Abstract
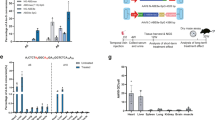
In contrast to traditional CRISPR–Cas9 homology-directed repair, base editing can correct point mutations without supplying a DNA-repair template. Here we show in a mouse model of tyrosinaemia that hydrodynamic tail-vein injection of plasmid DNA encoding the adenine base editor (ABE) and a single-guide RNA (sgRNA) can correct an A>G splice-site mutation. ABE treatment partially restored splicing, generated fumarylacetoacetate hydrolase (FAH)-positive hepatocytes in the liver, and rescued weight loss in mice. We also generated FAH+ hepatocytes in the liver via lipid-nanoparticle-mediated delivery of a chemically modified sgRNA and an mRNA of a codon-optimized base editor that displayed higher base-editing efficiency than the standard ABEs. Our findings suggest that adenine base editing can be used for the correction of genetic diseases in adult animals.





Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the paper and its Supplementary Information. The deep-sequencing data are available from the Sequence Read Archive (SRA) under accession number PRJNA513076 and PRJNA513291.
References
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Hess, G. T., Tycko, J., Yao, D. & Bassik, M. C. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 68, 26–43 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet.19, 770–788 (2018).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Chadwick, A. C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arterioscler. Thromb. Vasc. Biol. 37, 1741–1747 (2017).
Kim, K. et al. Highly efficient RNA-guided base editing in mouse embryos. Nat. Biotechnol. 35, 435–437 (2017).
Zong, Y. et al. Precise base editing in rice, wheat and maize with a Cas9–cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017).
Rossidis, A. C. et al. In utero CRISPR-mediated therapeutic editing of metabolic genes. Nat. Med. 24, 1513–1518 (2018).
Villiger, L. et al. Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat. Med. 24, 1519–1525 (2018).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2017).
Azuma, H. et al. Robust expansion of human hepatocytes in Fah −/−/Rag2 −/−/Il2rg −/− mice. Nat. Biotechnol. 25, 903–910 (2007).
Paulk, N. K. et al. Adeno-associated virus gene repair corrects a mouse model of hereditary tyrosinemia in vivo. Hepatology 51, 1200–1208 (2010).
Aponte, J. L. et al. Point mutations in the murine fumarylacetoacetate hydrolase gene: animal models for the human genetic disorder hereditary tyrosinemia type 1. Proc. Natl Acad. Sci. USA 98, 641–645 (2001).
Yin, H. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat. Biotechnol. 34, 328–333 (2016).
Song, C. Q. & Xue, W. CRISPR–Cas-related technologies in basic and translational liver research. Nat. Rev. Gastroenterol. Hepatol. 15, 251–252 (2018).
Shao, Y. et al. Cas9-nickase-mediated genome editing corrects hereditary tyrosinemia in rats. J. Biol. Chem. 293, 6883–6892 (2018).
Wang, D. et al. Cas9-mediated allelic exchange repairs compound heterozygous recessive mutations in mice. Nat. Biotechnol. 36, 839–842 (2018).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 6, 1258–1266 (1999).
Bateman, R. L. et al. Mechanistic inferences from the crystal structure of fumarylacetoacetate hydrolase with a bound phosphorus-based inhibitor. J. Biol. Chem. 276, 15284–15291 (2001).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Lee, H. K. et al. Targeting fidelity of adenine and cytosine base editors in mouse embryos. Nat. Commun. 9, 4804 (2018).
Liang, P. et al. Genome-wide profiling of adenine base editor specificity by EndoV-seq. Nat. Commun. 10, 67 (2019).
Zafra, M. P. et al. Optimized base editors enable efficient editing in cells, organoids and mice. Nat. Biotechnol. 36, 888–893 (2018).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Yin, H. et al. Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing. Nat. Biotechnol. 35, 1179–1187 (2017).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Xue, W. et al. Response and resistance to NF-kB inhibitors in mouse models of lung adenocarcinoma. Cancer Discov. 1, 236–247 (2011).
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007).
Zhu, L. J. et al. GUIDEseq: a Bioconductor package to analyze GUIDE-seq datasets for CRISPR–Cas nucleases. BMC Genomics 18, 379 (2017).
Yin, H. et al. Partial DNA-guided Cas9 enables genome editing with reduced off-target activity. Nat. Chem. Biol. 14, 311–316 (2018).
Acknowledgements
We thank C. Mello, P. Zamore, S. Wolfe and E. Sontheimer for discussions; J. Smith for editing the manuscript; M. Grompe (Oregon Health and Science University) for providing the Fah mice; Y. Liu and E. Kittler of the UMass Morphology and Deep Sequencing Cores for support. W.X. was supported by grants from the National Institutes of Health ((NIH) DP2HL137167, P01HL131471 and UG3HL147367), American Cancer Society (129056-RSG-16-093), the Lung Cancer Research Foundation, Hyundai Hope on Wheels, UMass CCTS and ALS Association. This work was supported by DARPA HR0011-17-2-0049; US NIH RM1 HG009490, R01 EB022376, U01 AI142756 and R35 GM118062; and HHMI (to D.R.L.), and R01 CA195787; K22 CA181280 (to L.E.D.). This work was supported in part by the Marble Center for Cancer Nanomedicine and a Cancer Center Support (core) grant P30-CA14051 from the National Cancer Institute. M.R. was supported by the HHMI Hanna H. Gray Fellowship. L.W.K. is an NSF Graduate Research Fellow and was supported NIH Training Grant T32 GM095450. H.Y. was supported by the National Natural Science Foundation of China 31871345, the Young Thousand Talented Program from Wuhan University and startup funding from Wuhan University.
Author information
Authors and Affiliations
Contributions
C.Q.S., T.J., M.R., D.R.L., H.Y. and W.X. designed the study. C.Q.S., T.J., M.R., L.H.R., L.W.K., M.P.Z., E.M.S., J.L.D., Y.C., L.J.Z., L.E.D. and D.G.A. performed experiments and analysed data. C.Q.S., T.J., H.Y. and W.X. wrote the manuscript with comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
D.R.L. is a consultant and co-founder of Editas Medicine, Pairwise Plants and Beam Therapeutics, which are companies that use genome editing. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and tables.
Supplementary Dataset 1
Guide-seq off-target sites.
Rights and permissions
About this article
Cite this article
Song, CQ., Jiang, T., Richter, M. et al. Adenine base editing in an adult mouse model of tyrosinaemia. Nat Biomed Eng 4, 125–130 (2020). https://doi.org/10.1038/s41551-019-0357-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-019-0357-8
- Springer Nature Limited
This article is cited by
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)
-
Engineering APOBEC3A deaminase for highly accurate and efficient base editing
Nature Chemical Biology (2024)
-
An adenine base editor variant expands context compatibility
Nature Biotechnology (2024)
-
Advances in functional lipid nanoparticles: from drug delivery platforms to clinical applications
3 Biotech (2024)
-
Genome editing: An insight into disease resistance, production efficiency, and biomedical applications in livestock
Functional & Integrative Genomics (2024)