Abstract
POLRMT (RNA polymerase mitochondrial) is essential for transcription of mitochondrial genome encoding components of oxidative phosphorylation process. The current study tested POLRMT expression and its potential function in osteosarcoma (OS). The Cancer Genome Atlas (TCGA) cohorts and Gene Expression Profiling Interactive Analysis (GEPIA) database both show that POLRMT transcripts are elevated in OS tissues. In addition, POLRMT mRNA and protein levels were upregulated in local OS tissues as well as in established and primary human OS cells. In different OS cells, shRNA-induced stable knockdown of POLRMT decreased cell viability, proliferation, migration, and invasion, whiling inducing apoptosis activation. CRISPR/Cas9-induced POLRMT knockout induced potent anti-OS cell activity as well. Conversely, in primary OS cells ectopic POLRMT overexpression accelerated cell proliferation and migration. In vivo, intratumoral injection of adeno-associated virus-packed POLRMT shRNA potently inhibited U2OS xenograft growth in nude mice. Importantly, levels of mitochondrial DNA, mitochondrial transcripts and expression of respiratory chain complex subunits were significantly decreased in U2OS xenografts with POLRMT shRNA virus injection. Together, POLRMT is overexpressed in human OS, promoting cell growth in vitro and in vivo. POLRMT could be a novel therapeutic target for OS.
Similar content being viewed by others
Introduction
Osteosarcoma (OS) is the most common bone malignancy among children and adolescents [1]. OS comprises 2.4% of all malignancies in pediatric patients [2,3,4]. It arises from the primitive transformed cells of mesenchymal origin [5, 6]. Under the current treatments, including systemic chemotherapy, radiation and local control surgery, the 5-year overall survival of OS is close to 70% [1]. For patients with metastatic and recurrent OS, the prognosis is often poor [5, 6]. One possibility is that OS cells are commonly resistant to chemotherapeutic agents [5, 6]. In recent years, molecularly-targeted therapies have drawn significant attentions for OS, and several of these agents are currently under different stages of clinical studies [6,7,8,9,10].
RNA polymerase mitochondrial (POLRMT) is a key component of mitochondrial transcription machinery [11, 12]. POLRMT and two other mitochondrial proteins, mitochondrial transcription factor A (TFAM) and transcription factor B2 (TFB2M), are essential for initiation and progression of promoter-specific mitochondrial DNA (mtDNA) transcription [11, 12]. Furthermore, the synthesis of RNA primers is also dependent on POLRMT, which is required for replication of mtDNA [13, 14].
Recent studies have proposed a cancer-promoting activity by POLRMT. In acute myeloid leukemia (AML) cells shRNA-induced knockdown of POLRMT reduced oxidative phosphorylation (OXPHOS), cell growth and viability [15]. Furthermore, 2-C-methyladenosine, a POLRMT inhibitor, exerted potent anti-tumor response in AML cells [15]. POLRMT and mitochondrial biogenesis levels are significantly elavated in breast cancer cells, important for cancer cell growth, and autophagy resistance [16, 17]. Bonekamp et al., have recently developed the first-in-class specific POLRMT inhibitors to block mitochondrial transcription [18]. These POLRMT inhibitors potently suppressed cancer cell growth in mice [18]. In the present study we showed that POLRMT is overexpressed in human OS, promoting OS cell growth in vitro and in vivo.
Results
POLRMT overexpression in human OS
Gene Expression Profiling Interactive Analysis (GEPIA) database was consulted to examine POLRMT mRNA expression profile in human OS. GEPIA parameters were set as follows: (|Log2FC| Cutoff: 1; P value Cutoff:0.05). Results showed that POLRMT mRNA transcripts in OS tumor tissues (n = 262) were significantly higher than those in normal bone tissues (n = 2) (Fig. 1A). We also downloaded the raw mRNA expression data of sarcoma from TCGA database (https://portal.gdc.cancer.gov/). A total of 264 samples were collected, including two normal samples and 262 sarcoma samples. Results showed that POLRMT mRNA upregulation in OS tumor tissues (P < 0.05 versus normal bone tissues) (Fig. 1B).
GEPIA database (A) and TCGA cohorts (B) showed the relative POLRMT mRNA expression in listed osteosarcoma (OS) tumor tissues and normal tissues. Expression of POLRMT mRNA and protein in OS tumor tissues (“T”) and the surrounding normal bone tissues (“N”) of eleven (11) OS patients was shown, with results quantified (C–E). Expression of POLRMT mRNA and protein in established OS cell lines (U2OS and MG63), primary human OS cells (pOS-1 and pOS-2, derived two patients), as well as in primary human osteoblasts (“Ob”) was shown, with results quantified (F, G). Data were presented as mean ± standard deviation (SD). *P < 0.05 versus normal tissues/“Ob” cells.
To confirm the bioinformatics observation, we examined POLRMT expression in local human OS tissues. OS tumor tissues (“T”) and surrounding normal bone tissues (“N”) were derived from a total of eleven (n = 11) primary OS patients [19]. qRT-PCR assay results, Fig. 1C, demonstrated that POLRMT mRNA levels in OS tumor tissues were significantly higher than those in normal bone tissues (P < 0.05, Fig. 1C). Analyzing POLRMT protein expression, by Western blotting assays, showed that POLRMT protein levels were elevated as well in OS tissues (Fig. 1D, E). POLRMT blotting data of four represent patients, Patient-1 to Patient-4, were presented (Fig. 1D). Quantification analyses integrating Western blotting data from all 11 sets of human tissues were presented as well (Fig. 1E).
We also tested POLRMT expression in human OS cells. As shown, POLRMT mRNA (Fig. 1F) and protein (Fig. 1G) levels in established OS cell lines (U2OS and MG63) and primary human OS cells (pOS-1 and pOS-2, derived from two different patients [19]) were significantly higher than those in the primary human osteoblasts (“Ob”) (Fig. 1F, G). These results show that POLRMT is overexpressed in human OS tissues and different human OS cells.
