Abstract
The present study aimed to elucidate the prognostic mutation signature (PMS) associated with long-term survival in a diffuse large B-cell lymphoma (DLBCL) cohort. All data including derivation and validation cohorts were retrospectively retrieved from The Cancer Genome Atlas (TCGA) database and whole-exome sequencing (WES) data. The Lasso Cox regression analysis was used to construct the PMS based on WES data, and the PMS was determined using the area under the receiver operating curve (AUC). The predictive performance of eligible PMS was analyzed by time-dependent receiver operating curve (ROC) analyses. After the initial evaluation, a PMS composed of 94 PFS-related genes was constructed. Notably, this constructed PMS accurately predicted the 12-, 36-, and 60-month PFS, with AUC values of 0.982, 0.983, and 0.987, respectively. A higher level of PMS was closely linked to a significantly worse PFS, regardless of the molecular subtype. Further evaluation by forest plot revealed incorporation of international prognostic index or tumor mutational burden into PMS increased the prediction capability for PFS. The drug-gene interaction and pathway exploration revealed the PFS-related genes were associated with DNA damage, TP53, apoptosis, and immune cell functions. In conclusion, this study utilizing a high throughput genetic approach demonstrated that the PMS could serve as a prognostic predictor in DLBCL patients. Furthermore, the identification of the key signaling pathways for disease progression also provides information for further investigation to gain more insight into novel drug-resistant mechanisms.
Similar content being viewed by others
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common aggressive non-Hodgkin lymphoma (NHL) worldwide, accounting for approximately 30–40% of annual newly diagnosed lymphoma cases1,2. The incorporation of the anti-CD20 monoclonal antibody, rituximab (R), into conventional anthracycline-based chemotherapy results in a high response rate and prolonged overall survival of patients with DLBCL3,4,5. For patients with newly diagnosed DLBCL, the standard R-CHOP (cyclophosphamide, doxorubicin, vincristine, and prednisone) regimen achieves a high complete remission rate (approximately 75%), and 60–70% of these patients remain relapse-free after 5 years of follow-up6,7. However, a proportion of DLBCL patients experience primary refractory disease or relapse after prior successful treatment. The prognosis of this subgroup is dismal, making the exploration of resistance mechanisms or new therapies an urgent medical need.
Accumulating evidence suggests that highly heterogeneous genetic alterations and the tumor microenvironment play crucial roles in treatment failure8. The gene expression profiling divides DLBCL into two distinct groups, namely, germinal center B-cell-like (GCB) and activated B-cell-like (ABC). Patients with ABC-DLBCL tend to have a poorer prognosis and a higher risk of treatment resistance9,10,11. In addition, a small subset of patients with MYC, BCL2, and/or BCL6 arrangement has been classified as double-hit lymphoma (DHL) or triple-hit lymphoma (THL), these patients with DHL/THL tend to respond poorly to R-CHOP regimen12,13,14. Advances in next-generation sequencing (NGS) technology and bioinformatics allow integrative genomic analyses in a large cohort of patients, enabling the identification of novel genetic subsets and modeling of novel genetic classifications15,16,17. Several genetic alterations related to relapsed or refractory DLBCL after R-CHOP treatment were identified, including epigenetic regulation, cell cycle regulation, signaling pathway activation, and oncogenes18,19,20,21. Moreover, the germinal center-related microenvironmental signature stratified DLBCL patients into different risk groups after R-CHOP treatment22.
Several models or genetic soring systems incorporating integrative gene expression analyses are under development, aiming to provide better prognostic information23. A four-gene signature-based score involving immune infiltration separated patients into high- and low-risk groups. Notably, the combination of the gene expression-based score with the international prognostic index (IPI) further improved the risk prediction24. Another study investigating NGS data has shown that MYC/BCL2, microenvironment biomarkers, and genetic subtyping are closely linked to the clinical outcome of DLBCL patients25. Despite these findings, investigation of gene mutation patterns and the degree of gene expression in the TME to explore new prognostic markers and novel therapeutic targets are still very critical.
In the present study, whole-exome sequencing (WES) data from a cohort of DLBCL patients was investigated and further validated by the TCGA database. In addition to the estimated tumor mutational burden, a panel composed of several prespecified gene expression signatures was incorporated as a panel of functional genes. These genes were then harnessed to formulate somatic mutation profiles that hold relevance to the prognosis of the disease, henceforth referred to as the prognostic mutation signature (PMS). Overall, this study aims to assess the utilization of both mutational signatures and common clinicopathological characteristics on prognostic outcomes for DLBCL patients.
