Abstract
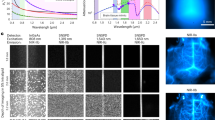
Applying rational design, we developed 17 kDa cyanobacteriochrome-based near-infrared (NIR-I) fluorescent protein, miRFP718nano. miRFP718nano efficiently binds endogenous biliverdin chromophore and brightly fluoresces in mammalian cells and tissues. miRFP718nano has maximal emission at 718 nm and an emission tail in the short-wave infrared (SWIR) region, allowing deep-penetrating off-peak fluorescence imaging in vivo. The miRFP718nano structure reveals the molecular basis of its red shift. We demonstrate superiority of miRFP718nano-enabled SWIR imaging over NIR-I imaging of microbes in the mouse digestive tract, mammalian cells injected into the mouse mammary gland and NF-kB activity in a mouse model of liver inflammation.


Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article and its Supplementary Information. All other data that support the findings of the study are available from the corresponding author upon request. The main plasmids constructed in this study, their maps and sequences are deposited to Addgene. For the structure of miRFP670nano: PDB ID 6MGH, https://www.rcsb.org/structure/6mgh. For the structure of miRFP709 PDB ID: 5VIQ, https://www.rcsb.org/structure/5VIQ.
Code availability
The miRFP718nano nucleotide sequence in GenBank is MW627296.1, https://www.ncbi.nlm.nih.gov/nuccore/MW627296.1. The miRFP718nano structural data were deposited at the Protein Data Bank (PDB ID 7LSD), https://www.rcsb.org/structure/7LSD.
Change history
02 February 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41592-023-01798-y
References
Frangioni, J. V. In vivo near-infrared fluorescence imaging. Curr. Opin. Chem. Biol. 7, 626–634 (2003).
Scholkmann, F. et al. A review on continuous wave functional near-infrared spectroscopy and imaging instrumentation and methodology. Neuroimage 85, 6–27 (2014).
Welsher, K. et al. A route to brightly fluorescent carbon nanotubes for near-infrared imaging in mice. Nat. Nanotechnol. 4, 773–780 (2009).
Smith, A. M., Mancini, M. C. & Nie, S. Bioimaging: second window for in vivo imaging. Nat. Nanotechnol. 4, 710–711 (2009).
Feng, Z. et al. Perfecting and extending the near-infrared imaging window. Light Sci. Appl. 10, 197 (2021).
Shcherbakova, D. M., Stepanenko, O. V., Turoverov, K. K. & Verkhusha, V. V. Near-infrared fluorescent proteins: multiplexing and optogenetics across scales. Trends Biotechnol. 36, 1230–1243 (2018).
Oliinyk, O. S., Chernov, K. G. & Verkhusha, V. V. Bacterial phytochromes, cyanobacteriochromes and allophycocyanins as a source of near-infrared fluorescent probes. Int. J. Mol. Sci. 18, 1691 (2017).
Oliinyk, O. S., Shemetov, A. A., Pletnev, S., Shcherbakova, D. M. & Verkhusha, V. V. Smallest near-infrared fluorescent protein evolved from cyanobacteriochrome as versatile tag for spectral multiplexing. Nat. Commun. 10, 279 (2019).
Oliinyk, O. S. et al. Single-domain near-infrared protein provides a scaffold for antigen-dependent fluorescent nanobodies. Nat. Methods 19, 740–750 (2022).
Shcherbakova, D. M. et al. Bright monomeric near-infrared fluorescent proteins as tags and biosensors for multiscale imaging. Nat. Commun. 7, 12405 (2016).
Yu, D. et al. A naturally monomeric infrared fluorescent protein for protein labeling in vivo. Nat. Methods 12, 763–765 (2015).
Komatsu, N. et al. Development of an optimized backbone of FRET biosensors for kinases and GTPases. Mol. Biol. Cell 22, 4647–4656 (2011).
Shcherbakova, D. M. Near-infrared and far-red genetically encoded indicators of neuronal activity. J. Neurosci. Methods 362, 109314 (2021).
Shcherbakova, D. M., Cox Cammer, N., Huisman, T. M., Verkhusha, V. V. & Hodgson, L. Direct multiplex imaging and optogenetics of Rho GTPases enabled by near-infrared FRET. Nat. Chem. Biol. 14, 591–600 (2018).
Li, L., Hsu, H. C., Verkhusha, V. V., Wang, L. V. & Shcherbakova, D. M. Multiscale photoacoustic tomography of a genetically encoded near-infrared FRET biosensor. Adv. Sci. 8, e2102474 (2021).
Shemetov, A. A. et al. A near-infrared genetically encoded calcium indicator for in vivo imaging. Nat. Biotechnol. 39, 368–377 (2021).
Baloban, M. et al. Designing brighter near-infrared fluorescent proteins: insights from structural and biochemical studies. Chem. Sci. 8, 4546–4557 (2017).
Chang, B. et al. A phosphorescent probe for in vivo imaging in the second near-infrared window. Nat. Biomed. Eng. 6, 629–639 (2021).
Jeong, S. et al. Multiplexed in vivo imaging using size-controlled quantum dots in the second near-infrared window. Adv. Healthc. Mater. 7, e1800695 (2018).
Robinson, J. T. et al. In vivo fluorescence imaging in the second near-infrared window with long circulating carbon nanotubes capable of ultrahigh tumor uptake. J. Am. Chem. Soc. 134, 10664–10669 (2012).