POLRMT silencing produces significant anti-cancer activity in OS cells
Next shRNA strategy was employed to silence POLRMT in OS cells. As described, two lentiviral shRNAs, targeting non-overlapping sequences of POLRMT, sh-POLRMT-1# and sh-POLRMT-2#, were separately transduced to pOS-1 primary cancer cells. Following selection by puromycin, stable cells were established. Analyzing POLRMT mRNA expression, using qRT-PCR assays, confirmed that each of the applied shRNA resulted in over 95% knockdown of POLRMT mRNA in pOS-1 cells (Fig. 2A). POLRMT protein levels were dramatically reduced as well in POLRMT shRNA-expressing pOS-1 cells (Fig. 2B). Testing cell viability, using CCK-8 assays, demonstrated that POLRMT shRNA significantly decreased the viability of pOS-1 cells (Fig. 2C). Moreover, clonogenicity assay results, Fig. 2D, showed that the number of viable pOS-1 cell colonies was largely decreased after POLRMT silencing. Evidenced by decreased EdU-positive nuclei ratio (Fig. 2E), we demonstrated that POLRMT shRNA inhibited pOS-1 cell proliferation. Further experiment results showed that POLRMT silencing potently suppressed pOS-1 cell migration and invasion, which were tested by “Transwell” (Fig. 2F) and “Matrigel Transwell” (Fig. 2G) assays, respectively. As expected, the scramble control shRNA, c-sh, failed to significantly alter POLRMT expression (Fig. 2A, B) and pOS-1 cell behaviors (Fig. 2C–G).
The primary human OS cells (pOS-1 and pOS-2, derived from two different patients) or established OS cell lines (U2OS and MG63), were cultured in polybrene-containing medium and infected with POLRMT shRNA lentivirus (sh-POLRMT-1#/sh-POLRMT-2#) or scramble control shRNA lentivirus (c-sh); Stable cells were established after puromycin selection. Expression of POLRMT mRNA and protein was shown (A, B, H); Cells were further cultured for applied time periods, cell viability (CCK-8 OD, C, I), colony formation (D) and proliferation (by recording nuclear EdU ratios, E, J) as well as cell migration and invasion (“Transwell” assays, F, G, K) were tested by the described methods, with results quantified. Expression of listed proteins was quantified and normalized to the loading control (B). “Pare” stands for the parental control cells. Error bars stand for mean ± standard deviation (SD, n = 5). *P < 0.05 versus “Pare” cells or “c-sh” cells. Experiments in this figure were repeated five times. Scale bar = 100 μm (E, F).
In addition, the sh-POLRMT-1# lentivirus was added to another primary human OS cells (pOS-2) and established OS cell lines (U2OS and MG63). Stable cells were again established after puromycin selection. Results from qRT-PCR assays demonstrated that sh-POLRMT-1# resulted in 80–90% silencing of POLRMT mRNA in the primary and established OS cells (Fig. 2H). In these OS cells, POLRMT shRNA potently inhibited cell viability, proliferation and migration, tested by CCK-8 (Fig. 2I), nuclear EdU staining (Fig. 2J) and “Transwell” (Fig. 2K) assays, respectively. These results showed that POLRMT shRNA inhibited OS cell viability, proliferation, migration, and invasion.
POLRMT silencing provokes apoptosis in OS cells
Experiments were performed to test whether POLRMT silencing could provoke apoptosis activation in OS cells. As shown in Fig. 3A, the relative caspase-3 activity was significantly increased in POLRMT shRNA-expressing pOS-1 primary cancer cells. Moreover, cleavages of caspase-3 and its substrate PARP (poly ADP-ribose polymerase) were significantly increased in POLRMT-silenced pOS-1 cells (Fig. 3B). Indicating DNA damage, we found that single strand DNA (ssDNA) contents were significantly increased in stable pOS-1 cells with POLRMT shRNA (Fig. 3C). In addition, JC-1 green monomer accumulation was detected in pOS-1 cells expressing the POLRMT shRNA (Fig. 3D), indicating mitochondrial depolarization [20, 21]. To confirm apoptosis activation we found that TUNEL-positive nuclei ratio was significantly increased in POLRMT-silenced pOS-1 cells (Fig. 3E). These results confirmed that POLRMT silencing by targeted shRNA provoked caspase-apoptosis activation in pOS-1 cells. The scramble control shRNA (c-sh), unsurprisingly, failed to induce caspase activation and apoptosis in pOS-1 cells (Fig. 3A–E).
The primary human OS cells (pOS-1 and pOS-2, derived from two different patients) or established OS cell lines (U2OS and MG63), were cultured in polybrene-containing medium and then infected with applied POLRMT shRNA lentivirus (sh-POLRMT-1#/sh-POLRMT-2#) or scramble control shRNA lentivirus (c-sh); Stable cells were established after puromycin selection; Cells were further cultured for applied time periods, the relative caspase-3 activity (A, F), caspase-3-PARP cleavages (B), single strand DNA (ssDNA) contents (ELISA assays, C) and mitochondrial depolarization (by testing JC-1 green monomer intensity, D, G) were tested by the listed assays; Cell apoptosis was tested by recording nuclear TUNEL ratio (E, H). The primary human osteoblasts (“Ob”) were cultured in polybrene-containing medium and then infected with applied POLRMT shRNA lentivirus (sh-POLRMT-1#/sh-POLRMT-2#) or scramble control shRNA lentivirus (c-sh); Stable cells were established after puromycin selection; Expression of POLRMT mRNA and listed proteins was shown (I); Cells were further cultured for 96 h, with cell viability and apoptosis tested by CCK-8 (J) and the nuclear TUNEL staining (K) assays, respectively. “Pare” stands for the parental control cells. Error bars stand for mean ± standard deviation (SD, n = 5). *P < 0.05 versus “Pare” cells or “c-sh” cells. Experiments in this figure were repeated five times. Scale bar = 100 μm (D, E).
