Abstract
The canonical model of agonist-stimulated phosphatidylinositol 3-kinase (PI3K)–Akt signalling proposes that PI3K is activated at the plasma membrane, where receptors are activated and phosphatidylinositol-4,5-bisphosphate is concentrated. Here we show that phosphatidylinositol-3,4,5-trisphosphate generation and activated Akt are instead largely confined to intracellular membranes upon receptor tyrosine kinase activation. Microtubule-associated protein 4 (MAP4) interacts with and controls localization of membrane vesicle-associated PI3Kα to microtubules. The microtubule-binding domain of MAP4 binds directly to the C2 domain of the p110α catalytic subunit. MAP4 controls the interaction of PI3Kα with activated receptors at endosomal compartments along microtubules. Loss of MAP4 results in the loss of PI3Kα targeting and loss of PI3K–Akt signalling downstream of multiple agonists. The MAP4–PI3Kα assembly defines a mechanism for spatial control of agonist-stimulated PI3K–Akt signalling at internal membrane compartments linked to the microtubule network.








Similar content being viewed by others
Data availability
Mass spectrometry data of PI3Kα immunoprecipitates and identification of MAP4 have been deposited at ProteomeXchange with primary accession code PXD021306. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Change history
24 November 2020
In the version of this Article originally published, the expansion of “PI3K” was incorrect in the title and the abstract. Instead of “Phosphatidylinositol-3-OH kinase signalling is spatially organized at endosomal compartments by microtubule-associated protein 4,” the title should read “Phosphatidylinositol 3-kinase signalling is spatially organized at endosomal compartments by microtubule-associated protein 4”. Furthermore, in the first sentence of the abstract, PI3K should again be expanded as “Phosphatidylinositol 3-kinase” and not “Phosphatidylinositol-3-OH kinase”. The errors have been corrected in the HTML version of the paper.
References
Zhang, Y. et al. A pan-cancer proteogenomic atlas of PI3K/AKT/mTOR pathway alterations. Cancer Cell 31, 820–832 (2017).
Fruman, D. A. et al. The PI3K pathway in human disease. Cell 170, 605–635 (2017).
Fruman, D. A. & Rommel, C. PI3K and cancer: lessons, challenges and opportunities. Nat. Rev. Drug Discov. 13, 140–156 (2014).
LoRusso, P. M. Inhibition of the PI3K/AKT/mTOR Pathway in solid tumors. J. Clin. Oncol. 34, 3803–3815 (2016).
Balla, T. Phosphoinositides: tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, 1019–1137 (2013).
Vadas, O., Burke, J. E., Zhang, X., Berndt, A. & Williams, R. L. Structural basis for activation and inhibition of class I phosphoinositide 3-kinases. Sci. Signal. 4, re2 (2011).
Thapa, N. et al. The hidden conundrum of phosphoinositide signaling in cancer. Trends Cancer 2, 378–390 (2016).
Burke, J. E. & Williams, R. L. Synergy in activating class I PI3Ks. Trends Biochem. Sci. 40, 88–100 (2015).
Thapa, N., Horn, H. T. & Anderson, R. A. Phosphoinositide spatially free AKT/PKB activation to all membrane compartments. Adv. Biol. Regul. 72, 1–6 (2019).
Clark, A. R. & Toker, A. Signalling specificity in the Akt pathway in breast cancer. Biochem. Soc. Trans. 42, 1349–1355 (2014).
Kelly, K. L., Ruderman, N. B. & Chen, K. S. Phosphatidylinositol-3-kinase in isolated rat adipocytes. Activation by insulin and subcellular distribution. J. Biol. Chem. 267, 3423–3428 (1992).
Kapeller, R., Chakrabarti, R., Cantley, L., Fay, F. & Corvera, S. Internalization of activated platelet-derived growth factor receptor-phosphatidylinositol-3′ kinase complexes: potential interactions with the microtubule cytoskeleton. Mol. Cell. Biol. 13, 6052–6063 (1993).
