCGA-READ, while GEO accession numbers are GSE39582, GSE38832, GSE132465, GSE28702, GSE19860, and GSE45404. The datasets analyzed and R language codes employed in the current study are accessible upon reasonable request from the corresponding author.
References
Xi, Y. & Xu, P. Global colorectal cancer burden in 2020 and projections to 2040. Transl. Oncol. 14, 101174 (2021).
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Siegel, R. L. et al. Colorectal cancer statistics, 2020. CA Cancer J Clin 70, 145–164 (2020).
Andre, T. et al. Pembrolizumab in microsatellite-instability-high advanced colorectal cancer. N. Engl. J. Med. 383, 2207–2218 (2020).
Koopman, M. et al. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br. J. Cancer 100, 266–273 (2009).
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Angelova, M. et al. Evolution of metastases in space and time under immune selection. Cell 175, 751–765 (2018).
Jin, M. Z. & Jin, W. L. The updated landscape of tumor microenvironment and drug repurposing. Signal Transduct. Target Ther. 5, 166 (2020).
Hernandez-Camarero, P., Lopez-Ruiz, E., Marchal, J. A. & Peran, M. Cancer: A mirrored room between tumor bulk and tumor microenvironment. J. Exp. Clin. Cancer Res. 40, 217 (2021).
Belli, C. et al. Targeting cellular components of the tumor microenvironment in solid malignancies. Cancers (Basel) 14, 4278 (2022).
Pelka, K. et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 184, 4734–4752 (2021).
Mellman, I., Chen, D. S., Powles, T. & Turley, S. J. The cancer-immunity cycle: Indication, genotype, and immunotype. Immunity 56, 2188–2205 (2023).
Anderson, P., Aptsiauri, N., Ruiz-Cabello, F. & Garrido, F. HLA class I loss in colorectal cancer: Implications for immune escape and immunotherapy. Cell Mol. Immunol. 18, 556–565 (2021).
Chen, Q. et al. Demethylase JMJD2D induces PD-L1 expression to promote colorectal cancer immune escape by enhancing IFNGR1-STAT3-IRF1 signaling. Oncogene 41, 1421–1433 (2022).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Fridman, W. H. et al. B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nat. Rev. Clin. Oncol. 19, 441–457 (2022).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Lee, H. O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Lause, J., Berens, P. & Kobak, D. Analytic Pearson residuals for normalization of single-cell RNA-seq UMI data. Genome Biol. 22, 258 (2021).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: A class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Kanehisa, M. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Charoentong, P. et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 18, 248–262 (2017).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Geeleher, P., Cox, N. & Huang, R. S. pRRophetic: An R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One 9, e107468 (2014).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Hu, C. et al. Cell Marker 2.0: An updated database of manually curated cell markers in human/mouse and web tools based on scRNA-seq data. Nucleic Acids Res. 51, D870–D876 (2023).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Tu, W. et al. Elucidating the role of T-cell exhaustion-related genes in colorectal cancer: A single-cell bioinformatics perspective. Funct. Integr. Genomics 23, 259 (2023).
Chen, J. et al. Comprehensive analysis and experimental verification of the mechanism of action of T cell-mediated tumor-killing related genes in Colon adenocarcinoma. Transl. Oncol. 43, 101918 (2024).
Liao, K. et al. Identification of signature of tumor-infiltrating CD8 T lymphocytes in prognosis and immunotherapy of colon cancer by machine learning. Clin. Immunol. 257, 109811 (2023).
Li, Q., Chu, Y., Yao, Y. & Song, Q. A Treg-related riskscore model may improve the prognosis evaluation of colorectal cancer. J. Gene Med. 26, e3668 (2024).
Li, X. et al. Identification of an immune signature predicting prognosis risk and lymphocyte infiltration in colon cancer. Front. Immunol. 11, 1678 (2020).
Bockelman, C., Engelmann, B. E., Kaprio, T., Hansen, T. F. & Glimelius, B. Risk of recurrence in patients with colon cancer stage II and III: A systematic review and meta-analysis of recent literature. Acta Oncol. 54, 5–16 (2015).