Methods
Data source
The DFCI dataset of DLBCL patients (DFCI, Nat Med 2018)16 encompassing clinicopathological characteristics and genomic data was collected for the derivation dataset, which was accessed through cBioPortal (http://www.cbioportal.org). The DFCI derivation cohort consisted of 135 patients with DLBCL including 120 patients who underwent standard R-CHOP therapy. The clinicopathological characteristics, somatic mutation profiles, and survival outcomes of the study cohort were also acquired. Furthermore, the data of DLBCL patients from TCGA database was also collected as the validation cohort. The TCGA validation cohort consisted of 48 patients with DLBCL including 25 patients who ever treated with CHOP-based therapy. The study flowchart is presented in Fig. 1.
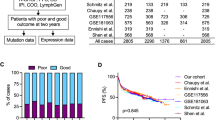
The study flowchart. (a) The genes in the eleven gene sets were screened by utilizing the data from whole exome sequencing in the DFCI cohort. (b) A total of 299 genes were identified for the mutation signature evaluation. Then sixty-one genes without mutations were excluded, resulting in 238 candidate genes. (c) Using Lasso regression, a total of 94 genes related to long-term survival were identified. (d) For validation, another cohort (TCGA) was utilized, and thirty genes were mapped. (e) Finally, the prognostic mutational signature (PMS) was constructed for further investigation.
The clinicopathological characteristics included age at diagnosis, sex, molecular subtype, and IPI (international prognostic index) score. Progression-free survival (PFS) was defined as the time from first treatment until disease progression or death. Because most of the relapses of DLBCL were observed within the first 2 years of diagnosis and the progression rate at five years was low, we observed the five-year survival outcome of the study cohort26,27. Patients who experienced relapse/progression of the disease or died within the study observed period (5 years) were considered the “Cases” group, and patients who achieved durable progression-free status were considered the “Controls” group.
Somatic mutation profiles
The somatic mutation profiles were obtained from the WES of tumor-normal matching sample pairs, and the details of the samples and WES procedure have been well described in a previous publication16. The number of genetic mutations in the cancer cells was computed and is presented as tumor mutational burden (TMB, mut/MB). The somatically mutated status of each gene in candidate functional gene sets was selected and used to derive a PMS for the study cohort according to their PFS status.
Functional gene sets
To investigate the complex genetic event and tumor microenvironment, a panel containing 11 gene functional signatures was utilized28, including DLBCL common genes, proliferation, stroma/EMT/TGF-β, RAS, MYC, WNT, hypoxia, glycolysis, angiogenesis, gMDSC (granulocytic myeloid-derived suppressor cells), and mMDSC (monocytic myeloid-derived suppressor cells) gene sets. The frequent mutated genes in DLBCL, including BCL2, TP53, MYC, MCM5, TSHZ3, KLHL6, MYD88, CD79B, and CREBBP, were defined as DLBCL common gene set (Supplementary Table S1).
Prognostic mutational signature (PMS)
The Lasso Cox model is particularly suitable for high-dimensional somatic mutation data due to its ability to handle sparsity, prevent overfitting, and provide a more interpretable and relevant subset of genomic features for predicting survival outcomes in the context of cancer genomics. Lasso Cox regression analysis was performed to select the optimal gene combination for prognostic risk prediction. The optimal gene combination of candidate functional genes was selected using the Lasso Cox model. The somatic mutation rate of the selected genes in study cohorts was summarized using oncoprints. Afterward, the estimated coefficients (\(\beta\)) of eligible genes in optimal gene combination were computed using Cox regression. The \(\beta\) and mutation status (mut) of each gene (g) were used to generate the candidate prognostic mutational signature (PMS) according to Eq. (1) as follows:
The time-dependent prognostic predictive performance at 12-, 36-, and 60-months of derived PMS, TMB, and IPI were evaluated using ROC analysis, and the area under the ROC curve (AUC) was reported. The threshold values in the AUC for ROC analysis represent the spectrum of sensitivity and specificity trade-offs. The AUC represents the overall discriminatory power of the corresponding feature, while AUC thresholds of 0.7, 0.8, and 0.9 indicate acceptable, good, and excellent dichotomous predictive performance. In addition, the PMS was dichotomized into corresponding low- and high-PMS subgroups according to the optimal cutoff point estimated by ROC. The survival difference between the low- and high-PMS subgroups was further evaluated using the Kaplan‒Meier estimator and tested using the log-rank test.