In the primary pOS-2 cells and OS cell lines (U2OS and MG63), POLRMT silencing by shRNA (sh-POLRMT-1#) induced similar results, causing caspase-3 activation (Fig. 3F) and mitochondrial depolarization (JC-1 green monomer accumulation, Fig. 3G). Significant apoptosis activation, evidenced by increased TUNEL-positive nuclei ratio, was detected as well in the OS cells expressing POLRMT shRNA (Fig. 3H). Therefore, POLRMT silencing provoked apoptosis activation in OS cells.
In the primary human osteoblasts (“Ob”), stable transfection of the lentiviral POLRMT shRNA (sh-POLRMT-1#/-2#) resulted in robust silencing of POLRMT mRNA and protein (Fig. 3I). Significantly, POLRMT shRNA only resulted in minor but significant viability (CCK-8 OD) reduction (Fig. 3J) and apoptosis activation (tested by the TUNEL-positive nuclei ratio increase, Fig. 3K) in the human osteoblasts.
CRISPR/Cas9-induced POLRMT knockout produces significant anti-cancer activity in OS cells
To further support the function of POLRMT in OS cells, we next utilized CRISPR/Cas9 strategy to KO POLRMT in OS cells. As described, the Cas9-expressing stable pOS-1 cells were further transfected with lenti-POLRMT-sgRNA construct. Following puromycin selection and POLRMT KO screening, the single stable POLRMT KO pOS-1 cells were established: ko-POLRMT cells. qRT-PCR and Western blotting assays were performed to examine POLRMT expression, showing complete POLRMT KO in ko-POLRMT cells (Fig. 4A, B). In pOS-1 cells CRISPR/Cas9-induced POLRMT KO potently inhibited cell proliferation (decreased EdU-positive nuclei ratio, Fig. 4C), migration (results quantified in Fig. 4D) and invasion(Fig. 4E). Moreover, mitochondrial depolarization (tested by JC-1 green monomer intensity increase, Fig. 4F) and apoptosis activation (tested by the TUNEL-positive nuclei ratio increase, Fig. 4G) were detected in ko-POLRMT pOS-1 cells. As expected, the control lenti-sgRNA construct, koC, failed to significantly alter POLRMT expression (Fig. 4A, B) and pOS-1 cell function (Fig. 4C–G).
The Cas9-expressing stable pOS-1 cells were further transfected with the lenti-POLRMT-sgRNA construct (ko-POLRMT) or the empty vector (“koC”), stable cells were established. Expression of POLRMT mRNA and protein was shown (A, B); Cells were further cultured for applied time periods, cell proliferation (nuclear EdU staining assays, C), migrated and invaded cell number (D, E), as well as mitochondrial depolarization (by testing JC-1 green monomer intensity, F) and cell apoptosis (by recording TUNEL-positive nuclei ratio, G) were tested. “Pare” stands for the parental control cells. Error bars stand for mean ± standard deviation (SD, n = 5). *P < 0.05 versus “Pare” cells. Experiments in this figure were repeated five times. Scale bar = 100 μm (C).
Ectopic overexpression of POLRMT accelerates OS cell proliferation, migration, and invasion
Since POLRMT silencing or KO induced robust anti-OS cell activity, we proposed that ectopic overexpression of POLRMT might exert opposite function. Therefore, a lentiviral construct encoding full-length POLRMT cDNA was stably transduced to pOS-1 cells. Via selection by puromycin two stable cell lines with PLORMT overexpression, oe-POLRMT-sL1 and oe-POLRMT-sL2, were established. Testing POLRMT expression, via qRT-PCR and Western blotting assays, showed that PLORMT mRNA (Fig. 5A) and protein (Fig. 5B) levels were significantly increased in oe-POLRMT pOS-1 cells (P < 0.05 versus control cells with empty vector/“EV”). Ectopic overexpression of POLRMT accelerated OS cell proliferation, as the EdU-positive nuclei ratio was significantly increased in oe-POLRMT-sL1 cells and oe-POLRMT-sL2 cells (Fig. 5C). Moreover, POLRMT overexpression in pOS-1 cells augmented cell migration and invasion, tested by “Transwell” (Fig. 5D) and “Matrigel Transwell” (Fig. 5E) assays, respectively.
POLRMT-overexpressed pOS-1 cells (oe-POLRMT-sL1 and oe-POLRMT-sL2, two stable cell lines) or the control pOS-1 cells with empty vector (EV) were established; POLRMT mRNA and protein were tested (A, B); Cells were further cultured for applied time periods, cell proliferation (nuclear EdU staining assays, C), migration and invasion (D, E, “Transwell” assays) were tested, with results quantified. Error bars stand for mean ± standard deviation (SD, n = 5). *P < 0.05 versus “EV” cells. Experiments in this figure were repeated five times.
In OS cells mtDNA contents, mitochondrial transcripts and expression respiratory chain complex subunits are decreased after POLRMT depletion
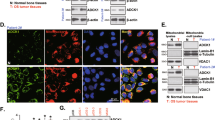
Studies have implied that POLRMT silencing or inhibition would result in decreases in mtDNA copies and mitochondrial transcripts [18]. As shown, in pOS-1 cells with POLRMT shRNA (sh-POLRMT-1#, see Figs. 2 and 3) or the lenti-POLRMT-sgRNA construct (ko-POLRMT, see Fig. 4), mtDNA contents were significantly decreased (P < 0.05 versus parental control cells, Fig. 6A). Levels of mitochondrial transcripts, ND1 and CYTB, were decreased as well after POLRMT silencing or KO (Fig. 6B). Moreover, mRNA levels of respiratory chain complex subunits, NDUFB8 and UQCRC2 [18], were reduced in sh-POLRMT and ko-POLRMT pOS-1 cells (Fig. 6B). Conversely, mtDNA contents (Fig. 6C) as well as mRNA levels of ND1, CYTB, NDUFB8 and UQCRC2 (Fig. 6D) were significantly increased in POLRMT-overexpressed pOS-1 cells: oe-POLRMT-sL1 and oe-POLRMT-sL1 (P < 0.05 versus “EV” control cells).
