Abstract
RNase P is the essential activity that performs the 5′ maturation of transfer RNA (tRNA) precursors. Beyond the ancestral form of RNase P containing a ribozyme, protein-only RNase P enzymes termed PRORP were identified in eukaryotes. In human mitochondria, PRORP forms a complex with two protein partners to become functional. In plants, although PRORP enzymes are active alone, we investigate their interaction network to identify potential tRNA maturation complexes. Here we investigate functional interactions involving the Arabidopsis nuclear RNase P PRORP2. We show, using an immuno-affinity strategy, that PRORP2 occurs in a complex with the tRNA methyl transferases TRM1A and TRM1B in vivo. Beyond RNase P, these enzymes can also interact with RNase Z. We show that TRM1A/TRM1B localize in the nucleus and find that their double knockout mutation results in a severe macroscopic phenotype. Using a combination of immuno-detections, mass spectrometry and a transcriptome-wide tRNA sequencing approach, we observe that TRM1A/TRM1B are responsible for the m22G26 modification of 70% of cytosolic tRNAs in vivo. We use the transcriptome wide tRNAseq approach as well as RNA blot hybridizations to show that RNase P activity is impaired in TRM1A/TRM1B mutants for specific tRNAs, in particular, tRNAs containing a m22G modification at position 26 that are strongly downregulated in TRM1A/TRM1B mutants. Altogether, results indicate that the m22G-adding enzymes TRM1A/TRM1B functionally cooperate with nuclear RNase P in vivo for the early steps of cytosolic tRNA biogenesis.



Similar content being viewed by others
Data availability
Sequences from TAIR v10 (27222 forward A. thaliana protein sequences) and Niben version 1.0.1 (57140 forward N. benthamiana protein sequences, Sol Genomics) databases were used here. Mass spectrometric data were deposited with the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifiers PXD040022 and PXD040029. Mim-tRNAseq data were deposited in the NCBI Gene Expression Omnibus under accession number GSE223876. Data supporting the findings of this work are available within the paper and its supplementary information files. A reporting summary for this Article is available as a supplementary information file. The clones and plant materials generated and analysed during the current study are available from the corresponding authors upon request. The source data underlying Figs. 2 and 3 as well as Extended Data Figs. 2, 4 and 7 are provided as a source data file. Source data are provided with this paper.
References
Chapeville, F. et al. On the role of soluble ribonucleic acid in coding for amino acids. Proc. Natl Acad. Sci. Usa. 48, 1086–1092 (1962).
Korostelev, A., Trakhanov, S., Laurberg, M. & Noller, H. F. Crystal structure of a 70S ribosome–tRNA complex reveals functional interactions and rearrangements. Cell 126, 1065–1077 (2006).
Altman, S. A view of RNase P. Mol. Biosyst. 3, 604–607 (2007).
Nickel, A. I. et al. Minimal and RNA-free RNase P in Aquifex aeolicus. Proc. Natl Acad. Sci. USA 114, 11121–11126 (2017).
Teramoto, T. et al. Minimal protein-only RNase P structure reveals insights into tRNA precursor recognition and catalysis. J. Biol. Chem. 297, 101028 (2021).
Göβringer, M., Wäber, N. B., Wiegard, J. C. & Hartmann, R. K. Characterization of RNA-based and protein-only RNases P from bacteria encoding both enzyme types. RNA 29, 376–391 (2023).
Howard, M. J., Lim, W. H., Fierke, C. A. & Koutmos, M. Mitochondrial ribonuclease P structure provides insight into the evolution of catalytic strategies for precursor-tRNA 50 processing. Proc. Natl Acad. Sci. USA 109, 16149–16154 (2012).
Gobert, A. et al. Structural insights into protein-only RNase P complexed with tRNA. Nat. Commun. 4, 1353 (2013).
Pinker, F. et al. Biophysical analysis of Arabidopsis protein-only RNase P alone and in complex with tRNA provides a refined model of tRNA binding. J. Biol. Chem. 292, 13904–13913 (2017).
Anantharaman, V. & Aravind, L. The NYN domains: novel predicted RNAses with a PIN domain-like fold. RNA Biol. 3, 18–27 (2006).