Statistical analyses
The clinicopathological characteristics, somatic mutation profiles, and survival outcomes were summarized and the difference between Cases and Controls groups was estimated using chi-squared, Fisher’s exact test, or Wilcoxon rank-sum test. Univariate and multivariate Cox proportional hazard regression analyses were performed to evaluate the association between PFS and PMS, while TMB and IPI were considered as covariates for model adjustment. The estimated Cox models were further summarized and illustrated using the forest plot. All p values were two-sided, and p < 0.05 was considered statistically significant. All analyses were conducted using R 4.1.229.
Results
Baseline characteristics of the study cohort
The clinicopathological characteristics, TMB, somatic mutation, and all-cause mortality status of the DFCI derivation cohort according to PFS status are summarized in Supplementary Table S2. There were 52 patients with disease progression (Cases group) and 68 progression-free patients (Controls group). The case group was older and had a higher proportion of females. Both groups had a similar percentage in the molecular subtype. The Cases group also showed a higher proportion of high IPI scores than the Controls group. Notably, 42 (80.8%) patients in the Cases group died during the follow-up period. The basic characteristics of the TCGA validation cohort were also summarized (Supplementary Table S3). The distribution of clinicopathological characteristics, TMB, and survival status between Cases and Controls from the TCGA validation cohort did not show a significant difference.
Somatic mutation profiles
The somatic mutation of the derivation cohort (n = 120) was first investigated. The most common somatic mutations were BCL2, TP53, and CREBBP (Supplementary Table S1). Among the patients with the ABC subtype, the most frequent mutations are in MYD88, CD79B, and TP53. In the GCB group, the most frequent mutations included BCL2, TP53, and CREBBP. Besides, BCL2, CREBBP, and TP53 are the most mutated genes in the unclassified group (Supplementary Table S1).
Next, to investigate survival-related somatic mutations, the genes in the 11 gene sets were first selected, and then the selected genes without any mutation variants in this study cohort were excluded. After the initial evaluation, the candidate genes were selected for further analysis, including DLBCL common (9 genes), proliferation (75 genes), stroma/EMT/TGF-β (34 genes), RAS (2 genes), MYC (8 genes), WNT (9 genes), hypoxia (4 genes), glycolysis (8 genes), angiogenesis (3 genes), gMDSC (15 genes), and mMDSC (80 genes).
Significant prognostic mutational signature
The results of the Lasso Cox regression analysis are shown in Fig. 2. The optimal gene combination for prognostic risk prediction was selected based on the log lambda (λ) validation obtained from the Lasso Cox model (Fig. 2a,b). We demonstrated that the estimated C-index from 94 genes combination exhibited optimal prediction performance (Fig. 2b). Figure 2c summarizes the mutation rate of 94 selected genes for PMS estimation in both relapse-free (blue bar) and relapse (red bar) patients, employing stacked bar plots. In the DFCI derivation cohort, BCL2, TP53, CREBBP2, MYD88, and CD79B were the five most common mutated genes (Fig. 2c). The \(\beta\)-value (standardized regression coefficients) of each gene in the optimal gene combination was presented in Fig. 2d, and the details were summarized in Supplementary Table S4. There were 51 genes related to an elevated risk of progression, whereas 43 genes showed a lower risk of progression (Fig. 2d). The somatic mutation profiles of the 94 PMS genes in both DFCI derivation and TCGA validation cohorts were also illustrated using oncoprints (Fig. 3). The common mutated genes in DFCI and TCGA cohorts include BCL2 (DFCI: 24%, TCGA: 11%), TP53 (DFCI: 22%, TCGA: 16%), CREBBP2 (DFCI: 21%, TCGA: 21%), MYD88 (DFCI: 18%, TCGA: 11%), and CD79B (DFCI:16%, TCGA:5%).
Lasso Cox regression analysis results. (a) Coefficients profiles of 238 candidate functional genes. (b) C-index profiles of estimated gene combinations. (c) The mutation rate of 94 selected genes according to relapse status, the blue bar indicates mutated relapse-free patients, and the red bar indicates mutated relapsed patients. (d) Estimated coefficients of 94 selected genes for PMS estimation (See details information in Table S4).