Sato, M., Ueda, Y., Takagi, T. & Umezawa, Y. Production of PtdInsP3 at endomembranes is triggered by receptor endocytosis. Nat. Cell Biol. 5, 1016–1022 (2003).
Naguib, A. et al. PTEN functions by recruitment to cytoplasmic vesicles. Mol. Cell 58, 255–268 (2015).
Liu, S. L. et al. Quantitative lipid imaging reveals a new signaling function of phosphatidylinositol-3,4-bisphophate: isoform- and site-specific activation of Akt. Mol. Cell 71, 1092–1104 (2018).
Ebner, M., Lucic, I., Leonard, T. A. & Yudushkin, I. PI(3,4,5)P3 engagement restricts Akt activity to cellular membranes. Mol. Cell 65, 416–431 (2017).
Varnai, P. et al. Selective cellular effects of overexpressed pleckstrin-homology domains that recognize PtdIns(3,4,5)P3 suggest their interaction with protein binding partners. J. Cell Sci. 118, 4879–4888 (2005).
Semenova, I. et al. Regulation of microtubule-based transport by MAP4. Mol. Biol. Cell 25, 3119–3132 (2014).
Wills, R. C., Goulden, B. D. & Hammond, G. R. V. Genetically encoded lipid biosensors. Mol. Biol. Cell 29, 1526–1532 (2018).
Murphy, J. E., Padilla, B. E., Hasdemir, B., Cottrell, G. S. & Bunnett, N. W. Endosomes: a legitimate platform for the signaling train. Proc. Natl Acad. Sci. USA 106, 17615–17622 (2009).
Choi, S. et al. Agonist-stimulated phosphatidylinositol-3,4,5-trisphosphate generation by scaffolded phosphoinositide kinases. Nat. Cell Biol. 18, 1324–1335 (2016).
Chen, M. et al. The nuclear phosphoinositide response to stress. Cell Cycle 19, 268–289 (2020).
Chen, M. et al. The specificity of EGF-stimulated IQGAP1 scaffold towards the PI3K–Akt pathway is defined by the IQ3 motif. Sci. Rep. 9, 9126 (2019).
Sun, Y., Thapa, N., Hedman, A. C. & Anderson, R. A. Phosphatidylinositol 4,5-bisphosphate: targeted production and signaling. Bioessays 35, 513–522 (2013).
Chapin, S. J. & Bulinski, J. C. Non-neuronal 210 × 103 Mr microtubule-associated protein (MAP4) contains a domain homologous to the microtubule-binding domains of neuronal MAP2 and tau. J. Cell Sci. 98, 27–36 (1991).
Seo, M. et al. MAP4-regulated dynein-dependent trafficking of BTN3A1 controls the TBK1–IRF3 signaling axis. Proc. Natl Acad. Sci. USA 113, 14390–14395 (2016).
Chapin, S. J., Lue, C. M., Yu, M. T. & Bulinski, J. C. Differential expression of alternatively spliced forms of MAP4: a repertoire of structurally different microtubule-binding domains. Biochemistry 34, 2289–2301 (1995).
Wu, H. et al. Regulation of class IA PI 3-kinases: C2 domain–iSH2 domain contacts inhibit p85/p110α and are disrupted in oncogenic p85 mutants. Proc. Natl Acad. Sci. USA 106, 20258–20263 (2009).
Huang, C. H. et al. The structure of a human p110α/p85α complex elucidates the effects of oncogenic PI3Kα mutations. Science 318, 1744–1748 (2007).
Bergeron, J. J., Di Guglielmo, G. M., Dahan, S., Dominguez, M. & Posner, B. I. Spatial and temporal regulation of receptor tyrosine kinase activation and intracellular signal transduction. Annu. Rev. Biochem. 85, 573–597 (2016).