Stable pOS-1 cells with sh-POLRMT-1# (“sh-POLRMT”), the lenti-POLRMT-sgRNA construct (“ko-POLRMT”) or scramble control shRNA plus the empty vector (“c-sh+koC”), as well as POLRMT-overexpressed pOS-1 cells (oe-POLRMT-sL1 and oe-POLRMT-sL2, two stable cell lines) or the control cells with empty vector (EV), were established; Levels of mtDNA (A, C) and listed mRNAs (qRT-PCR assays, B, D) were examined. “Pare” stands for parental control cells. Data were presented as mean ± standard deviation (SD, n = 5). *P < 0.05 versus “Pare”/“EV” cells. The experiments were repeated five times with similar results obtained.
POLRMT silencing inhibits U2OS xenograft growth in nude mice
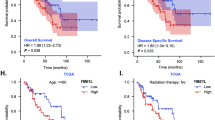
We tested the potential effect of POLRMT on OS cell growth in vivo. As described, U2OS cells, at 5 × 106 cells per mouse, were s.c. injected to the flanks of nude mice. Within 25 days, U2OS xenograft tumors were established with the volume close to 100 mm3 (“Day-0”). At 10 mice per group (n = 10), U2OS xenograft-bearing nude mice were randomly assigned into two groups. The treatment group received intratumoral injection of AAV-POLRMT shRNA (“AAV-sh-POLRMT”), daily for 6 consecutive days. The control group mice, on the other hand, received intratumoral injection of AAV scramble control shRNA (“AAV-c-sh”). In Fig. 7A, the tumor growth curve results showed that the growth of U2OS xenografts with AAV-sh-POLRMT injection was significantly slower than control xenografts with AAV-c-sh injection. The estimated daily tumor growth was calculated using the following formula: (Tumor volume at Day-42—Tumor volume at Day-0)/42. Results, Fig. 7B, demonstrated that intratumoral injection of AAV-sh-POLRMT potently inhibited U2OS xenograft growth in mice. At experimental Day-42, all xenograft tumors were carefully separated and weighted individually. As shown, the AAV-sh-POLRMT-injected U2OS xenografts were significantly lighter than control xenografts with AAV-c-sh injection (Fig. 7C). The mice body weights, shown in Fig. 7D, were not significantly different between the two groups. These results showed that POLRMT silencing inhibited U2OS xenograft growth in nude mice.
U2OS xenograft-bearing nude mice were randomly assigned into two groups (ten mice per group, n = 10), subject to intratumoral injection of AAV-POLRMT shRNA (“AAV-sh-POLRMT”) or AAV scramble control shRNA (“AAV-c-sh”). Viruses were injected daily for 6 consecutive days. Tumor volumes (A) and mice body weights (D) were recorded every 6 days (“Day-0 to Day-42”); Estimated daily tumor growth was calculated using the described formula (B). At Day-42 U2OS xenografts were isolated and individually weighted (C). At Day-6 and Day-12, one xenograft of each group was isolated (total four xenografts). Western blotting (E) and qRT-PCR (G) assays were employed to examine listed genes in fresh tumor tissue lysates, with mtDNA contents tested as well (F). Data were presented as mean ± standard deviation (SD). *P < 0.05 versus “AAV-c-sh” group.
At Day-6 and Day-12, one U2OS xenograft tumor from each group was isolated. The fresh tissue lysates from the four tumors were achieved and analyzed. Western blotting assay results, Fig. 7E, confirmed that POLRMT protein expression was significantly downregulated in AAV-sh-POLRMT-injected U2OS xenografts. In addition, mtDNA contents were decreased (Fig. 7F). Moreover, the mitochondrial transcripts (ND1 and CYTB) as well as mRNA expression of respiratory chain complex components (NDUFB8 and UQCRC2) were inhibited in U2OS xenografts with AAV-sh-POLRMT injection (Fig. 7G). Therefore, in line with the in vitro findings, POLRMT silencing decreased mtDNA contents, mitochondrial transcripts and expression of respiratory chain complex components in U2OS xenografts.
Discussion
OXPHOS generates ATP by transport of electrons to electron transport chain, the latter is composed of a number of transmembrane protein complexes in the mitochondrial inner membrane [22, 23]. Recent studies have implied that OXPHOS levels are upregulated in a number of different cancers, including leukemia, lymphomas, breast cancer, pancreatic cancer, certain melanomas, and endometrial carcinoma [22, 23]. OXPHOS inhibitors could therefore be utilized to target these cancers, showing promising anti-cancer efficiency in preclinical studies [22, 23]. Recent studies have implied that mtDNA is required for tumorigenesis and progression for cancer cells, mediating resistance to cytotoxic drugs [24]. Cancer cell growth could be largely diminished with mtDNA depletion [24, 25]. mtDNA transcription arrest could therefore result in significant anti-tumor activity [26,27,28].