Predictive performance of the prognostic model
The results of the time-dependent ROC analysis for the prognostic prediction of PMS, TMB, and IPI at 12-, 36-, and 60-months PFS are shown in Fig. 4a. Notably, the time-dependent AUC values of PMS (AUCs: 0.982 to 0.987) increased over time, while the time-dependent AUC values of TMB (AUCs: 0.677 to 0.501) and IPI (AUCs: 0.740 to 0.674) decreased slightly. Moreover, PMS also obtained better predictive ability for both short-term and long-term PFS prediction compared to TMB and IPI. Based on AUC evaluation, the optimal cutoff points of PMS, TMB, and IPI were 0.33, 2.23, and 3, respectively. The survival analysis revealed that the high PMS subgroups were associated with a significantly worse PFS (p < 0.001) as shown in Fig. 4b. Notably, these findings were consistently found in the validation cohort (Fig. 4c, p = 0.034). The 60-month PFS rate of the high PMS subgroup in the DFCI derivation cohort (18.7%, 95% CI 10.4–33.8) and TCGA validation cohort (53.3%, 95% CI 21.4–100) were significantly worse compared to low PMS subgroups (DFCI: 95.8%, 95% CI 91.2–100; TCGA 82.6%, 95% CI 66.3–100). The progression-free Controls group had significantly lower PMS than the Cases group regardless of molecular subtypes (Supplementary Fig. S1a). Moreover, high PMS subgroups were also related to a significantly poorer PFS regardless of the molecular subtypes (Supplementary Fig. S1b–d). Specifically, the 60-month PFS rate of high PMS subgroups had a worse PFS compared to low PMS subgroups in ABC (high PMS vs low PMS: 22.2%vs 92.6%), GCB (high PMS vs low PMS: 16.7% vs 100%), and unclassified subtype (high PMS vs low PMS: 16.7%, vs 96.0%) in DFCI derivation cohort. These results indicate the potential for the proposed PMS to predict long-term prognosis across different subtypes.
Furthermore, the association between PFS and PMS, TMB, and IPI in DFCI derivation and TCGA validation cohorts was illustrated in Fig. 5 using the forest plot. The findings suggested that high PMS could be linked to a higher risk of progression in the DFCI derivation cohort (HR = 1.02, 95% CI 1.02–1.03). Similar findings were also found in the TCGA validation cohort (HR = 1.01, 95% CI 0.98–1.02), although not statistically significant. However, when TMB and IPI were added to the evaluation, the risk prediction of PMS in both cohorts was enhanced. Specifically, when TMB was involved as the covariate, PMS could obtain more significant results in PFS in both DFCI derivation (HR = 2.72, 95% CI 1.99–3.71) and TCGA validation (HR = 1.01, 95% CI 1.00–1.03) cohorts. Concordant findings were also observed for the addition of IPI in DFCI derivation (HR = 2.72, 95% CI 1.99–3.72) cohorts.
To further explore the functional role of PMS-involved genes, we first annotated the 94 selected PMS genes to the drug-gene interaction database (DGIdb). The drug-gene interaction was visualized using a chord diagram as shown in Supplementary Fig. S2. Doxorubicin was more likely associated with TP53, followed by BCL2, BRCA1, EZH2, and FCGR3A. Cyclophosphamide was more likely associated with TP53, followed by BRCA1, ITGAL, and ITGB2. In addition, prednisolone was found related to FCGR3A and ITGB1. Based on these findings, the agents in the CHOP regimen showed interaction with genes related to DNA damage, TP53, apoptosis, and immune cell functions. Further pathway analysis showed the most relevant pathways of these 94 genes were associated with DNA damage, cell cycle, inflammation, and immune cell functions (Supplementary Table S5).
Discussion
DLBCL is characterized by heterogeneous genetic events and complicated interactions between lymphoma cells and tumor microenvironment. Through a combined approach utilizing next-generation sequencing and clinical data, we uncovered the somatic mutation profile of DLBCL with clinical significance. The most notable finding of the present study was the identification of PMS for the prediction of long-term survival. Importantly, some molecular pathways related to the constructed PMS may play an important role in the cellular functions of cancer cells including the cell cycle, DNA damage, inflammation, and immune function, were identified from the PMS.
