Abstract
Homologous recombination (HR) fulfils a pivotal role in the repair of DNA double-strand breaks and collapsed replication forks1. HR depends on the products of several paralogues of RAD51, including the tetrameric complex of RAD51B, RAD51C, RAD51D and XRCC2 (BCDX2)2. BCDX2 functions as a mediator of nucleoprotein filament assembly by RAD51 and single-stranded DNA (ssDNA) during HR, but its mechanism remains undefined. Here we report cryogenic electron microscopy reconstructions of human BCDX2 in apo and ssDNA-bound states. The structures reveal how the amino-terminal domains of RAD51B, RAD51C and RAD51D participate in inter-subunit interactions that underpin complex formation and ssDNA-binding specificity. Single-molecule DNA curtain analysis yields insights into how BCDX2 enhances RAD51–ssDNA nucleoprotein filament assembly. Moreover, our cryogenic electron microscopy and functional analyses explain how RAD51C alterations found in patients with cancer3,4,5,6 inactivate DNA binding and the HR mediator activity of BCDX2. Our findings shed light on the role of BCDX2 in HR and provide a foundation for understanding how pathogenic alterations in BCDX2 impact genome repair.






Similar content being viewed by others
Data availability
Atomic coordinates for apo BCDX2 and BCDX2–ssDNA complexes are available at the Research Collaboratory for Structural Bioinformatics PDB with accession codes 8FAZ and 8GBJ, respectively. The corresponding cryo-EM maps have been deposited into the Electron Microscopy Data Bank with accession numbers EMD-28961 (apo BCDX2) and EMD-29917 (BCDX2–ssDNA). The structure data used from the Research Collaboratory for Structural Bioinformatics are available at PDB 7EJC and PDB 7EJE. Source data are provided with this paper.
References
Sung, P. & Klein, H. Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nat. Rev. Mol. Cell Biol. 7, 739–750 (2006).
Sullivan, M. R. & Bernstein, K. A. RAD-ical new insights into RAD51 regulation. Genes https://doi.org/10.3390/genes9120629 (2018).
Meindl, A. et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat. Genet. 42, 410–414 (2010).
Kondrashova, O. et al. Secondary somatic mutations restoring RAD51C and RAD51D associated with acquired resistance to the PARP inhibitor rucaparib in high-grade ovarian carcinoma. Cancer Discov. 7, 984–998 (2017).
Garcin, E. B. et al. Differential requirements for the RAD51 paralogs in genome repair and maintenance in human cells. PLoS Genet. 15, e1008355 (2019).
Prakash, R. et al. Homologous recombination-deficient mutation cluster in tumor suppressor. Proc. Natl Acad. Sci. USA 119, e2202727119 (2022).
Clarke, T. L. & Mostoslavsky, R. DNA repair as a shared hallmark in cancer and ageing. Mol. Oncol. 16, 3352–3379 (2022).
Prakash, R., Zhang, Y., Feng, W. & Jasin, M. Homologous recombination and human health: the roles of BRCA1, BRCA2, and associated proteins. Cold Spring Harb. Perspect. Biol. 7, a016600 (2015).
Sigurdsson, S., Trujillo, K., Song, B., Stratton, S. & Sung, P. Basis for avid homologous DNA strand exchange by human Rad51 and RPA. J. Biol. Chem. 276, 8798–8806 (2001).
Zhao, W. et al. Promotion of BRCA2-dependent homologous recombination by DSS1 via RPA targeting and DNA mimicry. Mol. Cell 59, 176–187 (2015).
Masson, J. Y. et al. Identification and purification of two distinct complexes containing the five RAD51 paralogs. Genes Dev. 15, 3296–3307 (2001).
Martino, J. et al. The human Shu complex functions with PDS5B and SPIDR to promote homologous recombination. Nucleic Acids Res. 47, 10151–10165 (2019).
Yonetani, Y. et al. Differential and collaborative actions of Rad51 paralog proteins in cellular response to DNA damage. Nucleic Acids Res. 33, 4544–4552 (2005).
