Abstract
CRISPR–Cas systems defend prokaryotic cells from invasive DNA of viruses, plasmids and other mobile genetic elements. Here, we show using metagenomics, metatranscriptomics and single-cell genomics that CRISPR systems of widespread, uncultivated archaea can also target chromosomal DNA of archaeal episymbionts of the DPANN superphylum. Using meta-omics datasets from Crystal Geyser and Horonobe Underground Research Laboratory, we find that CRISPR spacers of the hosts Candidatus Altiarchaeum crystalense and Ca. A. horonobense, respectively, match putative essential genes in their episymbionts’ genomes of the genus Ca. Huberiarchaeum and that some of these spacers are expressed in situ. Metabolic interaction modelling also reveals complementation between host–episymbiont systems, on the basis of which we propose that episymbionts are either parasitic or mutualistic depending on the genotype of the host. By expanding our analysis to 7,012 archaeal genomes, we suggest that CRISPR–Cas targeting of genomes associated with symbiotic archaea evolved independently in various archaeal lineages.




Similar content being viewed by others
Data availability
Metagenomic datasets generated from the CG20,27 ecosystem in 2009, 2014 and 2015 (n = 66) and the HURL24 environment (n = 2) were downloaded from the NCBI SRA in April 2019 (Supplementary Table 1). SAGs generated in a previous study20 (n = 219) were retrieved from the Joint Genome Institute’s Integrated Microbial Genomes and Microbiomes database111 (Supplementary Table 3). The metagenome-derived genomes of Ca. A. crystalense and Ca. H. crystalense from CG are publicly accessible from NCBI (accession numbers in Supplementary Table 2). The genomes of Ca. A. horonobense and Ca. H. julieae from HURL were newly reconstructed in this investigation (Supplementary Table 2). All previously unpublished genomes used in this study are available in figshare https://doi.org/10.6084/m9.figshare.22339555(ref. 112) and all viral genomes are available at https://doi.org/10.6084/m9.figshare.22738568(ref. 113). All raw FISH images are deposited here: https://doi.org/10.6084/m9.figshare.22739849(ref. 114).
Code availability
The code used in this publication is based on previously published code. Please refer to the Methods for information regarding the software and versions used.
References
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67 (2010).
Andersson, A. F. & Banfield, J. F. Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320, 1047–1050 (2008).
Koonin, E. V. & Makarova, K. S. Evolutionary plasticity and functional versatility of CRISPR systems. PLoS Biol. 20, e3001481 (2022).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722 (2015).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Maniv, I., Jiang, W., Bikard, D. & Marraffini, L. A. Impact of different target sequences on type III CRISPR–Cas immunity. J. Bacteriol. 198, 941 (2016).
Marraffini, L. A. & Sontheimer, E. J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Dombrowski, N., Lee, J.-H., Williams, T. A., Offre, P. & Spang, A. Genomic diversity, lifestyles and evolutionary origins of DPANN archaea. FEMS Microbiol. Lett. 366, fnz008 (2019).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Castelle, C. J. et al. Biosynthetic capacity, metabolic variety and unusual biology in the CPR and DPANN radiations. Nat. Rev. Microbiol. 16, 629–645 (2018).
Sakai, H. D. et al. Insight into the symbiotic lifestyle of DPANN archaea revealed by cultivation and genome analyses. Proc. Natl Acad. Sci. USA 119, e2115449119 (2022).
Jahn, U. et al. Nanoarchaeum equitans and Ignicoccus hospitalis: new insights into a unique, intimate association of two archaea. J. Bacteriol. 190, 1743–1750 (2008).
Huber, H. et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417, 63–67 (2002).
Schwank, K. et al. An archaeal symbiont–host association from the deep terrestrial subsurface. ISME J. 13, 2135–2139 (2019).
Hamm, J. N. et al. Unexpected host dependency of Antarctic Nanohaloarchaeota. Proc. Natl Acad. Sci. USA 116, 14661 (2019).