The application of polygenic risk estimation in cancer patients has increased in popularity recently, which was driven by notable advancements in polygenic risk analysis. This approach enables the comprehensive assessment of the collective impact of multiple genetic variants associated with a specific condition30,31. When it comes to cancer, PMS can provide valuable insights into a patient's genetic susceptibility to disease development and progression32. According to the AUC evaluation, this study demonstrated that the constructed PMS involving 94 somatic mutation genes may exhibit better prediction capability than the conventional IPI score, a predictive model incorporating clinical parameters established around 30 years ago33. One possible explanation could be attributed to the complex genetic or epigenetic abnormalities in tumorigenesis, which can be meticulously analyzed through contemporary high-throughput sequencing technology and bioinformatics studies. Besides, previous studies suggested the accumulation of somatic mutations was related to the development of diseases, including cancer34,35,36. Hence, this analytical approach may hold the potential to provide comprehensive and invaluable prognostic information. For example, a study investigating NGS data for mutational status and its clinical relevance in patients with acute myeloid leukemia revealed a higher number of somatic mutations were associated with a worse outcome37. Another study utilizing NGS data to explore somatic mutation also identified certain gene mutations closely linked to overall survival in patients with anaplastic thyroid carcinoma38. Importantly, this study revealed the combination of genetic and clinical data can further augment the predictive capacity for long-term survival. Based on the above statement, with the advance of genetic and bioinformatic analysis, more prognostic prediction models would be developed and investigated in clinical studies39,40,41.
In the present study, we also explored the drug-gene interaction between the CHOP regimen and 94 genes in constructed PMS, the results revealed some genes like TP53, BCL2, BRCA1, EZH2, FCGR3A, ITGAL, and ITGB1 had significant interaction with therapeutic agents, which was concordant with the previous studies. For example, the existence of TP53 mutation was found to be negatively related to survival in patients with DLBCL who received R-CHOP treatment42. Another study indicated the expression of DNA damage response pathway and BCL-2 was linked to poorer outcome24,43. Moreover, the genetic and functional profile of immune cells also showed significant potential for outcome prediction24,44. Collectively, the constructed PMS would contain comprehensive parameters related to tumorigenesis and may provide more prognostic information.
The present study still had some limitations. First, the gene numbers in certain candidate gene sets were limited, causing potentially meaningful genes and PMS to be missed, which may have been due to a low incidence of mutation in these excluded genes. Second, the validation cohort had a comparably limited patient number and lacked consistent demographic data. In addition, approximately 30% of patients belonged to the unclassified molecular group, which might affect the result of survival analysis. Despite these issues, the identified PMS still showed satisfactory prediction performance in PFS.
Based on the above, the combination of polygenic risk estimation and clinical parameters would provide prognostic information for long-term survival in cancer patients. Regarding the perspectives, several new analytic models are also developed and investigated for mechanistic exploration and potential therapeutic therapeutic target identification. For example, a study utilizing ordinary differential equations-based modeling revealed the proteins in dynamic assembling/de-assembling of TNF signaling complexes and determination of cell death outcome45. Another study developed a novel mathematical model to investigate the establishment of molecular compositions within mRNA-driven protein droplets. The findings revealed that in a mixed system of two mRNAs sharing a common binding protein, the droplets preferentially assemble separately rather than colocalize, with competition occurring between them for protein recruitment46. Furthermore, the advances in computational biology like machine learning models can help us gain more insight into the complex crosstalk between genetic markers and related diseases47,48,49,50,51, as well as the development of genetic risk models52,53. Recently, the exploration of the interaction between long non-coding RNA and microRNA also provided valuable information49,54. Collectively, the substantial output of data produced by high-throughput sequencing represents an important breakthrough in biological research. Utilizing sophisticated bioinformatics investigative tools, the results can unveil novel mechanisms and guide subsequent functional studies.
In summary, the major contribution of this study was that we combined gene expression signatures with NGS data to identify novel molecular prognostic markers. We first identified frequent somatic mutations and then constructed the PMS, which may serve as predictors for long-term survival in DLBCL patients. The exploration of the relevant signaling pathways and genetic alterations may provide new information for further investigation to gain more insight into disease mechanisms.
Data availability
The data presented in this study are available downloaded from cBioPortal. 1. DFCI derivation cohort: https://cbioportal-datahub.s3.amazonaws.com/dlbcl_dfci_2018.tar.gz. 2. Validation cohort: https://cbioportal-datahub.s3.amazonaws.com/dlbc_tcga_pan_can_atlas_2018.tar.gz.
References
Morton, L. M. et al. Lymphoma incidence patterns by WHO subtype in the United States, 1992–2001. Blood 107, 265–276. https://doi.org/10.1182/blood-2005-06-2508 (2006).