POLRMT is essential for mtDNA transcription and OXPHOS [26,27,28], represents as a novel therapeutic target for human cancer [15, 18, 29]. POLRMT overexpression in AML is associated increased mtDNA copy number and lower overall survival [29]. In AML cells, POLRMT knockdown by shRNA reduced mitochondrial gene expression and inhibited OXPHOS and expression of respiratory chain complex subunits, causing significant cell death [15]. 2-C-methyladenosine (2-CM), a mitochondrial transcription inhibitor, resulted in significant cancer cell death [15]. A recent study by Bonekamp and colleagues developed a first-in-class POLRMT direct inhibitor, inhibitor of mitochondrial transcription 1 (IMT1). POLRMT inhibition by IMT1 induced robust cancer growth arrest [18]. Therefore, POLRMT inhibition or silencing shall impair mtDNA transcription and OXPHOS, resulting in significant cancer growth inhibition and cancer cell death [15, 18, 29].
Our results indicated that POLRMT could be an important therapeutic target of OS. GEPIA and TCGA database show that POLRMT transcripts are significantly elevated in human OS. Furthermore, POLRMT mRNA and protein are upregulated in local OS tissues as well as in established and primary human OS cells. But low expression is detected normal bone tissues and human osteoblasts. Oran et al., have also reported that POLRMT expression is elevated in U2OS cells, where it is a direct transcriptional target of the MYC oncoproteins [30].
In primary human OS cells and established cell lines, shRNA-induced knockdown of POLRMT largely inhibited cell viability, proliferation, migration and invasion, and inducing apoptosis activation. POLRMT KO, using CRISPR/Cas9 gene editing method, similarly induced profound anti-OS cell activity. Conversely, forced overexpression of POLRMT in primary OS cells accelerated cell proliferation, migration and invasion. In vivo, intratumoral injection of POLRMT shRNA AAV potently inhibited U2OS xenograft growth in nude mice. Importantly, the toxicity of the silencing for POLRMT was minor in the primary human osteoblasts as compared to that observed in OS cells. These results, together with results from other cancer studies [15, 16, 18, 31], strongly supported the pro-tumorigenic activity of POLRMT overexpression in human cancer. Targeting POLRMT could be a novel strategy to inhibit OS progression.
The mitochondrial functions are essential in tumorigenesis of OS [32, 33]. Bonuccelli et al. have found that OXPHOS and ATP-production were both significantly increased in OS cells after co-culture with mesenchymal stem cells, resulting in more aggressive behaviors of tumor cells, in particular enhanced cell migration [34]. Day et al., suggested that Bcl-xL can exert an anti-apoptotic effect by stimulating OXPHOS in OS cells [35]. Here we found that mtDNA contents, mitochondrial transcripts and expression of respiratory chain complex subunits were significantly reduced in POLRMT-silenced OS cells and U2OS xenografts, but were increased after POLRMT overexpression. Therefore, POLRMT-driven OS cell progression could be due to its effect in promoting OXPHOS. The underlying signaling mechanisms may warrant further characterizations.
Conclusion
Our results showed that POLRMT is overexpressed in human OS, promoting OS cell growth in vitro and in vivo. POLRMT could be a novel therapeutic target for better OS therapy.
Material and methods
Ethics
All the methods applied in this study were carried out according to the ethics guidelines by all authors’ institutions.
Chemicals, reagents, and antibodies
Puromycin, polybrene, fetal bovine serum (FBS), medium, antibiotics and other cell culture reagents were purchased from Sigma (St. Louis, Mo). Cell counting kit -8 (CCK-8) was provided by Dojindo (Kumamoto, Japan). The fluorescence dyes, including EdU (5-Ethynyl-2′-deoxyuridine), DAPI (4′,6-diamidino-2-phenylindole), TUNEL (Terminal deoxynucleotidyl transferase dUTP nick end labeling) and JC-1, were provided by Thermo-Fisher Invitrogen (Shanghai, China). The anti-POLRMT antibody was purchased from Abcam (ab228576, Shanghai, China). The cleaved caspase antibody sampler Kit (#9929) and anti-β-Tubulin (#2146) were provided by Cell Signaling Technologies (Beverly, MA).
Human tissues
OS tumor tissues and matched surrounding normal bone tissues were from a total of eleven (11) primary OS patients (provided by Dr. Cao’s group at Soochow University [19, 36]) with written-informed consents. All patients were administrated at authors institutions, received no prior therapies before surgeries. Fresh tissues were stored in liquid nitrogen. The protocols of using human tissues were approved by the Ethics Committee of The First Affiliated Hospital of Zhengzhou University, in according to the principles of Declaration of Helsinki.
OS cell culture
U2OS and MG-63 established human OS cell lines were from Dr. Zhang at Medical College of Southeast University [37,38,39]. The primary human OS cells, derived from two written-informed consent OS patients, were from Dr. Cao at Soochow University [19, 36]. The primary human osteoblasts were from Dr. Ji at Nanjing Medical University [40, 41]. The detailed protocols of culturing established and primary human OS cells as well as human osteoblasts were described early [37,38,39,40,41]. Mycoplasma-microbial contamination examination, STR profiling, population doubling time and morphology were checked to confirm the genotypes. The written-informed consent was obtained from each participant. The protocols of using primary human cells were approved by the Ethics Committee of The First Affiliated Hospital of Zhengzhou University, in according to the principles of Declaration of Helsinki.
Cell viability
In brief, OS cells with applied genetic modifications were seeded into 96-well plates (at 3 × 103 cells per well) and cultured for 96 h. Cell viability was examined by CCK-8 kits. In each well CCK-8 optical density (OD) at 490 nm was recorded.
Clonogenicity assay
OS cells with applied genetic modifications were re-suspended in complete medium with 0.5% agar (Sigma). Cell suspensions (3 × 104 cells in each plate) were thereafter added on top of the 100 mm-diameter culture dishes. The medium was renewed every 2 days. After 12 days, viable large colonies were stained and manually counted.