Salogiannis, J. & Reck-Peterson, S. L. Hitchhiking: A non-canonical mode of microtubule-based transport. Trends Cell Biol. 27, 141–150 (2017).
Bonifacino, J. S. & Neefjes, J. Moving and positioning the endolysosomal system. Curr. Opin. Cell Biol. 47, 1–8 (2017).
Tang, Y. C. et al. Functional genomics identifies specific vulnerabilities in PTEN-deficient breast cancer. Breast Cancer Res. 20, 22 (2018).
Jhawer, M. et al. PIK3CA mutation/PTEN expression status predicts response of colon cancer cells to the epidermal growth factor receptor inhibitor cetuximab. Cancer Res. 68, 1953–1961 (2008).
Xia, X., He, C., Wu, A., Zhou, J. & Wu, J. Microtubule-associated protein 4 is a prognostic factor and promotes tumor progression in lung adenocarcinoma. Dis. Markers 2018, 8956072 (2018).
Jiang, Y. Y. et al. Microtubule-associated protein 4 is an important regulator of cell invasion/migration and a potential therapeutic target in esophageal squamous cell carcinoma. Oncogene 35, 4846–4856 (2016).
Ou, Y. et al. Activation of cyclic AMP/PKA pathway inhibits bladder cancer cell invasion by targeting MAP4-dependent microtubule dynamics. Urol. Oncol. 32, 47 e21–47 e48 (2014).
Miao, B. et al. Small molecule inhibition of phosphatidylinositol-3,4,5-triphosphate (PIP3) binding to pleckstrin homology domains. Proc. Natl Acad. Sci. USA 107, 20126–20131 (2010).
Jethwa, N. et al. Endomembrane PtdIns(3,4,5)P3 activates the PI3K–Akt pathway. J. Cell Sci. 128, 3456–3465 (2015).
Betz, C. & Hall, M. N. Where is mTOR and what is it doing there? J. Cell Biol. 203, 563–574 (2013).
De Santis, M. C., Gulluni, F., Campa, C. C., Martini, M. & Hirsch, E. Targeting PI3K signaling in cancer: challenges and advances. Biochim. Biophys. Acta. Rev. Cancer 1871, 361–366 (2019).
Kim, J. & Guan, K. L. mTOR as a central hub of nutrient signalling and cell growth. Nat. Cell Biol. 21, 63–71 (2019).
Dibble, C. C. & Cantley, L. C. Regulation of mTORC1 by PI3K signaling. Trends Cell Biol. 25, 545–555 (2015).
Sun, Y., Hedman, A. C., Tan, X. & Anderson, R. A. An unexpected role for PI4,5P2 in EGF receptor endosomal trafficking. Cell Cycle 12, 1991–1992 (2013).
Henmi, Y. et al. PtdIns4KIIα generates endosomal PtdIns(4)P and is required for receptor sorting at early endosomes. Mol. Biol. Cell 27, 990–1001 (2016).
Sun, Y., Hedman, A. C., Tan, X., Schill, N. J. & Anderson, R. A. Endosomal type Iγ PIP 5-kinase controls EGF receptor lysosomal sorting. Dev. Cell 25, 144–155 (2013).
Tan, X. et al. LAPTM4B is a PtdIns(4,5)P2 effector that regulates EGFR signaling, lysosomal sorting, and degradation. EMBO J. 34, 475–490 (2015).
Tan, X., Thapa, N., Choi, S. & Anderson, R. A. Emerging roles of PtdIns(4,5)P2–beyond the plasma membrane. J. Cell Sci. 128, 4047–4056 (2015).
Tan, X., Thapa, N., Liao, Y., Choi, S. & Anderson, R. A. PtdIns(4,5)P2 signaling regulates ATG14 and autophagy. Proc. Natl Acad. Sci. USA 113, 10896–10901 (2016).