Hugenholtz, F. & de Vos, W. M. Mouse models for human intestinal microbiota research: a critical evaluation. Cell. Mol. Life Sci. 75, 149–160 (2018).
Hamano, N., Inada, T., Iwata, R., Asai, T. & Shingu, K. The alpha2-adrenergic receptor antagonist yohimbine improves endotoxin-induced inhibition of gastrointestinal motility in mice. Br. J. Anaesth. 98, 484–490 (2007).
Wang, D. et al. Trans-illumination intestine projection imaging of intestinal motility in mice. Nat. Commun. 12, 1682 (2021).
Liu, T., Zhang, L., Joo, D. & Sun, S. C. NF-κB signaling in inflammation. Signal Transduct. Target Ther. 2, 17023 (2017).
Osorio, F. G., de la Rosa, J. & Freije, J. M. Luminescence-based in vivo monitoring of NF-κB activity through a gene delivery approach. Cell Commun. Signal 11, 19 (2013).
Fushimi, K. & Narikawa, R. Phytochromes and cyanobacteriochromes: photoreceptor molecules incorporating a linear tetrapyrrole chromophore. Adv. Exp. Med Biol. 1293, 167–187 (2021).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Lamzin, V. S., Perrakis, A. & Wilson, K. S. in International Tables for Crystallography, Vol. F, Crystallography of Biological Macromolecules (eds Arnold E. et al.) 525–528 (Kluwer Academic Publishers, 2012).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D. Biol. Crystallogr. 67, 355–367 (2011).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Longo, P. A., Kavran, J. M., Kim, M. S. & Leahy, D. J. Transient mammalian cell transfection with polyethylenimine (PEI). Methods Enzymol. 529, 227–240 (2013).
Acknowledgements
We thank N. Peitsaro and N. Aarnio from the Flow Cytometry Core Facility of the University of Helsinki for the technical assistance. This work was supported by grants from the US National Institutes of Health (NIH) (grant nos. GM122567, EB028143, NS111039 and NS115581), Chan Zuckerberg Initiative (grant no. 226178), Cancer Foundation Finland and Magnus Ehrnrooth Foundation. S.P. was supported in part by the NIH Intramural Research Program for the Vaccine Research Center of the National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Contributions
O.S.O. developed and characterized the protein in vitro and in cultured cells. C.M. developed the SWIR imaging system, performed imaging and analyzed the data. C.T. developed the SWIR imaging system and measured fluorescence emission. H.S. performed animal surgeries. S.P. designed structural biology experiments and analyzed the data. M.B. purified the recombinant proteins and prepared cells for SWIR imaging. J.Y. planned and supervised the SWIR imaging experiments and analyzed the data. V.V.V. conceived, planned and supervised the whole project and together with J.Y., O.S.O. and S.P. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Takeharu Nagai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Structure of miRFP718nano and its red-shifted chromophore.
Overall structures of (a) miRFP670nano (PDB ID: 6MGH), (b) miRFP718nano, and (c) miRFP709 (PDB ID: 5VIQ). The biliverdin (BV) chromophores are shown in magenta. The PAS and GAF domains of miRFP709 are in cyan and yellow, respectively. The BV chromophores in (d) miRFP670nano, (e) miRFP718nano, and (f) miRFP709 bound to the respective Cys residues and their chemical formulas. Carbon, nitrogen, oxygen, and sulfur atoms are in white, blue, red, and yellow, respectively. Sticks representations show only rings A and B of the chromophores and Cys residues. In miRFP670nano, the BV chromophore (d) is bound to Cys86 via the C31 atom, miRFP718nano (e) and miRFP709 (f) have the same chromophore species bound to the Cys57 and Cys20, respectively. (g) Superposition of miRFP670nano (green) and miRFP718nano (yellow) structures. (h) miRFP718nano hydrogen bond network around the chromophores. (i) Stacking interactions between the chromophores and the surrounding residues in miRFP718nano. (j) The chromophores of miRFP718nano bound to the respective Cys57 residues in the 2Fo-Fc electron density map. The map is countered at 2.0σ-levels. (k) Superposition of the chromophores in miRFP670nano (green) and miRFP718nano (yellow). (l) Stabilizing mutations and hydrophobic clusters in miRFP718nano. The residues forming H-bonds are shown in green, hydrophobic clusters (one formed by residues Leu8, Ile11, Val12, Val26, Ile104, Leu114, Met140 and the other by Val15, Phe18, Leu19, Trp128, Phe132, Leu133) are in cyan and magenta.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2, Figs. 1–16, Tables 1–4 and References.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Oliinyk, O.S., Ma, C., Pletnev, S. et al. Deep-tissue SWIR imaging using rationally designed small red-shifted near-infrared fluorescent protein. Nat Methods 20, 70–74 (2023). https://doi.org/10.1038/s41592-022-01683-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01683-0
- Springer Nature America, Inc.
This article is cited by
-
Near-infrared II fluorescence imaging
Nature Reviews Methods Primers (2024)
-
A general strategy to develop fluorogenic polymethine dyes for bioimaging
Nature Chemistry (2024)
-
In vivo NIR-II fluorescence imaging for biology and medicine
Nature Photonics (2024)
-
High-angle deflection of metagrating-integrated laser emission for high-contrast microscopy
Light: Science & Applications (2023)