Nalepa, G. & Clapp, D. W. Fanconi anaemia and cancer: an intricate relationship. Nat. Rev. Cancer 18, 168–185 (2018).
Jacquinet, A. et al. Expanding the FANCO/RAD51C associated phenotype: cleft lip and palate and lobar holoprosencephaly, two rare findings in Fanconi anemia. Eur. J. Med. Genet. 61, 257–261 (2018).
Vaz, F. et al. Mutation of the RAD51C gene in a Fanconi anemia-like disorder. Nat. Genet. 42, 406–409 (2010).
Shamseldin, H. E., Elfaki, M. & Alkuraya, F. S. Exome sequencing reveals a novel Fanconi group defined by XRCC2 mutation. J. Med. Genet. 49, 184–186 (2012).
Miller, K. A., Sawicka, D., Barsky, D. & Albala, J. S. Domain mapping of the Rad51 paralog protein complexes. Nucleic Acids Res. 32, 169–178 (2004).
Xu, J. et al. Mechanisms of distinctive mismatch tolerance between Rad51 and Dmc1 in homologous recombination. Nucleic Acids Res. 49, 13135–13149 (2021).
Pellegrini, L. et al. Insights into DNA recombination from the structure of a RAD51-BRCA2 complex. Nature 420, 287–293 (2002).
Rajendra, E. & Venkitaraman, A. R. Two modules in the BRC repeats of BRCA2 mediate structural and functional interactions with the RAD51 recombinase. Nucleic Acids Res. 38, 82–96 (2010).
Yu, D. S. et al. Dynamic control of Rad51 recombinase by self-association and interaction with BRCA2. Mol. Cell 12, 1029–1041 (2003).
Short, J. M. et al. High-resolution structure of the presynaptic RAD51 filament on single-stranded DNA by electron cryo-microscopy. Nucleic Acids Res. 44, 9017–9030 (2016).
Xu, J. et al. Cryo-EM structures of human RAD51 recombinase filaments during catalysis of DNA-strand exchange. Nat. Struct. Mol. Biol. 24, 40–46 (2017).
Chi, P., Van Komen, S., Sehorn, M. G., Sigurdsson, S. & Sung, P. Roles of ATP binding and ATP hydrolysis in human Rad51 recombinase function. DNA Repair 5, 381–391 (2006).
Sullivan, M. R. et al. Long-term survival of an ovarian cancer patient harboring a RAD51C missense mutation. Cold Spring Harb. Mol. Case Stud. https://doi.org/10.1101/mcs.a006083 (2021).
Matsuo, Y., Sakane, I., Takizawa, Y., Takahashi, M. & Kurumizaka, H. Roles of the human Rad51 L1 and L2 loops in DNA binding. FEBS J. 273, 3148–3159 (2006).
Yang, H., Zhou, C., Dhar, A. & Pavletich, N. P. Mechanism of strand exchange from RecA-DNA synaptic and D-loop structures. Nature 586, 801–806 (2020).
Gibb, B., Silverstein, T. D., Finkelstein, I. J. & Greene, E. C. Single-stranded DNA curtains for real-time single-molecule visualization of protein-nucleic acid interactions. Anal. Chem. 84, 7607–7612 (2012).
Gibb, B. et al. Concentration-dependent exchange of replication protein A on single-stranded DNA revealed by single-molecule imaging. PLoS ONE 9, e87922 (2014).
Ma, C. J., Gibb, B., Kwon, Y., Sung, P. & Greene, E. C. Protein dynamics of human RPA and RAD51 on ssDNA during assembly and disassembly of the RAD51 filament. Nucleic Acids Res. 45, 749–761 (2017).
Ma, C. J., Steinfeld, J. B. & Greene, E. C. Single-stranded DNA curtains for studying homologous recombination. Methods Enzymol. 582, 193–219 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Gayarre, J. et al. Characterisation of the novel deleterious RAD51C p.Arg312Trp variant and prioritisation criteria for functional analysis of RAD51C missense changes. Br. J. Cancer 117, 1048–1062 (2017).