Gobert, A., Bruggeman, M. & Giegé, P. Involvement of PIN‐like domain nucleases in tRNA processing and translation regulation. IUBMB Life 71, 1117–1125 (2019).
Holzmann, J. et al. RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell 135, 462–474 (2008).
Gobert, A. et al. A single Arabidopsis organellar protein has RNase P activity. Nat. Struct. Mol. Biol. 17, 740–744 (2010).
Gutmann, B., Gobert, A. & Giegé, P. PRORP proteins support RNase P activity in both organelles and the nucleus in Arabidopsis. Genes Dev. 26, 1022–1027 (2012).
Lechner, M. et al. Distribution of ribonucleoprotein and protein-only RNase P in Eukarya. Mol. Biol. Evol. 32, 3186–3193 (2015).
Brillante, N. et al. Substrate recognition and cleavage-site selection by a single-subunit protein-only RNase P. Nucleic Acids Res. 44, 2323–2336 (2016).
Bouchoucha, A. et al. Determination of protein-only RNase P interactome in Arabidopsis mitochondria and chloroplasts identifies a complex between PRORP1 and another NYN domain nuclease. Plant J. 100, 549–561 (2019).
Howard, M. J. et al. Differential substrate recognition by isozymes of plant protein-only ribonuclease P. RNA 22, 782–792 (2016).
Kempenaers, M. et al. New archaeal methyltransferases forming 1-methyladenosine or 1-methyladenosine and 1-methylguanosine at position 9 of tRNA. Nucleic Acids Res. 38, 6533–6543 (2010).
Vilardo, E. et al. A Subcomplex of human mitochondrial RNase P Is a bifunctional methyltransferase-extensive moonlighting in mitochondrial tRNA biogenesis. Nucleic Acids Res. 40, 11583–11593 (2012).
Karasik, A., Fierke, C. A. & Koutmos, M. Interplay between substrate recognition, 5′ end tRNA processing and methylation activity of human mitochondrial RNase P. RNA 25, 1646–1660 (2019).
Bhatta, A., Dienemann, C., Cramer, P. & Hillen, H. S. Structural basis of RNA processing by human mitochondrial RNase P. Nat. Struct. Mol. Biol. 28, 713–723 (2021).
Funk, H. M. et al. Identification of the enzymes responsible for M2,2G and Acp3U formation on cytosolic tRNA from insects and plants. PLoS ONE 15, e0242737 (2020).
Chicois, C. et al. The UPF1 interactome reveals interaction networks between RNA degradation and translation repression factors in Arabidopsis. Plant J. 96, 119–132 (2018).
Canino, G. et al. Arabidopsis encodes four tRNase Z enzymes. Plant Physiol. 150, 1494–1502 (2009).
Glick, J. M., Averyhart, V. M. & Leboy, P. S. Purification and characterization of two tRNA-(guanine)-methyltransferases from rat liver. Biochim. Biophys. Acta 518, 158–171 (1978).
Ellis, S. R., Morales, M. J., Li, J. M., Hopper, A. K. & Martin, N. C. Isolation and characterization of the TRM1 locus, a gene essential for the N2,N2-dimethylguanosine modification of both mitochondrial and cytoplasmic tRNA in Saccharomyces cerevisiae. J. Biol. Chem. 261, 9703–9709 (1986).
Liu, J. & Strâby, K. B. The human tRNA(m22G26)dimethyltransferase: functional expression and characterization of a cloned hTRM1 gene. Nucleic Acids Res. 28, 3445–3451 (2000).
Takeda, H., Hori, H. & Endo, Y. Identification of Aquifex aeolicus tRNA (m2(2G26) methyltransferase gene. Nucleic Acids Res. 2, 229–230 (2002).
Pallan, P. S., Kreutz, C., Bosio, S., Micura, R. & Egli, M. Effects of N2,N2-dimethylguanosine on RNA structure and stability: crystal structure of an RNA duplex with tandem m2 2G:A pairs. RNA 14, 2125–2135 (2008).
Hooper, C. M., Castleden, I. R., Tanz, S. K., Aryamanesh, N. & Millar, A. H. SUBA4: the interactive data analysis centre for Arabidopsis subcellular protein locations. Nucleic Acids Res. 45, D1064–D1074 (2017).
