Abstract
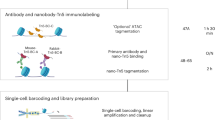
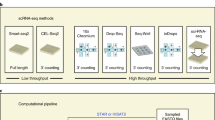
Single-cell combinatorial indexing (sci) with transposase-based library construction increases the throughput of single-cell genomics assays but produces sparse coverage in terms of usable reads per cell. We develop symmetrical strand sci (‘s3’), a uracil-based adapter switching approach that improves the rate of conversion of source DNA into viable sequencing library fragments following tagmentation. We apply this chemistry to assay chromatin accessibility (s3-assay for transposase-accessible chromatin, s3-ATAC) in human cortical and mouse whole-brain tissues, with mouse datasets demonstrating a six- to 13-fold improvement in usable reads per cell compared with other available methods. Application of s3 to single-cell whole-genome sequencing (s3-WGS) and to whole-genome plus chromatin conformation (s3-GCC) yields 148- and 14.8-fold improvements, respectively, in usable reads per cell compared with sci-DNA-sequencing and sci-HiC. We show that s3-WGS and s3-GCC resolve subclonal genomic alterations in patient-derived pancreatic cancer cell lines. We expect that the s3 platform will be compatible with other transposase-based techniques, including sci-MET or CUT&Tag.





Similar content being viewed by others
Data availability
The data discussed in this publication have been deposited in the National Center for Biotechnology Information’s (NCBI’s) Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE174226. External single-cell ATAC datasets were downloaded from GEO sample accession number GSM2668124 for snATAC, and external sites for dscATAC (https://github.com/buenrostrolab/dscATAC_analysis_code/blob/master/mousebrain/data/mousebrain-master_dataframe.rds) and 10X Genomics scATAC (https://cf.10xgenomics.com/samples/cell-atac/1.1.0/atac_v1_adult_brain_fresh_5k). The external single-cell WGS dataset was downloaded from NCBI BioProject PRJNA326698 (https://www.ncbi.nlm.nih.gov/sra/SRX2005587). Single-cell HiC datasets were downloaded from the 4D Nucleosome project (https://data.4dnucleome.org/publications/048d4558-2cac-41d2-ac6e-ff2ac3f007c4/#expsets-table). External bulk HiC datasets have been downloaded from the ENCODE consortium’s data portal, https://www.encodeproject.org/ via accession codes ENCSR194SRI, ENCSR346DCU, ENCSR444WCZ and ENCSR079VIJ. Source data are provided with this paper.
Code availability
Code and custom scripts used in this study are available at https://github.com/adeylab/scitools and https://mulqueenr.github.io/.
References
Cusanovich, D. A. et al. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome Biol. 11, R119 (2010).
Tan, L., Xing, D., Chang, C. H., Li, H. & Xie, X. S. Three-dimensional genome structures of single diploid human cells. Science 361, 924–928 (2018).
Sos, B. C. et al. Characterization of chromatin accessibility with a transposome hypersensitive sites sequencing (THS-seq) assay. Genome Biol. 17, 20 (2016).
Yin, Y. et al. High-throughput single-cell sequencing with linear amplification. Mol. Cell 76, 676–690.e10 (2019).
Chen, C. et al. Single-cell whole-genome analyses by linear amplification via transposon insertion (LIANTI). Science 356, 189–194 (2017).
Adey, A. & Shendure, J. Ultra-low-input, tagmentation-based whole-genome bisulfite sequencing. Genome Res. 22, 1139–1143 (2012).
Mulqueen, R. M. et al. Highly scalable generation of DNA methylation profiles in single cells. Nat. Biotechnol. 36, 428–431 (2018).
Wang, O. et al. Efficient and unique cobarcoding of second-generation sequencing reads from long DNA molecules enabling cost-effective and accurate sequencing, haplotyping, and de novo assembly. Genome Res. 29, 798–808 (2019).
Vitak, S. A. et al. Sequencing thousands of single-cell genomes with combinatorial indexing. Nat. Methods 14, 302–308 (2017).
Amini, S. et al. Haplotype-resolved whole-genome sequencing by contiguity-preserving transposition and combinatorial indexing. Nat. Genet. 46, 1343–1349 (2014).
Adey, A. et al. In vitro, long-range sequence information for de novo genome assembly via transposase contiguity. Genome Res. 24, 2041–2049 (2014).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation.Nat. Neurosci. 21, 432–439 (2018).
Single Cell ATAC: Official (10X Genomics Support, 2020); https://support.10xgenomics.com/single-cell-atac
Lareau, C. A. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility.Nat. Biotechnol. 37, 916–924 (2019).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Bravo González-Blas, C. et al. cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Stuart, T., Srivastava, A., Lareau, C. & Satija, R. Multimodal single-cell chromatin analysis with Signac. Preprint at bioRxiv https://doi.org/10.1101/2020.11.09.373613 (2020).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Thornton, C. A. et al. Spatially mapped single-cell chromatin accessibility. Nat. Commun. 12, 1274 (2021).
Domcke, S. et al. A human cell atlas of fetal chromatin accessibility. Science 370, eaba7721 (2020).
Wang, R., Lin, D. Y. & Jiang, Y. SCOPE: a normalization and copy-number estimation method for single-cell DNA sequencing. Cell Syst. 10, 445–452.e6 (2020).
Laks, E. et al. Clonal decomposition and DNA replication states defined by scaled single-cell genome sequencing. Cell 179, 1207–1221.e22 (2019).