Flowers, C. R., Sinha, R. & Vose, J. M. Improving outcomes for patients with diffuse large B-cell lymphoma. CA Cancer J. Clin. 60, 393–408. https://doi.org/10.3322/caac.20087 (2010).
Coiffier, B. et al. CHOP chemotherapy plus rituximab compared with CHOP alone in elderly patients with diffuse large-B-cell lymphoma. N. Engl. J. Med. 346, 235–242. https://doi.org/10.1056/NEJMoa011795 (2002).
Pfreundschuh, M. et al. CHOP-like chemotherapy with or without rituximab in young patients with good-prognosis diffuse large-B-cell lymphoma: 6-year results of an open-label randomised study of the MabThera International Trial (MInT) Group. Lancet Oncol. 12, 1013–1022. https://doi.org/10.1016/S1470-2045(11)70235-2 (2011).
Sehn, L. H. et al. Introduction of combined CHOP plus rituximab therapy dramatically improved outcome of diffuse large B-cell lymphoma in British Columbia. J. Clin. Oncol. 23, 5027–5033. https://doi.org/10.1200/JCO.2005.09.137 (2005).
Coiffier, B. et al. Long-term outcome of patients in the LNH-98.5 trial, the first randomized study comparing rituximab-CHOP to standard CHOP chemotherapy in DLBCL patients: A study by the Groupe d’Etudes des Lymphomes de l’Adulte. Blood 116, 2040–2045. https://doi.org/10.1182/blood-2010-03-276246 (2010).
Pfreundschuh, M. et al. Six versus eight cycles of bi-weekly CHOP-14 with or without rituximab in elderly patients with aggressive CD20+ B-cell lymphomas: A randomised controlled trial (RICOVER-60). Lancet Oncol. 9, 105–116. https://doi.org/10.1016/S1470-2045(08)70002-0 (2008).
He, M. Y. & Kridel, R. Treatment resistance in diffuse large B-cell lymphoma. Leukemia 35, 2151–2165. https://doi.org/10.1038/s41375-021-01285-3 (2021).
Rosenwald, A. et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N. Engl. J. Med. 346, 1937–1947. https://doi.org/10.1056/NEJMoa012914 (2002).
Lenz, G. et al. Stromal gene signatures in large-B-cell lymphomas. N. Engl. J. Med. 359, 2313–2323. https://doi.org/10.1056/NEJMoa0802885 (2008).
Scott, D. W. et al. Prognostic significance of diffuse large B-cell lymphoma cell of origin determined by digital gene expression in formalin-fixed paraffin-embedded tissue biopsies. J. Clin. Oncol. 33, 2848–2856. https://doi.org/10.1200/JCO.2014.60.2383 (2015).
Johnson, N. A. et al. Lymphomas with concurrent BCL2 and MYC translocations: The critical factors associated with survival. Blood 114, 2273–2279. https://doi.org/10.1182/blood-2009-03-212191 (2009).
Ennishi, D. et al. Genetic profiling of MYC and BCL2 in diffuse large B-cell lymphoma determines cell-of-origin-specific clinical impact. Blood 129, 2760–2770. https://doi.org/10.1182/blood-2016-11-747022 (2017).
Huang, W. et al. MYC/BCL2/BCL6 triple hit lymphoma: A study of 40 patients with a comparison to MYC/BCL2 and MYC/BCL6 double hit lymphomas. Mod. Pathol. 31, 1470–1478. https://doi.org/10.1038/s41379-018-0067-x (2018).
Lacy, S. E. et al. Targeted sequencing in DLBCL, molecular subtypes, and outcomes: A Haematological Malignancy Research Network report. Blood 135, 1759–1771. https://doi.org/10.1182/blood.2019003535 (2020).
Chapuy, B. et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 24, 679–690. https://doi.org/10.1038/s41591-018-0016-8 (2018).
Schmitz, R. et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N. Engl. J. Med. 378, 1396–1407. https://doi.org/10.1056/NEJMoa1801445 (2018).
Rushton, C. K. et al. Genetic and evolutionary patterns of treatment resistance in relapsed B-cell lymphoma. Blood Adv. 4, 2886–2898. https://doi.org/10.1182/bloodadvances.2020001696 (2020).
Wise, J. F. et al. Mutational dynamics and immune evasion in diffuse large B-cell lymphoma explored in a relapse-enriched patient series. Blood Adv. 4, 1859–1866. https://doi.org/10.1182/bloodadvances.2019001325 (2020).