Chen, Y. H. et al. Asymmetric PI3K activity in lymphocytes organized by a PI3K-mediated polarity pathway. Cell Rep. 22, 860–868 (2018).
Wallroth, A. & Haucke, V. Phosphoinositide conversion in endocytosis and the endolysosomal system. J. Biol. Chem. 293, 1526–1535 (2018).
Zewe, J. P. et al. Probing the subcellular distribution of phosphatidylinositol reveals a surprising lack at the plasma membrane. J. Cell Biol. 219, 201906127 (2020).
Pemberton, J. G. et al. Defining the subcellular distribution and metabolic channeling of phosphatidylinositol. J. Cell Biol. 219, 201906130 (2020).
Prinz, W. A., Toulmay, A. & Balla, T. The functional universe of membrane contact sites. Nat. Rev. Mol. Cell Biol. 21, 7–24 (2020).
Balla, T., Kim, Y. J., Alvarez-Prats, A. & Pemberton, J. Lipid dynamics at contact sites between the endoplasmic reticulum and other organelles. Annu. Rev. Cell Dev. Biol. 35, 85–109 (2019).
Ramkumar, A., Jong, B. Y. & Ori-McKenney, K. M. ReMAPping the microtubule landscape: how phosphorylation dictates the activities of microtubule-associated proteins. Dev. Dyn. 247, 138–155 (2018).
Gao, Y. L. et al. Tau in neurodegenerative disease. Ann. Transl. Med. 6, 175 (2018).
Iqbal, K., Liu, F. & Gong, C. X. Tau and neurodegenerative disease: the story so far. Nat. Rev. Neurol. 12, 15–27 (2016).
Xie, C. & Miyasaka, T. The role of the carboxyl-terminal sequence of tau and MAP2 in the pathogenesis of dementia. Front. Mol. Neurosci. 9, 158 (2016).
Thapa, N., Choi, S., Tan, X., Wise, T. & Anderson, R. A. Phosphatidylinositol phosphate 5-kinase iγ and phosphoinositide 3-kinase/Akt signaling couple to promote oncogenic growth. J. Biol. Chem. 290, 18843–18854 (2015).
Blackwell, J. R. & Horgan, R. A novel strategy for production of a highly expressed recombinant protein in an active form. FEBS Lett. 295, 10–12 (1991).
Samora, C. P. et al. MAP4 and CLASP1 operate as a safety mechanism to maintain a stable spindle position in mitosis. Nat. Cell Biol. 13, 1040–1050 (2011).
Thapa, N. et al. Phosphoinositide signaling regulates the exocyst complex and polarized integrin trafficking in directionally migrating cells. Dev. Cell 22, 116–130 (2012).
Costes, S. V. et al. Automatic and quantitative measurement of protein–protein co-localization in live cells. Biophys. J. 86, 3993–4003 (2004).
Choi, S., Chen, M., Cryns, V. L. & Anderson, R. A. A nuclear phosphoinositide kinase complex regulates p53. Nat. Cell Biol. 21, 462–475 (2019).
Asmari, M., Ratih, R., Alhazmi, H. A. & El Deeb, S. Thermophoresis for characterizing biomolecular interaction. Methods 146, 107–119 (2018).
Acknowledgements
We thank J. Feltenberger and L. Rodenkirch for technical support, and A. Rapraeger for the constructive comments and suggestions on the manuscript. This work is supported by NIH Grant RO1GM57549 and NIH Grant R35GM134955 to R.A.A. at the University of Wisconsin–Madison.