Ganesh, K. et al. Immunotherapy in colorectal cancer: Rationale, challenges and potential. Nat. Rev. Gastroenterol. Hepatol. 16, 361–375 (2019).
Liao, R. et al. Identification of biomarkers related to Tumor-Infiltrating Lymphocytes (TILs) infiltration with gene co-expression network in colorectal cancer. Bioengineered 12, 1676–1688 (2021).
Liu, H., Shi, H. & Sun, Y. Identification of a novel lymphangiogenesis signature associated with immune cell infiltration in colorectal cancer based on bioinformatics analysis. BMC Med. Genomics 17, 2 (2024).
Peng, Q. et al. ZNF385A and ZNF346 serve as prognostic biomarkers associated with an inflamed immunosuppressive tumor microenvironment in hepatocellular carcinoma. Int. J. Mol. Sci. 24, 3155 (2023).
Wang, X. et al. Robust machine-learning based prognostic index using cytotoxic T lymphocyte evasion genes highlights potential therapeutic targets in colorectal cancer. Cancer Cell Int. 24, 52 (2024).
Blees, A. et al. Structure of the human MHC-I peptide-loading complex. Nature 551, 525–528 (2017).
Ling, A. et al. TAP1 down-regulation elicits immune escape and poor prognosis in colorectal cancer. Oncoimmunology 6, e1356143 (2017).
Yang, P. et al. TCONS_00012883 promotes proliferation and metastasis via DDX3/YY1/MMP1/PI3K-AKT axis in colorectal cancer. Clin. Transl. Med. 10, e211 (2020).
Ng, L. et al. CD26 induces colorectal cancer angiogenesis and metastasis through CAV1/MMP1 signaling. Int J Mol Sci 23, 1181 (2022).
Cheng, X. et al. The role of hypoxia-related genes in TACE-refractory hepatocellular carcinoma: Exploration of prognosis, immunological characteristics and drug resistance based on onco-multi-OMICS approach. Front. Pharmacol. 13, 1011033 (2022).
Hung, H. C. et al. Effect of chimeric antigen receptor T cells against protease-activated receptor 1 for treating pancreatic cancer. BMC Med. 21, 338 (2023).
Cremonesi, E. et al. Gut microbiota modulate T cell trafficking into human colorectal cancer. Gut 67, 1984–1994 (2018).
Dong, S., Guo, X., Han, F., He, Z. & Wang, Y. Emerging role of natural products in cancer immunotherapy. Acta Pharm. Sin. B 12, 1163–1185 (2022).
Vincent, J. et al. 5-Fluorouracil selectively kills tumor-associated myeloid-derived suppressor cells resulting in enhanced T cell-dependent antitumor immunity. Cancer Res. 70, 3052–3061 (2010).
Limagne, E. et al. Accumulation of MDSC and Th17 Cells in patients with metastatic colorectal cancer predicts the efficacy of a FOLFOX-bevacizumab drug treatment regimen. Cancer Res. 76, 5241–5252 (2016).
Pfirschke, C. et al. Immunogenic chemotherapy sensitizes tumors to checkpoint blockade therapy. Immunity 44, 343–354 (2016).
Acknowledgements
The authors thank researchers and contributors of the TCGA and GEO databases for their valuable efforts and contributions.
Author information
Authors and Affiliations
Contributions
P.C. and Z.G.B. conceptualized and designed the study. P.C. was responsible for data collection, analysis, and initial manuscript drafting. H.B.W. reviewed the statistical analysis and performed manuscript editing. Z.G.B. oversaw the project, contributed to manuscript editing, provided critical feedback, and supervised the study. All authors contributed to the critical revision of the manuscript and approved the final version for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cui, P., Wang, H. & Bai, Z. Integrated single-cell and bulk RNA-seq analysis identifies a prognostic T-cell signature in colorectal cancer. Sci Rep 14, 20177 (2024). https://doi.org/10.1038/s41598-024-70422-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70422-6
- Springer Nature Limited