Jardin, F. et al. Diffuse large B-cell lymphomas with CDKN2A deletion have a distinct gene expression signature and a poor prognosis under R-CHOP treatment: A GELA study. Blood 116, 1092–1104. https://doi.org/10.1182/blood-2009-10-247122 (2010).
Morin, R. D. et al. Genetic landscapes of relapsed and refractory diffuse large B-cell lymphomas. Clin. Cancer Res. 22, 2290–2300. https://doi.org/10.1158/1078-0432.CCR-15-2123 (2016).
Miyawaki, K. et al. A germinal center-associated microenvironmental signature reflects malignant phenotype and outcome of DLBCL. Blood Adv. 6, 2388–2402. https://doi.org/10.1182/bloodadvances.2021004618 (2022).
Reddy, A. et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell 171(481–494), e415. https://doi.org/10.1016/j.cell.2017.09.027 (2017).
Merdan, S. et al. Gene expression profiling-based risk prediction and profiles of immune infiltration in diffuse large B-cell lymphoma. Blood Cancer J. 11, 2. https://doi.org/10.1038/s41408-020-00404-0 (2021).
Xu-Monette, Z. Y. et al. Genetic subtyping and phenotypic characterization of the immune microenvironment and MYC/BCL2 double expression reveal heterogeneity in diffuse large B-cell lymphoma. Clin. Cancer Res. 28, 972–983. https://doi.org/10.1158/1078-0432.CCR-21-2949 (2022).
Wang, Y. et al. Late relapses in patients with diffuse large B-cell lymphoma treated with immunochemotherapy. J. Clin. Oncol. 37, 1819–1827. https://doi.org/10.1200/JCO.19.00014 (2019).
Larouche, J. F. et al. Lymphoma recurrence 5 years or later following diffuse large B-cell lymphoma: Clinical characteristics and outcome. J. Clin. Oncol. 28, 2094–2100. https://doi.org/10.1200/JCO.2009.24.5860 (2010).
Cristescu, R. et al. Transcriptomic determinants of response to pembrolizumab monotherapy across solid tumor types. Clin. Cancer Res. 28, 1680–1689. https://doi.org/10.1158/1078-0432.CCR-21-3329 (2022).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/ (2021).
Janssens, A. Validity of polygenic risk scores: Are we measuring what we think we are?. Hum. Mol. Genet. 28, R143–R150. https://doi.org/10.1093/hmg/ddz205 (2019).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224. https://doi.org/10.1038/s41588-018-0183-z (2018).
Lewis, C. M. & Vassos, E. Polygenic risk scores: From research tools to clinical instruments. Genome Med. 12, 44. https://doi.org/10.1186/s13073-020-00742-5 (2020).
International Non-Hodgkin’s Lymphoma Prognostic Factors P. A predictive model for aggressive non-Hodgkin’s lymphoma. N. Engl. J. Med. 329, 987–994. https://doi.org/10.1056/NEJM199309303291402 (1993).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489. https://doi.org/10.1126/science.aab4082 (2015).
Stephens, P. J. et al. Complex landscapes of somatic rearrangement in human breast cancer genomes. Nature 462, 1005–1010. https://doi.org/10.1038/nature08645 (2009).
Forbes, S. A. et al. COSMIC: Exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805-811. https://doi.org/10.1093/nar/gku1075 (2015).
Yu, J. et al. Gene mutational analysis by NGS and its clinical significance in patients with myelodysplastic syndrome and acute myeloid leukemia. Exp. Hematol. Oncol. 9, 2. https://doi.org/10.1186/s40164-019-0158-5 (2020).
Wang, J. R. et al. Impact of somatic mutations on survival outcomes in patients with anaplastic thyroid carcinoma. JCO Precis. Oncol. 6, e2100504. https://doi.org/10.1200/PO.21.00504 (2022).
Yang, D., Ma, X. & Song, P. A prognostic model of non small cell lung cancer based on TCGA and ImmPort databases. Sci. Rep. 12, 437. https://doi.org/10.1038/s41598-021-04268-7 (2022).
Huang, S. et al. Comprehensive analysis of prognostic genes in gastric cancer. Aging (Albany NY) 13, 23637–23651. https://doi.org/10.18632/aging.203638 (2021).