Author information
Authors and Affiliations
Contributions
N.T., M.C., H.T.H., S.C., T.W. and R.A.A. designed and discussed experiments. N.T., M.C., H.T.H., S.C. and T.W performed experiments. N.T., M.C., H.T.H., S.C., T.W. and R.A.A. discussed and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Colocalization of Activated Akt and PI3,4,5P3 with Endosomal Compartments.
a, Spatial localization of activated Akt with endosomal vesicles. MDA-MB-231 cells stimulated with EGF were immunostained with antibodies for pAkt and clathrin (CHC) or early endosome antigen 1 (EEA1) or transferrin receptor (TFR). The colocalization of pAkt with CHC/EEA1/TFR was quantified in unstimulated vs EGF stimulated cells by Pearson’s correlation coefficient (Pearson’s r). Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. b, c, Abrogation of EGF stimulated Akt activation and PI3,4,5P3 generation by PI3K inhibitors. MDA-MB-231 cells were treated with wortmannin or LY294002 compound before EGF stimulation. Cells were harvested and processed for immunoblotting or immunostaining using antibodies specific for activated Akt or PI3,4,5P3. Activated Akt and PI3,4,5P3 levels were quantified. Scale bar, 5 μm; Error bars denote mean±SD; n=4 independent experiments (b), n=30 cells from representative experiments (c). d, e, f, Examination of the activation level of GFP-tagged full- length Akt1 and their localization upon EGF stimulation. Cos-7 cells transiently transfected with GFP-tagged full-length Akt1 were pre-treated with PI3K inhibitors before EGF stimulation. Localization of GFP-tagged full-length Akt1 with different endosomal vesicles were quantified. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. g, Detection of PI3,4,5P3 in EGF stimulated cells by three different antibodies specific to PI3,4,5P3. Scale bar: 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments.
Extended Data Fig. 2 Detection of PI3,4,5P3 by PH domain GFP Reporter; Localization of activated EGFR upon EGF stimulation; Effect of Endocytosis Inhibitor in Colocalization of Activated Akt and PI3,4,5P3 with Clathrin Vesicles.
a, b, Spatial localization of GFP-tagged PH domain of Akt1 in EGF stimulated cells. Hs578T cells stably expressing GFP- tagged PH domain of Akt1 were stimulated with EGF for 5- minutes and colocalization of GFP signal with CHC/EEA1/TFR was quantified in unstimulated vs stimulated cells by Pearson’s r. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. c, d, Spatial localization of activated EGFR in EGF stimulated cells. MDA-MB-231 cells were stimulated with EGF and localization of activated EGFR at different time points was examined by immunostaining with an antibody specific for phospho-EGFR and tubulin. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. e, f, g, Effect of endocytosis inhibitor Dynasore in EGF stimulated activation level of Akt and PI3,4,5P3. The activation level of Akt in MDA-MB-231 cells pretreated with the Dynasore inhibitor before EGF stimulation was quantified by western blotting. Activated Akt and PI3,4,5P3 were also examined by immunostaining and quantified. The image shown is the representative images of multiple reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD; n=3. independent experiments (e), n=30 cells from representative experiments (g).
Extended Data Fig. 3 PI3Kα Vesicle Distribution in PI3Kα Mutant Expressing Cells; Specificity of PI3Kα Antibody Used and Distribution of Ectopically Expressed p110α.
a, Immunostaining of mutant PI3Kα expressing Cal51 and T47D cells with p85α or p110α specific antibodies. PI3Kα were distributed in small vesicle-like structures and its co- localization with microtubules was quantified by Pearson’s r. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. b, c, Specificity of PI3Kα antibody used. MDA-MB-231 cells transfected with siRNA for knockdown of p85α or p110α were used to demonstrate the loss of signals in knockdown cells by immunoblotting and immunofluorescence study. The immunoblot is the representative of reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. d, Localization of stably expressed haemagglutinin-tagged p110α WT or mutant forms in MDA-MB-231 cells. Wild type or mutant forms of p110α cloned into pWPT lentiviral vector were stably expressed into MAD-MB-231 cells by retroviral infection. Cells were processed for immunofluorescence study to examine the localization of ectopically expressed p110α using rabbit anti-haemagglutinin and rat anti-tubulin antibodies. The image shown is the representative images of multiple reproducible experiments. Scale bar: 5 μm. e, Effect of microtubule depolymerization in the distribution of PI3Kα. HeLa cells were either treated with DMSO or nocodazole (1 uM) for 3 hours before fixing the cells with paraformaldehyde. Cells were immunostained with anti-tubulin and anti-p85α or anti-p110α antibodies to examine the distribution of PI3Kα vesicles along microtubules. The image shown is the representative images of multiple reproducible experiments. Scale bar: 5 μm.