Belan, O. et al. Single-molecule analysis reveals cooperative stimulation of Rad51 filament nucleation and growth by mediator proteins. Mol Cell 81, 1058–1073 (2021).
Roy, U. et al. The Rad51 paralog complex Rad55-Rad57 acts as a molecular chaperone during homologous recombination. Mol. Cell 81, 1043–1057 (2021).
Špírek, M., Taylor, M. R. G., Belan, O., Boulton, S. J. & Krejci, L. Nucleotide proofreading functions by nematode RAD51 paralogs facilitate optimal RAD51 filament function. Nat. Commun. 12, 5545 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Delano, W. The PyMOL molecular graphics system http://www.pymol.org (DeLano Scientific, 2002).
Gaines, W. A. et al. Promotion of presynaptic filament assembly by the ensemble of S. cerevisiae Rad51 paralogues with Rad52. Nat. Commun. 6, 7834 (2015).
De Tullio, L., Kaniecki, K. & Greene, E. C. Single-stranded DNA curtains for studying the Srs2 helicase using total internal reflection fluorescence microscopy. Methods Enzymol. 600, 407–437 (2018).
Greene, E. C., Wind, S., Fazio, T., Gorman, J. & Visnapuu, M. L. DNA curtains for high-throughput single-molecule optical imaging. Methods Enzymol. 472, 293–315 (2010).
Roy, U. & Greene, E. C. Single-stranded DNA curtains for single-molecule visualization of Rad51-ssDNA filament dynamics. Methods Mol. Biol. 2281, 193–207 (2021).
Acknowledgements
We thank members of the laboratories of P.S., E.C.G. and S.K.O. for discussions. Research reported in this publication was supported by NIH grants R01 GM115568 and R01 GM128731 (S.K.O.); R01 CA168635, R01 ES007061, P01 CA92584 and R35 CA241801 (P.S.); R00 GM140264 (E.V.W.); R01 CA221858 and R35 GM118026 (E.C.G.); R01 GM141091 and RSG-22-721675-01-DMC (W.Z.); R01 CA246807 and R01 CA258381 (S.B.); R01 GM136717, R01 CA23728, R01 CA188347 and Congressionally Directed Medical Research Programs BC191160 (A.V.M.); R50 CA265315 (Y.K.); R35 CA253174 (M.J.); R01 ES030335, ES031796 and the Department of Defense BC201356 (K.A.B.); and R01 CA139429 and Cancer Prevention and Research Institute of Texas RP220269 (R.H.). P.S. is the holder of the Robert A. Welch Distinguished Chair in Chemistry (AQ-0012) and the recipient of a CPRIT REI Award (RR180029). A.V.M. is the holder of the Joe R. and Teresa Lozano Long Chair in Cancer Research and is the recipient of a CPRIT REI Award (RR210023). S.K.O. is the recipient of a CPRIT Rising Star Award (RR200030), and E.V.W. is the recipient of a CPRIT Recruitment of First Time Tenure Track Faculty Award (RR220068). E.C.G. is a recipient of a Wellcome Trust Collaborative Award in Science (206292/D/17/Z). The cryo-EM data used for the BCDX2 reconstruction were collected at the UT Health San Antonio Cryo-EM Facility on a Glacios transmission electron microscope equipped with a Falcon IV camera and a Selectris energy filter purchased with the support of University of Texas STARs awards 402-1288 (P.S.) and 402-1317 (S.K.O.). We thank A. Brilot for assistance screening cryo-EM grids at the University of Texas at Austin Sauer Structural Biology Laboratory (RRID:SCR_022951). We thank B. Hunter at the UT Health San Antonio Electron Microscopy Laboratory of South Texas Reference Laboratories, Department of Pathology and Laboratory Medicine, for assistance with negative-stain EM. The SPR assay was carried out in the Center for Innovative Drug Discovery and the Mays Cancer Center Drug Discovery and Structural Biology Shared Resource supported by CPRIT Core Facility Award RP160844 (D.Z.) and NIH-NCI P30 CA054174 (S.K.O. and D.Z.), respectively. The content of this study is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
Protein purification was conducted by Y.R. and H.K. Structural experiments and analysis were conducted by L.J., E.A.R., E.V.W. and S.K.O. Y.R. conducted biochemical, mass photometry and negative-stain EM assays. S.Z. conducted SPR assays. A.M. conducted single-molecule experiments. K.A.B., M.J., Y.K., R.H., A.B.T., S.B., A.V.M., W.Z. and D.Z. assisted with experimental design and data interpretation. The figures and manuscript were prepared by Y.R., L.J., A.M., Y.K., E.V.W., E.C.G., P.S. and S.K.O., with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 BCDX2 reconstitution and cryo-EM data processing flowchart for apo structure.