Western blotting
Cells or tissues were incubated with cell lysis buffer as described [42, 43]. The achieved protein lysates (30 μg of each treatment) were separated by 10–12% SDS-PAGE and thereafter transferred to polyvinylidene fluoride blots (Millipore, Invitrogen, Shanghai, China). Afterwards, blots were blocked and incubated with applied primary and secondary antibodies. Signal detection and data quantification were described in detail in elsewhere [42, 43].
Quantitative reverse transcription PCR (qRT-PCR)
Total RNA was extracted via TRIzol reagents (Invitrogen; Thermo Fisher Scientific). Complementary DNA (cDNA) was then synthesized through a PrimeScript RT Reagent kit (Takara, Shanghai, China). qRT-PCR assays were carried out by an ABI 7900 Fast Real-Time PCR system (Applied Bioscience; Thermo Fisher Scientific.) through a Power SYBR Green PCR Master mix kit (Applied Biosystems, Shanghai, China). A 2−ΔΔCt method was utilized for data quantification. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was always examined as internal control. mRNA primers were provided by Dr. Shi’s group at Soochow University [31].
TUNEL staining
U2OS cells with applied genetic treatments were seeded into 12-well plates (5 × 104 cells per well) and cultured for 96 h. A TUNEL In Situ Cell Apoptosis Detection Kit (Invitrogen) was employed. Briefly, TUNEL and DAPI fluorescence dyes were added to OS cells, visualized under a fluorescent microscope (Leica). TUNEL ratio (% versus DAPI) was calculated from at least 1500 cell nuclei in five random views (1 × 200) in each condition [38].
Caspase-3 activity assay
U2OS cells with applied genetic treatments were seeded into six-well tissue plates at 1.2 × 105 cells per well. Twenty μg of cytosolic protein extracts were added to the attached caspase assay buffer [37, 38] along with the caspase-3 substrate Ac-DEVD-AFC. Using a spectrofluorometer the released AFC intensity was test at 450 nm.
EdU staining
OS cells with applied genetic treatments were seeded into 12-well plates (at 5 × 104 cells per well) for 96 h. To test cell proliferation, an EdU Apollo-567 Kit (Invitrogen) was employed. Briefly, EdU and DAPI fluorescence dyes were added to OS cells, visualized under a fluorescent microscope (Leica). For each condition, at least 1500 cell nuclei from five random views (1 × 200) were included to calculate EdU-positive nuclei ratio (% versus DAPI).
Transwell assays
“Transwell” chambers with12 μm pore (Corning, Shanghai, China) were utilized for in vitro cell migration and invasion assays. OS cells with applied genetic modifications were placed on the upper chambers (at 2 × 104 cells per chamber, in 200 µL DMEM). The lower chambers were filled with complete medium with 10% FBS. Following 24 h, the non-migrated OS cells on the upper surface were removed carefully. On the lower surface migrated cells were fixed and stained (with crystal violet). For invasion assays, the chambers were coated with Matrigel (BD Biosciences, Shanghai, China).
Mitochondrial depolarization
OS cells with applied genetic treatments were seeded into 12-well plates (at 3 × 104 cells per well) for 72 h. JC-1 dye was then added for 2 h. JC-1 green monomer intensity, reflecting mitochondrial depolarization, was detected at the test-wavelength of 490 nm. JC-1 images, integrating both green (490 nm) and red (625 nm) fluorescence channels, were presented.
POLRMT silencing or overexpression
GV369 lentiviral constructs encoding shRNAs against non-overlapping sequences of POLRMT (sh-POLRMT-1# and sh-POLRMT-2#), as well as the GV369 lentiviral construct encoding full-length POLRMT cDNA were provided by Dr. Shi at Soochow University [31]. Each was transduced to HEK-293T cells together with the lentivirus helper plasmids. Within 48 h lentivirus was generated, enriched and filtered. OS cells were seeded into six-well plates (at 1 × 105 cells per well), cultured in polybrene-containing complete medium and infected with the lentivirus (MOI = 10). After 24 h, cells were cultured in fresh complete medium. Puromycin was added to select stable cells. POLRMT expression in stable OS cells was verified by qRT-PCR and Western blotting assays.
POLRMT knockout (KO)
OS cells were initially seeded into six-well plates (at 1 × 105 cells per well), cultured in polybrene-containing complete medium and transfected with a LentiCas9-puro construct (Genechem). Stable Cas9 cells were established after puromycin selection. Cells were further transfected with Lenti-POLRMT-sgRNA-puro construct (Genechem) for 24 h. Afterwards, cells were cultured in fresh complete medium. Puromycin was added to select stable cells. Stable cells were distributed into 96-well plates and subjected to POLRMT KO screening. POLRMT expression in the single stable OS cells was verified by qRT-PCR and Western blotting assays.
mtDNA contents
Total DNA in OS cells was extracted, purified and quantified. For each treatment 7.5 ng/μl DNA was analyzed by quantitative PCR via a Taqman 2× Universal PCR mastermix (Applied Biosystems, Shanghai, China) and commercially available Taqman assay probes for COX1 mtDNA (18 S). mtDNA contents in genetic modified OS cells were normalized to those in control OS cells.
Xenograft assay
The nude mice were purchased from the experimental animal center of Soochow University (Suzhou, China): Half male and hale female, 7-week old, 18.3–19.2 grams. Mice were maintained under the 12 h light/dark cycle in standard environmental conditions (22.2 ± 1.5 °C, 46–55% r humidity). U2OS cells, at 5 × 106 cells per mouse, were dissolved in 100 μL of RPMI and Matrigel (1:1) and were subcutaneously (s.c.) injected to right flanks of nude mice. Within 25 days, U2OS xenograft-bearing nude mice were established and the volume of each tumor close to 100 mm3. Afterward, mice were randomly assigned into two groups, with ten mice per group. One group mice were intratumorally injected with POLRMT shRNA adeno-associated virus (“AAV-sh-POLRMT”) and the other group mice received AAV packed scramble control shRNA (“AAV-c-sh”). AAV was injected daily for 6 consecutive days. Tumor dimensions were measured and tumor volume calculated using the following formula: Volume (V) = length × width × height × 0.5236. The animal protocols were according to the regulations of the IACUC and with the approval of Zhengzhou University.