Extended Data Fig. 4 PI3Kα is Responsible for Agonist-stimulated Akt Activation and PI3,4,5P3 Generation in Internal Membrane Vesicles.
a, Effect of PI3Kα knockdown in EGF stimulated Akt activation. MDA-MB-231 cells were individually transfected with three different siRNAs specific for p110α. 48-72 hours post-transfection, cells were stimulated with EGF and activated Akt was examined by immunoblotting. Error bars denote mean±SD; n=3 independent experiments. b, c, d, Expression of HA-tagged p110α rescue the effect on endogenous p110α knockdown in EGF stimulated Akt activation. The siRNAs specific to 3’ UTR (sip110α III) wereused to knockdown p110α in mock or HA-p110α expressing cells. 48-72 hours post-transfection, cells were stimulated with EGF to examine the activation level of Akt by immunoblotting or immunofluorescence microscopy. Scale bar, 5 μm; Error bars denote mean±SD; n=4 independent experiments (b), n=30 cells from representative experiments (c). e, f, g, Effect of PI3Kα inhibitor in EGF stimulated activation level of Akt and PI3,4,5P3. MDA-MB-231 cells were pretreated with PI3K inhibitors (BKM120 or BYL719) before EGF stimulation. The activated Akt level was examined by immunoblotting. Similarly, activated Akt and PI3,4,5P3 co- localized with clathrin vesicles were examined by immunofluorescence microscopy and quantified. The image shown is the representative images of multiple reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD. n=3 independent experiments (e), n=30 cells from representative experiments (g). h, EGF stimulation promotes PI3Kα association with PIP5Kα and PIP5Kγ. PI3Kα was immunoprecipitated from MDA-MB-231 cells stimulated with EGF for 5 minutes and coimmunoprecipitation of PIP5Kα and PIP5Kγ was examined by immunoblotting using specific antibodies. The data shown is the representative of multiple reproducible experiments. The immunoblot shown is the representative of reproducible experiments. i, j, k Knockdown of PIP5Kα or PIP5Kγ affects EGF stimulated PI3,4,5P3 generation MDA-MB-231 cells were transfected with siRNAs for PIP5Kα or PIP5Kγ. 48-72 hours post-transfection, cells were stimulated with EGF and the activation level of Akt was examined by immunoblotting. Activated Akt and PI3,4,5P3 co-localized with clathrin vesicles (CHC) in control vs PIP5Kα or PIP5Kγ knockdown cells were examined by immunofluorescence microscopy by Pearson’s r. The image and immunoblot shown is the representatives of multiple reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD. n=30 cells from representative experiments.
Extended Data Fig. 5 Colocalization of MAP4 and PI3Kα in different cell types.
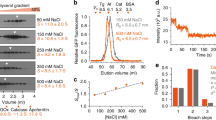
a, b, Coomassie staining of proteins coimmunoprecipitated with PI3Kα antibody (anti-p85α and anti-p110α antibodies used together) showed a distinct band above 170 kDa. Mass- spectrometry analysis of the isolated protein band revealed it as microtubule-associated protein 4 (MAP4). The image shown is the representative images of reproducible experiments. Arrow in image indicates the band of interest for mass spectrometry analysis. c, PI3Kα in small vesicles distribute along MAP4 that mimics microtubules in different cell types. Immunofluorescence study was performed in different cell types using mouse anti-MAP4 and rabbit anti-p110α or p85α antibodies. The image shown is the representative images of reproducible experiments. Scale bar, 5 μm. d, Amino acid alignment of MAP4 MTBD along with that of Tau and MAP2. All four MTBD repeats of MAP4 (MTBD I- MTBD IV) and that of Tau and MAP2 (other microtubule- associated proteins expressed in neuronal cells) show highly similar amino acid order and microtubule-binding motif. e, Representative MST binding affinity graphs for interaction between His-PI3Kα and GST-MAP4 proteins.