Behrens, A., Rodschinka, G. & Nedialkova, D. D. High-resolution quantitative profiling of tRNA abundance and modification status in eukaryotes by mim-tRNAseq. Mol. Cell. 81, 1802–1815.e7 (2021).
O’Connor, J. P. & Peebles, C. L. In vivo pre-tRNA processing in Saccharomyces cerevisiae. Mol. Cell. Biol. 11, 425–439 (1991).
Hopper, A. K. Transfer RNA post-transcriptional processing, turnover, and subcellular dynamics in the yeast Saccharomyces cerevisiae. Genetics 194, 43–67 (2013).
Blewett, N. H. & Maraia, R. J. La involvement in tRNA and other RNA processing events including differences among yeast and other eukaryotes. Biochim. Biophys. Acta 1861, 361–372 (2018).
Pinker, F. et al. PPR proteins shed a new light on RNase P biology. RNA Biol. 10, 1457–1468 (2013).
Arimbasseri, A. G. et al. RNA polymerase III output is functionally linked to tRNA dimethyl-G26 modification. PLoS Genet. 11, e1005671 (2015).
Wilusz, J. E. Controlling translation via modulation of tRNA levels. Wiley Interdiscip. Rev. RNA 6, 453–470 (2015).
Steiner, R. E. & Ibba, M. Regulation of tRNA-dependent translational quality control. IUBMB Life 71, 1150–1157 (2019).
Awai, T. et al. Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure. J. Biol. Chem. 286, 35236–35246 (2011).
Teramoto, T. et al. Pentatricopeptide repeats of protein-only RNase P use a distinct mode to recognize conserved bases and structural elements of pre- tRNA. Nucleic Acids Res. 48, 11815–11826 (2020).
Li de la Sierra-Gallay, I., Mathy, N., Pellegrini, O. & Condon, C. Structure of the ubiquitous 3′ processing enzyme RNase Z bound to transfer RNA. Nat. Struct. Mol. Biol. 13, 376–377 (2006).
Fleurdépine, S., Deragon, J. M., Devic, M., Guilleminot, J. & Bousquet-Antonelli, C. A bona fide La protein is required for embryogenesis in Arabidopsis thaliana. Nucleic Acids Res. 35, 3306–3321 (2007).
Maraia, R. J., Mattijssen, S., Cruz-Gallardo, I. & Conte, M. R. The La and related RNA-binding proteins (LARPs): structures, functions, and evolving perspectives. Wiley Interdiscip. Rev. RNA https://doi.org/10.1002/wrna.1430 (2017).
Waltz, F. et al. Small is big in Arabidopsis mitochondrial ribosome. Nat. Plants 5, 106–117 (2019).
Kuhn, L., Vincent, T., Hammann, P. & Zuber, H. Exploring protein interactome data with IPinquiry: statistical analysis and data visualization by spectral counts. Methods Mol. Biol. 2426, 243–265 (2023).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson, J. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Acknowledgements
This work was supported by the French ‘Centre National de la Recherche Scientifique’, by the University of Strasbourg, by ANR research grants ‘MITRA, ANR-16-CE11-0024’, ‘DAMIA, ANR-20-CE11-0021’ and ‘PROPHAN, ANR-22-CE12-0008-01’ and by the LabEx consortium ‘MitoCross’. The MS instrumentations were funded by the IdEx ‘Equipement mi-lourd’ 2015 from the University of Strasbourg. We thank J. Zumsteg from the Plant Imaging Mass Spectrometry platform, ibmp, Strasbourg, S. Koechler from the Gene Expression Analysis platform, ibmp, Strasbourg. Finally, we thank D. D. Nedialkova, Max Planck Institute of Biochemistry, Martinsried, Germany for their precious advice on the adaptation of mim-tRNAseq technology to plant samples.