a, Chromatogram of an analytical gel filtration of the BCDX2 complex in the presence of ATP/Mg2+ on a Superdex200 Increase 10/300 column expressed and purified as described in the Methods (top). Coomassie-stained SDS-PAGE gel of the indicated fractions representing multiple replicates (bottom). b, Mass photometry analysis of the BCDX2 complex purified in the presence (red) and absence (blue) of ATP and Mg2+. Both samples were rapidly diluted into PBS immediately prior to data acquisition. c, Auto-picked particles were first sorted by three rounds of 2D classification, six initial models were made first then followed by four rounds of heterogenous refinement. A final step of stringent heterogenous refinement and tight mask homogeneous refinement led to a final higher resolution map. All data analysis were performed using Cryosparc v3.3.1.
Extended Data Fig. 2 Quality of the apo BCDX2 complex cryo-EM reconstruction.
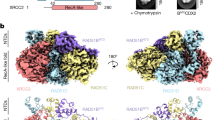
a, The final Euler angle distribution of particles used for 3D refinement. b, Fourier Shell Correlation (FSC) plot from two half maps, with an overall resolution of 2.3 Å as determined at 0.143 criterion. c, Local resolution map color-coded from lower (red) to higher resolution (blue). d, Overall map (left) and detailed views of local cryo-EM density for selected regions of the BCDX2 complex (right). e, Cryo-EM map of the BCDX2 complex with two views related by a 180° rotation about the y-axis (left). Maps of the individual subunits are shown to the right with RAD51B (pink), RAD51C (green), RAD51D (yellow), XRCC2 (purple).
Extended Data Fig. 3 Subunit-subunit interaction networks within the BCDX2 complex.
a, Cartoon representation of the indicated RAD51B/RAD51C interface with residues involved in intermolecular interaction shown as sticks. An overview of the complex is shown on the left with magnified views of the indicated interfaces presented in the three right panels. b–d, The indicated complexes are presented as in panel a with RAD51C/RAD51D (panel b), RAD51D/XRCC2 (panel c), and RAD51/RAD51 from the presynaptic filament (PDB: 7EJC) (panel d). e–g, Structure-based sequence alignment of the indicated proteins. Residue numbers to the left of the sequence, secondary structure elements (RAD51 as a reference) are shown on top of the sequence. Contacts to RAD51 (shaded cyan), RAD51B (salmon), RAD51C (green), RAD51D (gold), and XRCC2 (magenta). In panel g, solid shade indicates ATPase domain contacts to the NTD and boxed shaded regions indicate contacts to the linker. h, Surface electrostatic representation of the complete BCDX2 complex with the ATPase domain of RAD51B modeled using AlphaFold and residues comprising the basic patch which serves as the putative nucleic acid binding site labeled.
Extended Data Fig. 4 Structural features and functional analysis of BCDX2 subunit active sites.