Extended Data Fig. 6 Colocalization of MAP4/IQGAP1 with EGFR, Effect of MAP4 Knockdown in Distribution of PI3Kα Vesicles along Microtubules in cells expressing WT MAP4 and MTBD Deletion Mutant.
a, b, PLA shows induced association of MAP4 and IQGAP1 and that they are localized with EGFR. MDA-MB-231 cells were stimulated with EGF and processed for MAP4-IQGAP1 PLA followed by immunostaining with EGFR. The image shown is the representative images of multiple reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. c, d, e, MAP4 knockdown demonstrated by immunofluorescence study and immunoblotting. Three different siRNAs were used individually to knockdown MAP4 in MDA-MB-231 cells. 48-72 hours after siRNA transfection, cells were processed for immunofluorescence study using an antibody specific to MAP4 and tubulin. MAP4 knockdown was also shown by immunoblotting. The image and blot shown is the representative of reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. f, g, Localization of PI3Kα vesicles along microtubules in WT and MTBD deletion mutant of MAP4 expressing cells after knocking down endogenous MAP4. As described above, siRNAs targeting 3’ UTR region of MAP4 were used to knockdown endogenous MAP4. Cells were processed for immunofluorescence study using antibodies specific for p110α and tubulin. Distribution of PI3Kα vesicles along microtubules was quantified. n=30 cells from representative experiments. h, i, Localization of ectopically expressed WT and MTBD deletion mutant of MAP4 after knockdown of endogenous MAP4. The siRNAs targeting the 3’ UTR region of MAP4 (siMAP4#3) were used to knockdown endogenous MAP4 in MDA-MB-231 cells expressing MAP4 WT or MTBD deletion mutant of MAP4. 48-72 hours post-transfection, cells were examined via immunofluorescence study using antibodies specific to MAP4 and tubulin. The colocalization of ectopically expressed MAP4 with tubulin was quantified by Pearson’s r. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments.
Extended Data Fig. 7 Effect of MAP4 Loss in Phosphorylation Level of EGFR and its Distribution in Endosomes.
a, b, Effect of MAP4 knockdown in endosomal localization of PI3Kα vesicles. siRNAs targeting the 3’ UTR region of MAP4 (siMAP4#3) were used to knockdown endogenous MAP4 in MDA-MB-231 cells expressing WT or the MTBD deletion mutant of MAP4. 48-72 hours post-transfection, cells were processed for immunofluorescence study using antibodies specific for endogenous p110α and clathrin or TFR. The quantification of the colocalization of p110α and CHC/TFR by Pearson’s r was shown in Fig. 5c. The images shown are the representative images of multiple reproducible experiments. Scale bar, 5 μm. c, d, The phosphorylation level of EGFR in MAP4 knockdown cells. 48-72 hours post siRNA transfection, cells were stimulated with EGF before examining the phosphorylation level of EGFR by immunoblotting and immunofluorescence microscopy. The immunoblot shown is the representative images of multiple reproducible experiments. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. e, Localization of activated EGFR in endosomes in MAP4 knockdown cells. 48-72 hours post siRNA transfection for MAP4 knockdown, cells were stimulated with EGF and processed for immunofluorescence study to examine the co- localization of activated EGFR with endosomes (EEA1 and TFR) by Pearson’s r. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments.