Author information
Authors and Affiliations
Contributions
M.A., M.B., A.G. and P.G. designed and coordinated the experiments. M.A., M.B., V.S., L.C., Y.-F.Q., C.S., V.C., P.H., J.C., P.W. and A.G. performed experiments and analysed results. M.A., M.B. and P.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Markos Koutmos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of double knock-out plants for PRORP2 and PRORP3 complemented by the insertion of a cDNA encoding PRORP2 fused with an HA tag.
a Gene models for AtPRORP2, AtPRORP3 and PRORP2-HA. Colour arrows and their names indicate primers used in b. T-DNA insertions, determined by Gutmann et al.14 are indicated by triangles. Expected sizes of the PCR products are indicated by the colour boxes under the gene models. b PCR products analysed on 1 % agarose gels. Colours in each image refer to the products indicated in gene models shown in a. WT plants are Columbia (Col-0) ecotype. PRORP2-HA was amplified from a cDNA representing the coding sequence without introns, which explains the difference in size in the PRORP2 gene PCR between WT plants and PRORP2-HA plants. These genotyping experiments were repeated three times.
Extended Data Fig. 2 Analysis by western blot using anti-HA antibodies of fractions obtained during co-immunoprecipitation (IP) experiments.
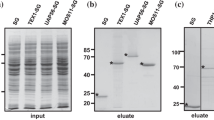
Analysis by western blot using anti-HA antibodies of fractions obtained during co-immunoprecipitation (IP) experiments using a flower buds extract from an Arabidopsis stable plant line expressing a PRORP2-HA fusion protein (HA) and control wild-type plants (WT). IP were performed with anti-HA beads. Lysate, Flow through (FT), Wash (W) and elution (EL) samples were run on a 8 % SDS-PAGE gel. Proteins were transferred to a PVDF membrane. M indicate molecular weight markers shown in kDa. The respective positions of the IgG heavy and light chains are indicated by grey arrows. A red arrow indicates the band corresponding to AtPRORP2-HA. This experiment was repeated three times.
Extended Data Fig. 3 RNase treatment results in reduced levels of TRM1A and TRM1B in anti-HA immunoprecipitations using PRORP2-HA as a bait.
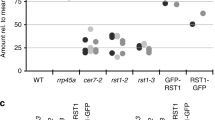
Histogram show the proportion of spectra corresponding to TRM1A and TRM1B, normalized to the total number of spectra detected in the respective IPs and expressed as a percentage of the number of spectra representing the PRORP2-HA bait. Results obtained with untreated samples are shown on the left, whereas results obtained after benzonase treatment are shown on the right.
Extended Data Fig. 4 TRM1A and TRM1B are both localized to the nucleus.
Arabidopsis protoplasts were isolated, transfected with constructs expressing TRM1A-CFP and YFP-TRM1B and visualized by confocal microscopy. CFP signal is shown in cyan and YFP signal in yellow. DAPI signal in blue is used as a control to identify nuclei, that is nucleoplasm and the autofluorescence of chlorophyll in purple shows chloroplasts. Scale bars represent 10 mm. These microscopy observations were repeated three times.
Extended Data Fig. 5 Assessment of mim-tRNAseq output quality.
a Table indicating the raw number of reads for the 6 libraries sequenced and the number of reads of sufficient quality, selected as described in the methods section, that were kept for mim-tRNAseq analysis. b Histogram indicating the proportion of reads corresponding to tRNAs in the respective libraries. Green bars represent the proportion of reads uniquely mapped to tRNA sequences. Grey bars represent the proportion of reads that did not map to tRNA sequences. This shows that between 70 and 80 % of reads from the respective libraries corresponded to tRNAs. c Average distribution of sequence lengths in the mim-tRNAseq libraries. Sequence lengths are indicated on the x-axis in base pairs and numbers of reads, in thousands, are indicated in the y-axis. Most reads had a length between 70 and 90 nucleotides, in accordance with tRNA and pre-tRNA molecules.
Extended Data Fig. 6 Analysis of chloroplastic tRNAs modifications in wild-type and TRM1A/B mutants.
Heat maps representing the frequency of mismatches between mim-tRNAseq reads and genomic sequences in wild-type and TRM1A/B double mutant samples, at each tRNA positions (1 to 76) in the 23 hierarchically clustered Arabidopsis chloroplast encoded tRNAs, revealing RNA modifications in individual tRNAs. S stands for stem, L for loop, Acc for acceptor, AC for anticodon V for variable region. Darker blue shade represents increased mismatch frequencies with 0 to 1 representing 0 to 100% mismatch frequencies.