Munson-McGee, J. H. et al. Nanoarchaeota, their Sulfolobales host, and Nanoarchaeota virus distribution across Yellowstone National Park hot springs. Appl. Environ. Microbiol. 81, 7860–7868 (2015).
Jarett, J. K. et al. Single-cell genomics of co-sorted Nanoarchaeota suggests novel putative host associations and diversification of proteins involved in symbiosis. Microbiome 6, 161 (2018).
Wurch, L. et al. Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment. Nat. Commun. 7, 12115 (2016).
Hamm, J. N. et al. The parasitic lifestyle of an archaeal symbiont. Preprint at bioarXiv https://doi.org/10.1101/2023.02.24.5298342.24.529834v2 (2023).
Probst, A. J. et al. Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface. Nat. Microbiol. 3, 328–336 (2018).
Heimerl, T. et al. A complex endomembrane system in the archaeon Ignicoccus hospitalis tapped by Nanoarchaeum equitans. Front. Microbiol. 8, 1072 (2017).
Comolli, L. R. & Banfield, J. F. Inter-species interconnections in acid mine drainage microbial communities. Front. Microbiol. 5, 367 (2014).
Baker, B. J. et al. Enigmatic, ultrasmall, uncultivated Archaea. Proc. Natl Acad. Sci. USA 107, 8806–8811 (2010).
Hernsdorf, A. W. et al. Potential for microbial H2 and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sediments. ISME J. 11, 1915–1929 (2017).
Probst, A. J. et al. Biology of a widespread uncultivated archaeon that contributes to carbon fixation in the subsurface. Nat. Commun. 5, 5497 (2014).
Probst, A. J. et al. Genomic resolution of a cold subsurface aquifer community provides metabolic insights for novel microbes adapted to high CO2 concentrations. Environ. Microbiol. 19, 459–474 (2017).
Emerson, J. B., Thomas, B. C., Alvarez, W. & Banfield, J. F. Metagenomic analysis of a high carbon dioxide subsurface microbial community populated by chemolithoautotrophs and bacteria and archaea from candidate phyla. Environ. Microbiol. 18, 1686–1703 (2016).
Rahlff, J. et al. Lytic archaeal viruses infect abundant primary producers in Earth’s crust. Nat. Commun. 12, 4642 (2021).
Wimmer, F., Mougiakos, I., Englert, F. & Beisel, C. L. Rapid cell-free characterization of multi-subunit CRISPR effectors and transposons. Mol. Cell 82, 1210–1224.e6 (2022).
Marshall, R. et al. Rapid and scalable characterization of CRISPR technologies using an E. coli cell-free transcription-translation system. Mol. Cell 69, 146–157.e3 (2018).
Heussler, G. E. & O’Toole, G. A. Friendly fire: biological functions and consequences of chromosomal targeting by CRISPR–Cas systems. J. Bacteriol. 198, 1481–1486 (2016).
Stern, A., Keren, L., Wurtzel, O., Amitai, G. & Sorek, R. Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet. 26, 335–340 (2010).
Aklujkar, M. & Lovley, D. R. Interference with histidyl-tRNA synthetase by a CRISPR spacer sequence as a factor in the evolution of Pelobacter carbinolicus. BMC Evol. Biol. 10, 230 (2010).
Bhaya, D., Davison, M. & Barrangou, R. CRISPR–Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu. Rev. Genet. 45, 273–297 (2011).
Wilson, G. G. Organization of restriction-modification systems. Nucleic Acids Res. 19, 2539–2566 (1991).
Bornemann, T. L. V. et al. Genetic diversity in terrestrial subsurface ecosystems impacted by geological degassing. Nat. Commun. 13, 284 (2022).
Turgeman-Grott, I. et al. Pervasive acquisition of CRISPR memory driven by inter-species mating of archaea can limit gene transfer and influence speciation. Nat. Microbiol. 4, 177–186 (2019).
Stachler, A.-E. et al. High tolerance to self-targeting of the genome by the endogenous CRISPR–Cas system in an archaeon. Nucleic Acids Res. 45, 5208–5216 (2017).