a, Overview of the BCDX2 structure shown as a semitransparent cartoon with. ATP and BCDX2 residues contacting ATP are shown as solid sticks (center). Active sites are boxed. Magnified views of BCDX2 active sites are shown in the left and right panels. b, Structure-based sequence alignment of ATP binding sites of BCDX2 subunits and RAD51. Nucleotides are buried at the interface between subunits and the primary ATP binding site is defined as the subunit with its Walker A motif contacting ATP while the secondary ATP binding site is defined as the subunit contributing interacting residues on the other side of ATP. Interacting residues from primary and secondary ATP binding sites are colored green and pink, respectively. RAD51 (PDB: 7EJC) is shown for reference. c, ATPase assays of 1μM indicated BCDX2 variants in the presence or absence of ssDNA after 60 min incubation. Pi indicates released inorganic phosphate after hydrolysis. The histograms show results from three independent experiments, data are presented as mean values ±SD and p-values for the significance of differences in mean values were calculated by the two-sided t-test are indicated. d. ATP hydrolysis with 1 μM WT and indicated mutant BCDX2 complexes with or without ssDNA were assessed for indicated time points and measurements for three independent experiments were plotted as mean values ±SD. ATPase images in panels c and d were derived from same experiment and processed in parallel.
Extended Data Fig. 5 Cryo-EM data processing flowchart for the BCDX2/ssDNA structure.
Auto-picked particles were first sorted by three rounds of 2D classification. Next, six initial models were generated, followed by four rounds of heterogenous refinement. A subsequent step of stringent heterogenous refinement and tight mask homogeneous refinement of class 3 resulted in a map with 3.12 Å resolution. A final non-uniform refinement yields final map of 3.11Å resolution. All data processing was performed using Cryosparc v3.3.1.
Extended Data Fig. 6 Quality of the BCDX2-ssDNA complex cryo-EM reconstruction.
a, The final Euler angle distribution of particles used for 3D refinement. b, Fourier Shell Correlation (FSC) plot from two half maps, with an overall resolution of 3.1 Å as determined at 0.143 criterion. c, Local resolution map color-coded from lower (red) to higher resolution (blue). d, Overall map (left) and detailed views of local cryo-EM density for AMP-PNP and ssDNA from the BCDX2/ssDNA structure (right). e, Cryo-EM map of the BCDX2 complex with two views related by a 180° rotation about the y-axis (left). Maps of the individual subunits are shown to the right with Rad51B (pink), Rad51C (green), Rad51D (yellow), XRCC2 (purple), ssDNA (cyan).
Extended Data Fig. 7 A potential mechanism for the ssDNA binding specificity of BCDX2.
a, RAD51C from BCDX2 was superposed onto RAD51 from the RAD51/ssDNA postsynaptic structure (PDB: 7EJE). Three subunits from the RAD51 filament are presented. An overview of the superposition is shown in the center panel, while magnified views of the RAD51C/RAD51D and RAD51D/XRCC2 subcomplexes are shown to the right and left, respectively. b, Magnified view of the RAD51D/XRCC2 module from panel a, rotated 180 degrees about the y-axis (left). Right, dsDNA from the RAD51/dsDNA postsynaptic complex is shown as spheres with the RAD51 structure omitted for clarity. Note clashes between dsDNA and XRCC2 due to 11 and 23 degree tilts at the RAD51C/RAD51D and RAD51D/XRCC2 interfaces, respectively, compared to RAD51. c, Surface electrostatic representation of the BCDX2 complex with dsDNA from the superposition presented in panels a and b. Note clashes between dsDNA and XRCC2 and close proximity of the NTDs of BCDX2 to dsDNA. d, Superposition of XRCC2 onto RAD51. Note structural differences in length and conformation of the loop 1 region including helices H12 and H14, which clash with dsDNA in the model.
Extended Data Fig. 8 Comparison of ATPase/ATPase domain interactions in BCDX2 and RAD51.
a–c, Analysis of the amino acid interaction networks at the ATPase/ATPase interfaces of the RAD51C/RAD51D (panel a), and RAD51D/XRCC2 (panel b) subcomplexes of BCDX2, and RAD51/RAD51 from the RAD51/ssDNA presynaptic filament complex (PDB:7EJC) (panel c). Structures are shown as cartoon representations with residues involved in intermolecular interactions shown as sticks. Two views of the complexes related by a 180 degree rotation about the y-axis are shown in the top and bottom panels. d, Structure-based sequence alignment highlighting ATPase domain/ATPase domain interactions at the interfaces highlighted in panels a-c. Residues are colored by the subunit to which they interact: RAD51C (green), RAD51D (gold), XRCC2 (magenta), and RAD51 (cyan). The ATPase domain of RAD51B is not visible in the BCDX2 structure but is included in the sequence alignment for comparison.