Statistical studies
Data with normal distribution were presented as mean ± standard deviation (SD). Statistical differences between multiple groups were analyzed by one-way ANOVA with post hoc Bonferroni test (SPSS version 20.0, SPSS Co., Chicago, CA). The two-tailed unpaired T-test was applied (Excel 2007, Microsoft) for the comparison of two groups. P < 0.05 was considered statistically significant.
Data availability
The original contributions presented in the study are included in the paper. Further inquiries can be directed to the corresponding authors.
References
Anderson ME. Update on Survival in Osteosarcoma. Orthop Clin North Am. 2016;47:283–92.
Ottaviani G, Jaffe N. The epidemiology of osteosarcoma. Cancer Treat Res. 2009;152:3–13.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34.
Isakoff MS, Bielack SS, Meltzer P, Gorlick R. Osteosarcoma: current treatment and a collaborative pathway to success. J Clin Oncol. 2015;33:3029–35.
Kansara M, Teng MW, Smyth MJ, Thomas DM. Translational biology of osteosarcoma. Nat Rev Cancer. 2014;14:722–35.
Gill, J & Gorlick, R. Advancing therapy for osteosarcoma. Nat Rev Clin Oncol. 2021;18:609–24.
Sayles LC, Breese MR, Koehne AL, Leung SG, Lee AG, Liu HY, et al. Genome-Informed Targeted Therapy for Osteosarcoma. Cancer Disco. 2019;9:46–63.
Shaikh AB, Li F, Li M, He B, He X, Chen G, et al. Present Advances and Future Perspectives of Molecular Targeted Therapy for Osteosarcoma. Int J Mol Sci. 2016;17:506.
Yang J, Zhang W. New molecular insights into osteosarcoma targeted therapy. Curr Opin Oncol. 2013;25:398–406.
Shi Y, Dierckx A, Wanrooij PH, Wanrooij S, Larsson NG, Wilhelmsson LM, et al. Mammalian transcription factor A is a core component of the mitochondrial transcription machinery. Proc Natl Acad Sci USA. 2012;109:16510–5.
Larsson NG, Wang J, Wilhelmsson H, Oldfors A, Rustin P, Lewandoski M, et al. Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat Genet. 1998;18:231–6.
Posse V, Al-Behadili A, Uhler JP, Clausen AR, Reyes A, Zeviani M, et al. RNase H1 directs origin-specific initiation of DNA replication in human mitochondria. PLoS Genet. 2019;15:e1007781.
Fuste JM, Wanrooij S, Jemt E, Granycome CE, Cluett TJ, Shi Y, et al. Mitochondrial RNA polymerase is needed for activation of the origin of light-strand DNA replication. Mol Cell. 2010;37:67–78.
Bralha FN, Liyanage SU, Hurren R, Wang X, Son MH, Fung TA, et al. Targeting mitochondrial RNA polymerase in acute myeloid leukemia. Oncotarget 2015;6:37216–28.
Salem AF, Whitaker-Menezes D, Howell A, Sotgia F, Lisanti MP. Mitochondrial biogenesis in epithelial cancer cells promotes breast cancer tumor growth and confers autophagy resistance. Cell Cycle. 2012;11:4174–80.
Sotgia F, Whitaker-Menezes D, Martinez-Outschoorn UE, Salem AF, Tsirigos A, Lamb R, et al. Mitochondria “fuel” breast cancer metabolism: fifteen markers of mitochondrial biogenesis label epithelial cancer cells, but are excluded from adjacent stromal cells. Cell Cycle. 2012;11:4390–401.
Bonekamp NA, Peter B, Hillen HS, Felser A, Bergbrede T, Choidas A, et al. Small-molecule inhibitors of human mitochondrial DNA transcription. Nature 2020;588:712–6.
Shan HJ, Zhu LQ, Yao C, Zhang ZQ, Liu YY, Jiang Q, et al. MAFG-driven osteosarcoma cell progression is inhibited by a novel miRNA miR-4660. Mol Ther Nucleic Acids. 2021;24:385–402.
Salvioli S, Ardizzoni A, Franceschi C, Cossarizza A. JC-1, but not DiOC6(3) or rhodamine 123, is a reliable fluorescent probe to assess delta psi changes in intact cells: implications for studies on mitochondrial functionality during apoptosis. FEBS Lett. 1997;411:77–82.
Cossarizza A, Baccarani-Contri M, Kalashnikova G, Franceschi C. A new method for the cytofluorimetric analysis of mitochondrial membrane potential using the J-aggregate forming lipophilic cation 5,5′,6,6′-tetrachloro-1,1′,3,3′-tetraethylbenzimidazolcarbocyanine iodide (JC-1). Biochem Biophys Res Commun. 1993;197:40–5.
Weinberg SE, Chandel NS. Targeting mitochondria metabolism for cancer therapy. Nat Chem Biol. 2015;11:9–15.
Ashton TM, McKenna WG, Kunz-Schughart LA, Higgins GS. Oxidative Phosphorylation as an Emerging Target in Cancer Therapy. Clin Cancer Res. 2018;24:2482–90.
Cavalli LR, Varella-Garcia M, Liang BC. Diminished tumorigenic phenotype after depletion of mitochondrial DNA. Cell Growth Differ. 1997;8:1189–98.
Viale A, Corti D, Draetta GF. Tumors and mitochondrial respiration: a neglected connection. Cancer Res. 2015;75:3685–6.