Vink, J. N. A., Baijens, J. H. L. & Brouns, S. J. J. PAM-repeat associations and spacer selection preferences in single and co-occurring CRISPR–Cas systems. Genome Biol. 22, 281 (2021).
Pyenson, N. C., Gayvert, K., Varble, A., Elemento, O. & Marraffini, L. A. Broad targeting specificity during bacterial type III CRISPR–Cas immunity constrains viral escape. Cell Host Microbe 22, 343–353 (2017).
Chabas, H., Müller, V., Bonhoeffer, S. & Regoes, R. R. Epidemiological and evolutionary consequences of different types of CRISPR-Cas systems. PLoS Comput. Biol. 18, e1010329 (2022).
Brodt, A., Lurie-Weinberger, M. N. & Gophna, U. CRISPR loci reveal networks of gene exchange in archaea. Biol. Direct 6, 65 (2011).
Paper, W. et al. Ignicoccus hospitalis sp. nov., the host of ‘Nanoarchaeum equitans’. Int. J. Syst. Evol. Microbiol. 57, 803–808 (2007).
Dombrowski, N., Teske, A. P. & Baker, B. J. Expansive microbial metabolic versatility and biodiversity in dynamic Guaymas Basin hydrothermal sediments. Nat. Commun. 9, 4999 (2018).
Hohenester, E. & Yurchenco, P. D. Laminins in basement membrane assembly. Cell Adhes. Migr. 7, 56–63 (2013).
Hohenester, E. Laminin G-like domains: dystroglycan-specific lectins. Curr. Opin. Struct. Biol. 56, 56–63 (2019).
Benner, S. A., Ellington, A. D. & Tauer, A. Modern metabolism as a palimpsest of the RNA world. Proc. Natl Acad. Sci. USA 86, 7054–7058 (1989).
Joshi, N. A. & Fass, J. N. Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (v.1.33) Github https://github.com/najoshi/sickle (2011).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Bornemann, T. L. V., Esser, S. P., Stach, T. L., Burg, T. & Probst, A. J. uBin—a manual refining tool for genomes from metagenomes. Environ. Microbiol. 25, 1077–1083 (2023).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Darling, A. E. et al. PhyloSift: phylogenetic analysis of genomes and metagenomes. PeerJ 2, e243 (2014).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Wang, H.-C., Minh, B. Q., Susko, E. & Roger, A. J. Modeling site heterogeneity with posterior mean site frequency profiles accelerates accurate phylogenomic estimation. Syst. Biol. 67, 216–235 (2017).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2017).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Anisimova, M., Gil, M., Dufayard, J.-F., Dessimoz, C. & Gascuel, O. Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Syst. Biol. 60, 685–699 (2011).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 47, W256–W259 (2019).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59 (2014).
Gouy, M., Tannier, E., Comte, N. & Parsons, D. P. in Multiple Sequence Alignment: Methods and Protocols (ed. Katoh, K.) 241–260 (Springer, 2021).
Couvin, D. et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 46, W246–W251 (2018).
Moller, A. G. & Liang, C. MetaCRAST: reference-guided extraction of CRISPR spacers from unassembled metagenomes. PeerJ 5, e3788 (2017).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Biswas, A., Fineran, P. C. & Brown, C. M. Accurate computational prediction of the transcribed strand of CRISPR non-coding RNAs. Bioinformatics 30, 1805–1813 (2014).
Roux, S., Enault, F., Hurwitz, B. L. & Sullivan, M. B. VirSorter: mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Xie, Z. & Tang, H. ISEScan: automated identification of insertion sequence elements in prokaryotic genomes. Bioinformatics 33, 3340–3347 (2017).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Cook, R. et al. INfrastructure for a PHAge REference. Database: identification of large-scale biases in the current collection of cultured phage genomes. Phage 2, 214–223 (2021).
Bolduc, B. et al. vConTACT: an iVirus tool to classify double-stranded DNA viruses that infect archaea and bacteria. PeerJ 5, e3243 (2017).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Meier-Kolthoff, J. P. & Göker, M. VICTOR: genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 33, 3396–3404 (2017).