Extended Data Fig. 9 Characterization of BCDX2 recombination mediator activity.
a, Representative kymographs of single ssDNA molecules (left panel) showing GFP-BC + DX2 plus ATP (reproduced from Fig. 4a, for comparison), GFP-BC + DX2 minus ATP, BC + GFP-DX2 minus ATP, GFP-BC + DX2 plus ATPγS and GFP-BC + DX2 (green channel; 10 nM each protein complex) plus AMP-PNP (2.5 mM ATP or ATP analogue) binding to mCherry-RPA-ssDNA (red channel). The graph (right panel) shows quantification of GFP-BCDX2 binding to the mCherry-RPA-ssDNA. The graph for GFP-BC + DX2 was reproduced from Fig. 4b for comparison. Data are represented as mean normalized GFP signal intensity; error bars represent 95% CI. Data were derived from three flow cells per reaction condition. b, Representative kymographs of single ssDNA molecules (left panel) showing GFP-BC + DX2 (reproduced from Fig. 4a, for comparison), GFP-BC-R258H + DX2, GFP-BC-R312W + DX2 (10 nM each protein complex) in the presence of 2.5 mM ATP binding to and dissociating from a mCherry-RPA-ssDNA. The graph (right panel) shows quantification of GFP-BCDX2 binding to the mCherry-RPA-ssDNA. The graph for GFP-BC + DX2 (+ATP) was reproduced from Fig. 4b for comparison. Data are represented as mean normalized GFP signal intensity; error bars represent 95% CI. Data were derived using three flow cells per reaction condition. c, Representative kymographs of single RPA-ssDNA molecules showing the behavior of GFP-BC+DX2 in reactions with ATP (left kymograph), ATPγS (center kymograph) and AMP-PNP (right kymograph) during RAD51 filament assembly. d, Quantification of mCherry-RPA dissociation during RAD51 filament assembly in the presence and absence of BCDX2 (left panel), as indicated. Data represented the mean normalized mCherry signal intensity. Data were derived using three flow cells per reaction condition. Calculated rates for RPA-mCherry dissociation from ssDNA during RAD51 filament assembly (right panel). Error bars represent 95% CI. n = 3 flow cells per data set; p-values were calculated by unpaired t-test (two-sided).
Extended Data Fig. 10 Biophysical and biochemical characterization of BCDX2 functions.
a, Representative micrographs depicting RAD51 nucleoprotein filaments seen by negative stain electron microscopy in absence or presence of BCDX2, CDX2 and BCDX2 with RAD51C mutants R258H and R312W. Data represent at least 40 electron micrographs for each sample captured in two independent experiments. b,c, Mass photometry analysis of the RAD51 interactions with (panel b) BCDX2 complex and (panel c) BC subcomplex in the presence of ATP/Mg2+. Plots representing molecular mass profiles RAD51, BCDX2, BC alone and RAD51 when mixed with BCDX2 or BC are differently color coded and molecular mass of most prominent peak in each sample is indicated. d, A two protomer fragment of the RAD51 presynaptic filament (RAD51.1 in slate and RAD51.2 in cyan) were superimposed onto the XRCC2 subunit of XRCC2 subunit of the BCDX2 complex. The overall model is shown in the left panel and a magnified view of the ATP binding site at the interface between XRCC2 and RAD51.1 is shown in the right panel. Note that the β8 strand of XRCC2 is extended by several residues and the loop between β7 and β8 adops a significantly different conformation from RAD51 which clashes with ATP in the model. e, ATP hydrolysis with 1 μM WT and indicated mutant BCDX2 complexes with or without ssDNA were assessed for indicated time points and measurements for three independent experiments were plotted as mean values ±SD. ATPase images were derived from same experiment and processed in parallel.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rawal, Y., Jia, L., Meir, A. et al. Structural insights into BCDX2 complex function in homologous recombination. Nature 619, 640–649 (2023). https://doi.org/10.1038/s41586-023-06219-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06219-w
- Springer Nature Limited