Smith, J. C. & Sheltzer, J. M. Genome-wide identification and analysis of prognostic features in human cancers. Cell Rep. 38, 110569. https://doi.org/10.1016/j.celrep.2022.110569 (2022).
Xu-Monette, Z. Y. et al. Mutational profile and prognostic significance of TP53 in diffuse large B-cell lymphoma patients treated with R-CHOP: Report from an International DLBCL Rituximab-CHOP Consortium Program Study. Blood 120, 3986–3996. https://doi.org/10.1182/blood-2012-05-433334 (2012).
Rossi, A. et al. Dual targeting of the DNA damage response pathway and BCL-2 in diffuse large B-cell lymphoma. Leukemia 36, 197–209. https://doi.org/10.1038/s41375-021-01347-6 (2022).
Autio, M. et al. Immune cell constitution in the tumor microenvironment predicts the outcome in diffuse large B-cell lymphoma. Haematologica 106, 718–729. https://doi.org/10.3324/haematol.2019.243626 (2021).
Li, X. et al. RIP1-dependent linear and nonlinear recruitments of caspase-8 and RIP3 respectively to necrosome specify distinct cell death outcomes. Protein Cell 12, 858–876. https://doi.org/10.1007/s13238-020-00810-x (2021).
Xu, F. et al. Specificity and competition of mRNAs dominate droplet pattern in protein phase separation. Phys. Rev. Res. 5, 023159. https://doi.org/10.1103/PhysRevResearch.5.023159 (2023).
Sun, F., Sun, J. & Zhao, Q. A deep learning method for predicting metabolite-disease associations via graph neural network. Brief. Bioinform. https://doi.org/10.1093/bib/bbac266 (2022).
Wang, T., Sun, J. & Zhao, Q. Investigating cardiotoxicity related with hERG channel blockers using molecular fingerprints and graph attention mechanism. Comput. Biol. Med. 153, 106464. https://doi.org/10.1016/j.compbiomed.2022.106464 (2023).
Wang, W., Zhang, L., Sun, J., Zhao, Q. & Shuai, J. Predicting the potential human lncRNA-miRNA interactions based on graph convolution network with conditional random field. Brief. Bioinform. https://doi.org/10.1093/bib/bbac463 (2022).
Hu, H. et al. Gene function and cell surface protein association analysis based on single-cell multiomics data. Comput. Biol. Med. 157, 106733. https://doi.org/10.1016/j.compbiomed.2023.106733 (2023).
Meng, R., Yin, S., Sun, J., Hu, H. & Zhao, Q. scAAGA: Single cell data analysis framework using asymmetric autoencoder with gene attention. Comput. Biol. Med. 165, 107414. https://doi.org/10.1016/j.compbiomed.2023.107414 (2023).
Chen, Z. et al. DCAMCP: A deep learning model based on capsule network and attention mechanism for molecular carcinogenicity prediction. J. Cell. Mol. Med. 27, 3117–3126. https://doi.org/10.1111/jcmm.17889 (2023).
Gao, H. et al. Predicting metabolite-disease associations based on auto-encoder and non-negative matrix factorization. Brief. Bioinform. https://doi.org/10.1093/bib/bbad259 (2023).
Zhang, L., Yang, P., Feng, H., Zhao, Q. & Liu, H. Using network distance analysis to predict lncRNA-miRNA interactions. Interdiscip. Sci. 13, 535–545. https://doi.org/10.1007/s12539-021-00458-z (2021).
Funding
This study was supported by a grant from Kaohsiung Medical University Hospital (KMUH110-0R21, KMUH111-1R15, KMUH112-2R23) and the Taiwan Ministry of Science and Technology (MOST 110-2628-B-037-009, 111-2314-B-037-050-MY2). This study is also partially supported by Kaohsiung Medical University Research Center Grant (KMU-TC112B04) and KMUH-DK(C)11300X. The funders had no role in the study design, data collection, data analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: SFC, TJY, HCW, JSD, YCG, YYL, TMC, YCL, HHH, and SHM. methodology: SFC and SHM. Data curation: SFC and SHM. Writing—original draft preparation: SFC, TJY, and SHM. Writing—review, and editing: SFC and SHM. Supervision: SFC and SHM. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cho, SF., Yeh, TJ., Wang, HC. et al. Prognostic mutation signature would serve as a potential prognostic predictor in patients with diffuse large B-cell lymphoma. Sci Rep 14, 6161 (2024). https://doi.org/10.1038/s41598-024-56583-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-56583-4
- Springer Nature Limited