Moraru, C., Varsani, A. & Kropinski, A. M. VIRIDIC—a novel tool to calculate the intergenomic similarities of prokaryote-infecting viruses. Viruses 12, 1268 (2020).
Meier-Kolthoff, J. P., Auch, A. F., Klenk, H.-P. & Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 14, 60 (2013).
Lefort, V., Desper, R. & Gascuel, O. FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol. Biol. Evol. 32, 2798–2800 (2015).
Farris, J. S. Estimating phylogenetic trees from distance matrices. Am. Nat. 106, 645–668 (1972).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Nishimura, Y. et al. ViPTree: the viral proteomic tree server. Bioinformatics 33, 2379–2380 (2017).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013).
Dufault-Thompson, K., Steffensen, J. L. & Zhang, Y. in Metabolic Network Reconstruction and Modeling: Methods and Protocols (ed. Fondi, M.) 131–150 (Springer, 2018).
Steffensen, J. L., Dufault-Thompson, K. & Zhang, Y. PSAMM: a portable system for the analysis of metabolic models. PLoS Comput. Biol. 12, e1004732–e1004732 (2016).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Gonnerman, M. C., Benedict, M. N., Feist, A. M., Metcalf, W. W. & Price, N. D. Genomically and biochemically accurate metabolic reconstruction of Methanosarcina barkeri Fusaro, iMG746. Biotechnol. J. 8, 1070–1079 (2013).
Goyal, N., Widiastuti, H., Karimi, I. A. & Zhou, Z. A genome-scale metabolic model of Methanococcus maripaludis S2 for CO2 capture and conversion to methane. Mol. Biosyst. 10, 1043–1054 (2014).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Saier, M. H. Jr et al. The transporter classification database (TCDB): recent advances. Nucleic Acids Res. 44, D372–D379 (2016).
Neidhardt, F. C., Neidhardt, F. C. N., Ingraham, J. L. & Schaechter, M. Physiology of the Bacterial Cell: A Molecular Approach (Sinauer Associates, 1990).
Nelson, D. L., Nelson, R. D. & Cox, M. M. Lehninger Principles of Biochemistry (W.H. Freeman, 2004).
Zhang, Y. & Sievert, S. Pan-genome analyses identify lineage- and niche-specific markers of evolution and adaptation in Epsilonproteobacteria. Front. Microbiol. 5, 110 (2014).
Biswas, A., Gagnon, J. N., Brouns, S. J. J., Fineran, P. C. & Brown, C. M. CRISPRTarget: bioinformatic prediction and analysis of crRNA targets. RNA Biol. 10, 817–827 (2013).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Schneider, T. D. & Stephens, R. M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Oberortner, E., Cheng, J.-F., Hillson, N. J. & Deutsch, S. Streamlining the design-to-build transition with build-optimization software tools. ACS Synth. Biol. 6, 485–496 (2017).
Garamella, J., Marshall, R., Rustad, M. & Noireaux, V. The All E. coli TX-TL Toolbox 2.0: a platform for cell-free synthetic biology. ACS Synth. Biol. 5, 344–355 (2016).
Shin, J. & Noireaux, V. An E. coli cell-free expression toolbox: application to synthetic gene circuits and artificial cells. ACS Synth. Biol. 1, 29–41 (2012).
Leenay, R. T. et al. Identifying and visualizing functional PAM diversity across CRISPR–Cas systems. Mol. Cell 62, 137–147 (2016).
Ondov, B. D., Bergman, N. H. & Phillippy, A. M. Interactive metagenomic visualization in a web browser. BMC Bioinform. 12, 385 (2011).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Parks, D. H. et al. A complete domain-to-species taxonomy for bacteria and archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Chen, I.-M. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Esser, S. P. & Probst, A. J. Genomes of Ca. Altiarchaeum and Ca. Huberiarchaeum from Crystal Geyser and Horonobe Underground Research Laboratory. figshare https://doi.org/10.6084/m9.figshare.22339555 (2023).