Raphael, B. J. et al. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, e13 (2017).
Lindenburger, K. et al. Drug responses of patient-derived cell lines in vitro that match drug responses of patient PDAc tumors in situ. Ann. Pancreat. Cancer 1, AB024 (2018).
Ramani, V. et al. Massively multiplex single-cell Hi-C. Nat. Methods 14, 263–266 (2017).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Ahmed, S., Bradshaw, A.-D., Gera, S., Dewan, M. & Xu, R. The TGF-β/smad4 signaling pathway in pancreatic carcinogenesis and its clinical significance. J. Clin. Med. 6, 5 (2017).
Wu, H. et al. PRSS1 genotype is associated with prognosis in patients with pancreatic ductal adenocarcinoma. Oncol. Lett. 19, 121–126 (2020).
Kim, H.-J. et al. Capturing cell type-specific chromatin compartment patterns by applying topic modeling to single-cell Hi-C data. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1008173 (2020).
Sritangos, P. et al. Plasma membrane Ca2+ atpase isoform 4 (PMCA4) has an important role in numerous hallmarks of pancreatic cancer. Cancers 12, https://doi.org/10.3390/cancers12010218 (2020).
Ahmad, M. K., Abdollah, N. A., Shafie, N. H., Yusof, N. M. & Razak, S. R. A. Dual-specificity phosphatase 6 (DUSP6): a review of its molecular characteristics and clinical relevance in cancer. Cancer Biol. Med. 15, 14–28 (2018).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Liu, X. et al. Conditional reprogramming and long-term expansion of normal and tumor cells from human biospecimens. Nat. Protoc. 12, 439–451 (2017).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2017).
Sinnamon, J. R. et al. The accessible chromatin landscape of the murine hippocampus at single-cell resolution. Genome Res. 29, 857–869 (2019).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Mcinnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection software review repository archive. J. Open Source Softw. https://doi.org/10.21105/joss.00861 (2018).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from single-cell chromatin accessibility data. Mol. Cell https://doi.org/10.1016/j.molcel.2018.06.044 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
González-Blas, C. B. et al. cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400 (2019).
Jiang, Y. et al. CODEX2: Full-spectrum copy number variation detection by high-throughput DNA sequencing. Genome Biol. 19, 202 (2018).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data.Bioinformatics 32, 2847–2849 (2016).
Drost, H.-G. Philentropy: information theory and distance quantification with R. J. Open Source Softw. 3, 765 (2018).
Galili, T. dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31, 3718–3720 (2015).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
pysam. https://github.com/pysam-developers/pysam (GitHub, 2020).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Acknowledgements
We thank other members of the Adey Laboratory as well as J. Shendure and C. Trapnell for helpful suggestions and feedback. This work was funded by grants R01DA047237 (NIH/NIDA) and R35GM124704 (NIH/NIGMS) to A.C.A. and R01MH113926 (NIH/NIMH) to B.J.O. We also thank the Oregon Brain Bank for the donated biological sample used in this study.
Author information
Authors and Affiliations
Contributions
R.M.M., D.P., F.J.S. and A.C.A. conceived the study. R.M.M. performed all s3 experiments and led all analysis under the supervision of A.C.A. D.P. and F.Z. performed additional experiments under the supervision of F.J.S. B.L.O. and G.G.Y. contributed to the design and analysis of chromatin conformation s3-GCC protocol and datasets. B.J.O. provided support for R.M.M. and advice on analysis. C.A.T. contributed to the analysis of cell types in the s3-ATAC datasets. J.L. generated PDCL cell lines and performed characterization of the lines under supervision of R.C.S. J.L. and R.C.S. contributed to the analysis of PDAC s3-WGS and s3-GCC datasets. The paper was written by R.M.M. and A.C.A. All authors reviewed and contributed to the paper.
Corresponding author
Ethics declarations
Competing interests
D.P., F.Z. and F.J.S. are employees of Scale Bio. R.M.M., D.P., F.Z., F.J.S. and A.C.A. are authors on licensed patents that cover components of the technologies described in this paper. This potential conflict of interest for A.C.A. and R.M.M. has been reviewed and managed by OHSU.
Additional information
Peer review information Nature Biotechnology thanks Kun Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3.
Supplementary Data 1
Mouse brain ATAC-seq peaks.
Supplementary Data 2
s3-ATAC differential accessibility and cell type classification.
Supplementary Data 3
s3-GCC A/B compartment eigenvector plots.
Supplementary Tables
Supplementary Tables 1–8.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Rights and permissions
About this article
Cite this article
Mulqueen, R.M., Pokholok, D., O’Connell, B.L. et al. High-content single-cell combinatorial indexing. Nat Biotechnol 39, 1574–1580 (2021). https://doi.org/10.1038/s41587-021-00962-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-00962-z
- Springer Nature America, Inc.
This article is cited by
-
scAbsolute: measuring single-cell ploidy and replication status
Genome Biology (2024)
-
txci-ATAC-seq: a massive-scale single-cell technique to profile chromatin accessibility
Genome Biology (2024)
-
Fast and flexible profiling of chromatin accessibility and total RNA expression in single nuclei using Microwell-seq3
Cell Discovery (2024)
-
Nano-CUT&Tag for multimodal chromatin profiling at single-cell resolution
Nature Protocols (2024)
-
TIME-seq reduces time and cost of DNA methylation measurement for epigenetic clock construction
Nature Aging (2024)