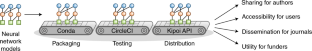
The Kipoi repository provides a home for trained machine-learning models that facilitates their sharing and reuse.
References
Avsec, Z. et al. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0140-0 (2019).
Beaulieu-Jones, B. K. & Greene, C. S. Nat. Biotechnol. 35, 342–346 (2017).
Brendel, W. & Bethge, M. Approximating CNNs with bag-of-local-features models works surprisingly well on ImageNet. Int. Conf. Learn. Represent. https://openreview.net/forum?id=SkfMWhAqYQ (2019).
Taroni, J.N. et al. MultiPLIER: a transfer learning framework for transcriptomics reveals systemic features of rare disease. Preprint at https://doi.org/10.1101/395947 (2019).
Mao, W., Harmann, B., Sealfon, S.C., Zaslavsky, E. & Chikina, M. Pathway-Level Information ExtractoR (PLIER) for gene expression data. Preprint at https://doi.org/10.1101/116061 (2017).
Collado-Torres, L. et al. Nat. Biotechnol. 35, 319–321 (2017).
Kelley, D. R., Snoek, J. & Rinn, J. L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Perou, C. M. Nat. Genet. 29, 373 (2001).
Barrett, T. et al. Nucleic Acids Res. 39, D1005–D1010 (2011).
Rustici, G. et al. Nucleic Acids Res. 41, D987–D990 (2013).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Rights and permissions
About this article
Cite this article
Greene, C.S. Show me the models. Nat Biotechnol 37, 623–625 (2019). https://doi.org/10.1038/s41587-019-0143-x
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0143-x
- Springer Nature America, Inc.