Bosc C, Selak MA, Sarry JE. Resistance Is Futile: Targeting Mitochondrial Energetics and Metabolism to Overcome Drug Resistance in Cancer Treatment. Cell Metab. 2017;26:705–7.
Kuntz EM, Baquero P, Michie AM, Dunn K, Tardito S, Holyoake TL, et al. Targeting mitochondrial oxidative phosphorylation eradicates therapy-resistant chronic myeloid leukemia stem cells. Nat Med. 2017;23:1234–40.
Birsoy K, Possemato R, Lorbeer FK, Bayraktar EC, Thiru P, Yucel B, et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 2014;508:108–12.
Chaudhary S, Ganguly S, Palanichamy JK, Singh A, Bakhshi R, Jain A, et al. PGC1A driven enhanced mitochondrial DNA copy number predicts outcome in pediatric acute myeloid leukemia. Mitochondrion 2021;58:246–54.
Oran AR, Adams CM, Zhang XY, Gennaro VJ, Pfeiffer HK, Mellert HS, et al. Multi-focal control of mitochondrial gene expression by oncogenic MYC provides potential therapeutic targets in cancer. Oncotarget 2016;7:72395–414.
Zhou Tong, Sang Yong-Hua, Cai Shang, Xu Chun, Shi Min-hua. The requirement of mitochondrial RNA polymerase for non-small cell lung cancer cell growth. Cell Death Dis. 2021;12:751.
Sotgia F, Martinez-Outschoorn UE, Lisanti MP. The reverse Warburg effect in osteosarcoma. Oncotarget 2014;5:7982–3.
Giang AH, Raymond T, Brookes P, de Mesy Bentley K, Schwarz E, O’Keefe R, et al. Mitochondrial dysfunction and permeability transition in osteosarcoma cells showing the Warburg effect. J Biol Chem. 2013;288:33303–11.
Bonuccelli G, Avnet S, Grisendi G, Salerno M, Granchi D, Dominici M, et al. Role of mesenchymal stem cells in osteosarcoma and metabolic reprogramming of tumor cells. Oncotarget 2014;5:7575–88.
Dey R, Moraes CT. Lack of oxidative phosphorylation and low mitochondrial membrane potential decrease susceptibility to apoptosis and do not modulate the protective effect of Bcl-x(L) in osteosarcoma cells. J Biol Chem. 2000;275:7087–94.
Gao YY, Ling ZY, Zhu YR, Shi C, Wang Y, Zhang XY, et al. The histone acetyltransferase HBO1 functions as a novel oncogenic gene in osteosarcoma. Theranostics 2021;11:4599–615.
Zhu YR, Xu Y, Fang JF, Zhou F, Deng XW, Zhang YQ. Bufotalin-induced apoptosis in osteoblastoma cells is associated with endoplasmic reticulum stress activation. Biochem Biophys Res Commun. 2014;451:112–8.
Zhu YR, Min H, Fang JF, Zhou F, Deng XW, Zhang YQ. Activity of the novel dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor NVP-BEZ235 against osteosarcoma. Cancer Biol Ther. 2015;16:602–9.
Zhu YR, Zhou XZ, Zhu LQ, Yao C, Fang JF, Zhou F, et al. The anti-cancer activity of the mTORC1/2 dual inhibitor XL388 in preclinical osteosarcoma models. Oncotarget 2016;7:49527–38.
Zhuang Y, Wang S, Fei H, Ji F, Sun P. miR-107 inhibition upregulates CAB39 and activates AMPK-Nrf2 signaling to protect osteoblasts from dexamethasone-induced oxidative injury and cytotoxicity. Aging (Albany NY). 2020;12:11754–67.
Zhao S, Mao L, Wang SG, Chen FL, Ji F, Fei HD. MicroRNA-200a activates Nrf2 signaling to protect osteoblasts from dexamethasone. Oncotarget 2017;8:104867–76.
Cao C, Rioult-Pedotti MS, Migani P, Yu CJ, Tiwari R, Parang K, et al. Impairment of TrkB-PSD-95 signaling in Angelman syndrome. PLoS Biol. 2013;11:e1001478.
Cao C, Huang X, Han Y, Wan Y, Birnbaumer L, Feng GS, et al. Galpha(i1) and Galpha(i3) are required for epidermal growth factor-mediated activation of the Akt-mTORC1 pathway. Sci Signal. 2009;2:ra17.
Acknowledgements
This work is supported by the National Natural Science Foundation of China (82002850) and by the Key R&D and Promotion Program of the Science and Technology Department of Henan Province (LHGJ20190173). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the paper.
Author information
Authors and Affiliations
Contributions
All the listed authors designed the study, performed the experiments and the statistical analysis, and wrote the paper and revise it. Authors have read the paper and approved the final submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All the methods applied in this study were carried out according to the ethics guidelines of Zhengzhou University, in according to the principles of Declaration of Helsinki.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, Qc., Zhang, Xy., Yan, Ph. et al. Identification of mitochondrial RNA polymerase as a potential therapeutic target of osteosarcoma. Cell Death Discov. 7, 393 (2021). https://doi.org/10.1038/s41420-021-00780-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-021-00780-x
- Springer Nature Limited
This article is cited by
-
Identification of the central role of RNA polymerase mitochondrial for angiogenesis
Cell Communication and Signaling (2024)
-
Targeting the mitochondrial protein YME1L to inhibit osteosarcoma cell growth in vitro and in vivo
Cell Death & Disease (2024)
-
Targeting POLRMT by IMT1 inhibits colorectal cancer cell growth
Cell Death & Disease (2024)
-
Mitochondria transcription and cancer
Cell Death Discovery (2024)
-
Targeting POLRMT by a first-in-class inhibitor IMT1 inhibits osteosarcoma cell growth in vitro and in vivo
Cell Death & Disease (2024)