Extended Data Fig. 8 Effect of MAP4 Knockdown in Akt activation, Cell Proliferation, and Cell Invasion.
a, MAP4 loss impairs activation of Akt downstream of EGFR. Three different siRNAs were used individually to knockdown MAP4 in MDA-MB-231 cells. EGF induced Akt activation was analyzed by immunofluorescence microscopy. Scale bar, 5 μm; Error bars denote mean±SD; n=30 cells from representative experiments. b, c, MAP4 loss impairs Akt activation downstream of integrin receptors. MDA-MB-231 cells were detached from culture plates 72-hours post-siRNA transfection for MAP4 knockdown (siMAP4#1) and resuspended in serum-free medium. Cells were seeded into culture plates-coated with type I collagen before harvesting at different time points followed by immunoblotting or immunofluorescence microscopy using an antibody specific for activated Akt and tubulin. Scale bar: 5 μm. Error bars denote mean±SD; n=4 independent experiments (b), n=30 cells from representative experiments (c). d, MAP4 overexpression promotes Akt signaling. MDA-MB-231 cells ectopically overexpressing MAP4 or Mock were serum-starved overnight before stimulating with EGF. Cells were harvested at different time points and activation levels of Akt were examined by immunoblotting using activated Akt specific antibody. Error bars denote mean±SD; n=3 independent experiments. e, Effect of MAP4 knockdown in cell proliferation. MDA-MB- 468 and Cal51, both showing higher activation levels of Akt were transfected with siRNA for MAP4 knockdown. 72-96 hours post-siRNA transfection, cell numbers were manually quantified. Scale bar, 100 μm; Error bars denote mean±SD; n=3 independent experiments. f. g, h, Effect of MAP4 knockdown in cell invasion and cell migration. 48-72 hours post-siRNA transfection for MAP4 knockdown, cell invasion and scratch-wound healing for cell migration were performed. The image shown is the representative images of multiple reproducible experiments. Scale bar, 100 μm; Error bars denote mean±SD; n=9 fields from representative experiments (g), n=15 fields from representative experiments (h).
Supplementary information
Supplementary Tables 1 and 2
Defining the interaction of GST-fusion proteins of MAP4 with purified PI3Kα by MST.
Source data
Source Data Fig. 1
Unprocessed western blots
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Unprocessed western blots and gel images
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Unprocessed western blots
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Unprocessed western blots
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Unprocessed western blots
Source Data Fig. 6
Statistical source data
Source Data Fig. 7
Unprocessed western blots
Source Data Fig. 7
Statistical source data
Source Data Fig. 8
Unprocessed western blots
Source Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Unprocessed gel image
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Unprocessed western blots
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 8
Unprocessed western blots
Source Data Extended Data Fig. 8
Statistical source data
Rights and permissions
About this article
Cite this article
Thapa, N., Chen, M., Horn, H.T. et al. Phosphatidylinositol 3-kinase signalling is spatially organized at endosomal compartments by microtubule-associated protein 4. Nat Cell Biol 22, 1357–1370 (2020). https://doi.org/10.1038/s41556-020-00596-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00596-4
- Springer Nature Limited
This article is cited by
-
Non-catalytic role of phosphoinositide 3-kinase in mesenchymal cell migration through non-canonical induction of p85β/AP2-mediated endocytosis
Nature Communications (2024)
-
Endocytic vesicles act as vehicles for glucose uptake in response to growth factor stimulation
Nature Communications (2024)
-
DTX3L Accelerates Pancreatic cancer Progression via FAK/PI3K/AKT Axis
Biochemical Genetics (2024)
-
MAP4 acts as an oncogene and prognostic marker and affects radioresistance by mediating epithelial–mesenchymal transition in lung adenocarcinoma
Journal of Cancer Research and Clinical Oncology (2024)
-
CHKB-AS1 enhances proliferation and resistance to NVP-BEZ235 of renal cancer cells via regulating the phosphorylation of MAP4 and PI3K/AKT/mTOR signaling
European Journal of Medical Research (2023)