Esser, S. P., Rahlff, J. & Probst, A. J. Viral operational taxonomic units (vOTUs) from Crystal Geyser. figshare https://doi.org/10.6084/m9.figshare.22738568.v1 (2023).
Turzynski, V., Esser, S. P. & Probst, A. J. Fluorescence in situ hybridization images of Ca. Altiarchaeum and Ca. Huberiarchaeu. figshare https://doi.org/10.6084/m9.figshare.22739849 (2023).
Sharrar, A. M. et al. Novel large sulfur bacteria in the metagenomes of groundwater-fed chemosynthetic microbial mats in the Lake Huron Basin. Front. Microbiol. 8, 791 (2017).
Bird, J. T., Baker, B. J., Probst, A. J., Podar, M. & Lloyd, K. G. Culture independent genomic comparisons reveal environmental adaptations for Altiarchaeales. Front. Microbiol. 7, 1221 (2016).
Posit team. Rstudio: Integrated development environment for R. https://posit.co/; version 2023.03.0+386 (2022).
Acknowledgements
This research was funded by the Ministerium für Kultur und Wissenschaft des Landes Nordrhein- Westfalen (Nachwuchsgruppe Dr. Alexander Probst) and the German Science Foundation under project NOVAC (grant no. DFG PR1603/2-1) and through SPP 2141 (grant no. DFG BE6703/1-1). Genome-scale metabolic modelling was supported by the National Science Foundation under grant no. 1553211. The Ministry of Economy, Trade and Industry of Japan funded a part of the work as ‘The project for validating assessment methodology in geological disposal system’ (2019 FY, grant no. JPJ007597). The work (proposal https://doi.org/10.46936/10.25585/60000800) conducted by the US Department of Energy (DOE) Joint Genome Institute (https://ror.org/04xm1d337), a DOE Office of Science User Facility, is supported by the Office of Science of the US DOE operated under contract no. DE-AC02-05CH11231. M.P. is supported by the Austrian Science Funds (project MAINTAIN, DOC 69 doc.funds). P.S.A. was supported by a postdoctoral fellowship from the Alexander von Humboldt Foundation. J.P. was supported by Aker BP within the framework of the GeneOil Project given to A.J.P. J.R. was supported by the German Science Foundation (grant no. RA3432/1-1, project no. 446702140). Support by the German Federal Ministry of Education and Research within the project ‘MultiKulti’ (BMBF funding code: 161L0285E) is acknowledged. We thank K. Dreger for exemplary server administration and B. Siebers, I. Berg, J. F. Banfield and B. Meyer for insightful discussions.
Author information
Authors and Affiliations
Contributions
S.P.E. and A.J.P. performed genome-resolved metagenomics, while S.P.E. and J.R. performed viromics. J.R. analysed viral genomes with input from M.K. CRISPR–Cas analyses were done by S.P.E., J.R. and A.J.P. SNP analysis was performed by M.P. and T.R. Genome-scale modelling was conducted by W.Z. and Y.Z. with input from S.P.E., P.A.F.G. and A.J.P. J.P. conducted the sliding window analysis with the input from S.P.E., J.R., and A.J.P. Phylogenomic analyses were carried out by P.S.A. T.L.V.B. provided bioinformatic assistance and K. Schwank, I.B. and V.T. performed microscopy and initial metabolic analyses. J.M. and W.B. resampled CG and J.L., T.W. and A.J.P. conducted RNA extraction and sequencing and S.P.E. analysed transcriptomes. J-F.C. synthesized the Cas genes with input from I.K.B., F.W. and C.B. performed binding, cleavage and PAM assays and J.L., J.J., Y.A., T.W. and A.J.P. generated/provided raw data. K. Sures and S.P.E. analysed the archaeal CRISPR–Cas interactions from published NCBI archaeal genomes. A.J.P. conceptualized the work. S.P.E., J.R., W.Z., Y.Z. and A.J.P. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Correlation of repeat abundance and abundance of Ca. Altiarchaea genomes.