Extended Data Fig. 7 RNA gel with 1 µg total RNA samples prepared from wild-type, TRM1 single mutants and TRM1A/B double mutants.
Electrophoresis was carried out in 17 % polyacrylamide and 7 M urea. RNA species were visualized by BET coloration under UV light. Molecular weight markers are shown on the left and their sizes indicated in nucleotides. A black bar represent the gel section containing tRNA and tRNA precursors. This analysis was repeated three times.
Extended Data Fig. 8 Histograms representing the sequence coverage for representative individual tRNA genes for the 3 wild-type and 3 TRM1A/B libraries analyzed by mim-tRNAseq.
Histograms representing the sequence coverage for the representative individual tRNA genes tRNA-Arg-TCT-1-1, tRNA-Ser-GCT-3-2, tRNA-Glu-CTC-1-1, tRNA-Cys-GCA-9-2, tRNA-Tyr-GTA-2-11 and tRNA-Pro-TGG-1-1 for the 3 wild-type and 3 TRM1A/B libraries analyzed by mim-tRNAseq. tRNA sequences are represented on the x-axis with S standing for stem, L for loop, Acc for acceptor, AC for anticodon V for variable region, while the number of reads are indicated on the y-axis. 5′ leader sequences are highlighted by red boxes. Likewise, reads corresponding to sequences containing either 3′ trailer sequences or CCA are also highlighted by red boxes.
Extended Data Fig. 9 TRM1A can form complexes with pre-tRNAs containing a G26 or a A26.
Size exclusion chromatograms of recombinant AtTRM1A in presence of Sc-tRNAPhe containing a G26, At-tRNAmtCys containing a A26 and a fragment of At-COX1 mRNA. Red traces correspond to RNA alone. Green traces correspond to RNA and AtTRM1A mixed at a molar ratio of 1:2, blues traces correspond to RNA and AtTRM1A mixed at a molar ratio of 1:10, orange trace to TRM1A alone and purple trace to molecular weight markers. M1 is thyroglobulin (670 kDa), M2 is g-globulin (158 kDa), M3 is ovalbumin (44 kDa), M4 is myoglobin (17 kDa) and M5 is vitamin B12 (1,35 kDa). Grey arrows represent peaks corresponding to AtTRM1A / pre-tRNA complexes. x-axis shows the size exclusion chromatography elution time while the y-axis shows the OD measured at 260 nm for RNA containing samples or 280 nm for protein samples.
Extended Data Fig. 10 Structural model for the interaction between Arabidopsis PRORP2, a pre-tRNA and TRM1A.
a PRORP2 experimental crystal structure is shown in brown shades, a representative pre-tRNA is shown in purple with the position of G26 highlighted. PRORP2 binding to tRNA is represented according to experimental biochemical and biophysical results9 and PRORP2 pre-tRNA cleavage site is shown by a red star. Arabidopsis TRM1A structure model was built with AlphaFold 2.050 and its docking to tRNA is represented according to the experimental biochemical analysis of Aquifex aeolicus TRM1 binding to tRNA40. TRM1A C-terminal domain is shown in pink, whereas its N-terminal catalytic domain is shown in light blue. TRM1 catalytic pocket is represented in yellow. b TRM1A model generated by AlphaFold is colored according to model confidence, that is with dark blue part of the protein including the catalytic domain and the protein regions shown to interact with tRNA in Aquifex aeolicus40 having very high model confidence. AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Some regions below 50 pLDDT may be unstructured.
Supplementary information
Supplementary Table
List Supplementary Tables 1a–g, 2, 3 and 4.
Source data
Source Data
Unprocessed microscopy images, gels, and hybridized blots for Figs. 2 and 3, and Extended Data Figs. 2, 4 and 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arrivé, M., Bruggeman, M., Skaltsogiannis, V. et al. A tRNA-modifying enzyme facilitates RNase P activity in Arabidopsis nuclei. Nat. Plants 9, 2031–2041 (2023). https://doi.org/10.1038/s41477-023-01564-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01564-0
- Springer Nature Limited