Spearman rank correlation (two-tailed) of logarithmic abundances of Ca. A. crystalense and logarithmic abundances of repeat sequences of the unassigned CRISPR array (p-value < 3.4 e−16) and the CRISPR system type I-B (p < 2.2 e−16) in metagenomes from CG (n = 66). The grey area depicts the confidence interval of 0.95. The line indicates that the correlation of the genome abundance and repeat abundance is linear. Visualization was performed with R87,117 (version 3.6.1).
Extended Data Fig. 2 Viral clusters predicted by VIRIDIC79.
Heatmap showing intergenomic similarity for viral scaffolds of viral clusters (VC_XY) and some singletons (black). Colouring of viral OTUs (vOTUs) according to Supplementary Table 6. VC_09, _12, _13 determined by the other tools were not found by VIRIDIC. Only scaffolds with intergenomic similarity of >10 between two viral scaffolds are shown.
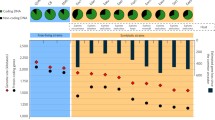
Extended Data Fig. 3 Coverage analyses of scaffolds targeted by spacers from Ca. Altiarchaea.
Coverage changes within targeted regions by CRISPR system type I-B of Ca. Altiarchaeum and Ca. Huberiarchaeum based on metagenomic read mapping. The vertically grey marked regions are spacer targeted regions of either Ca. Altiarchaeum or Ca. Huberiarchaeum, whereby the horizontally dark grey lines are showing the average coverage of the scaffold. The coloured graphs show the coverage across the spacer targeted region of three samples from the minor eruption phase, where Ca. Altiarchaeum is the most abundant organism (Supplementary Fig. 1).
Extended Data Fig. 4 Spacer targeting analyses of publicly available archaeal genomes.
Directed spacer analysis of 7,012 publicly available archaeal genomes (Supplementary Table 4) shows large clusters of spacers targeting at species level. The targeting spacers (edges) of the genomes Sulfolobus, Methanomicrobia and Halobacterium (nodes) form large clusters performing self-targeting or targeting other genomes of the same family. Edges are colored according to their relationship at least familiy level or lower. The clustering was illustrated with Cytoscape83 (version 3.9.1). Please note that targeting within the same genus might limit the interspecies recombination, as demonstrated in haloarchaea37, or reflect the presence of multiple conserved genomic regions between the genomes.
Supplementary information
Supplementary Information
Supplementary Results, Table 7 and Figs. 1–9.
Supplementary Data
Collection of all phylogenetic trees calculated for this study. (1) Phylogenetic tree of Ca. Altiarchaeum crystalense/horonobense and Ca. Huberiarchaeum crystalense/julieae located within the archaeal branch. (2) Phylogenetic analysis of the phenylalanine-tRNA synthetase of Ca. Huberiarchaeum crystalense located within the archaeal branch. Calculated to determine the closest relative within the archaeal branch. (3) Phylogenetic analysis of the phenylalanine-tRNA synthetase of Ca. Altiarchaeum crystalense located within the archaeal branch. Calculated to determine the closest relative within the archaeal branch. (4) Phylogenetic analysis of the lysine-tRNA synthetase of Ca. Huberiarchaeum crystalense located within the archaeal branch. Calculated to determine the closest relative within the archaeal branch. (5) Phylogenetic analysis of the lysine-tRNA synthetase of Ca. Altiarchaeum crystalense located within the archaeal branch. Calculated to determine the closest relative within the archaeal branch.
Supplementary Tables
Supplementary Tables 1–6 and 8–13.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Esser, S.P., Rahlff, J., Zhao, W. et al. A predicted CRISPR-mediated symbiosis between uncultivated archaea. Nat Microbiol 8, 1619–1633 (2023). https://doi.org/10.1038/s41564-023-01439-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01439-2
- Springer Nature Limited
This article is cited by
-
Soil microbial ecology through the lens of metatranscriptomics
Soil Ecology Letters (2024)
-
Neue Funktion von CRISPR-Systemen: Basis für Archaea-Symbiosen
BIOspektrum (2024)
-
CRISPR-influenced symbiosis
Nature Microbiology (2023)