Abstract
Background
The “watch-and-wait” approach is a common treatment option amongst patients with locally advanced rectal cancer (LARC). However, the diagnostic sensitivity of clinical modalities, such as colonoscopy and magnetic resonance imaging to determine pathological response, is not high. We analysed the clinical utility of circulating tumour DNA (ctDNA) of patients with LARC to predict response to preoperative therapy and postoperative recurrence.
Methods
A serial ctDNA analysis of 222 plasma samples from 85 patients with LARC was performed using amplicon-based deep sequencing on a cell-free DNA panel covering 14 genes with over 240 hotspots.
Results
ctDNA was detected in 57.6% and 22.3% of samples at baseline and after preoperative treatment, respectively, which was significantly different (P = 0.0003). Change in ctDNA was an independent predictor of complete response to preoperative therapy (P = 0.0276). In addition, postoperative ctDNA and carcinoembryonic antigen (CEA) were independent prognostic markers for risk of recurrence after surgery (ctDNA, P = 0.0127 and CEA, P = 0.0105), with a combined analysis having cumulative effects on recurrence-free survival (P = 1.0 × 10–16).
Conclusions
Serial ctDNA analysis may offer clinically useful predictive and prognostic markers for response to preoperative therapy and postoperative recurrence in patients with LARC.
Similar content being viewed by others
Background
Rectal cancer is the second-most common cancer of the large intestine, and a major health burden globally. Radiation therapy can improve local recurrence rates in patients with locally advanced rectal cancer (LARC),1,2,3 with preoperative chemoradiation therapy (CRT) considered the standard of care for these patients. Indeed, studies have shown that CRT before surgery can reduce tumour volume and invoke pathological complete response (pCR, ypT0N0M0) in 15–20% of patients.4,5,6 The achievement of pCR is associated with improved local and distant control, disease-free survival and overall survival; some patients who achieve pCR may not even require surgery.7 Although several studies have reported factors that are predictive of pCR after preoperative CRT,8,9,10,11 the utility of such factors to predict pCR in the clinical setting has not been identified. In addition, despite the range of options available to detect pCR, including digital rectal examinations, endoscopic assessments of mucosal integrity and magnetic resonance imaging (MRI) for changes in primary lesions, the accuracy of these modalities is questionable.12,13,14
The advent of liquid biopsies was expected to allow for an interrogation of the cancer genome while providing a minimally invasive approach for patients with cancer.15 Liquid biopsies are based on the analysis of circulating tumour DNA (ctDNA), which is secreted from cancer cells into the peripheral blood as a result of cell apoptosis and/or necrosis.15,16 However, ctDNA represents only a small fraction of the cell-free DNA (cfDNA), meaning that there is very little ctDNA with each blood sample drawn. Despite this limitation, various studies have reported the clinical utility of cfDNA and ctDNA for the treatment of colorectal cancer, in terms of monitoring treatment response, cancer recurrence and drug resistance.17,18,19,20 The clinical utility of ctDNA to predict patient responses to preoperative therapy, including CRT among patients with LARC, has not yet been reported.
A diagnostic system that could predict patient responses to preoperative therapy would help to select those patients with LARC who would be expected to achieve pCR, and provide the option for these patients to be managed without immediate surgery (i.e., the “watch-and-wait” approach).7,21 Thus, the aim of the present study was to investigate the clinical utility of serial ctDNA analysis to predict responses to preoperative therapy and clinical outcomes after surgical intervention in patients with LARC. To this end, we examined mutant allele fractions of 14 genes that are frequently mutated in colorectal cancer using targeted next-generation sequencing (NGS). Our results may aid in the development of a diagnostic system for personalised preoperative therapy, and help to improve survival prognostication and the stratification of patients with LARC.
Methods
Patients
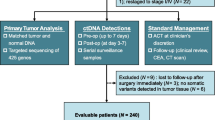
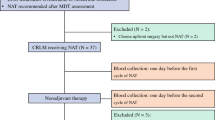
This study prospectively enrolled patients with LARC between February 2017 and November 2018. All patients were diagnosed as clinical stage II or III (cT3-4N0, or cTanyN+) using the Union for International Cancer Control (UICC) TNM classification, and received preoperative therapy at the Cancer Institute Hospital, Japanese Foundation for Cancer Research. Thirty-three patients received standard CRT, comprising a total dose of 50.4 Gy in 28 fractions over 5 weeks and concurrent Tegafur/Gimeracil/Oteracil (80–120 mg/m2/day) orally administered over 4 weeks. Nine patients received short-course radiotherapy (SRT), with 25 Gy administered in 5 fractions over 1 week. Four patients with upper rectal cancer diagnosed as cT4 or cN2 received neoadjuvant chemotherapy (NAC), consisting of four cycles of capecitabine (2000 mg/m2/day), oxaliplatin (130 mg/m2) and bevacizumab (7.5 mg/kg) (CapeOx + Bmab). Twenty-three patients with lower rectal cancer diagnosed as cT4 or cN2 received a combination of 6 cycles of fluorouracil (bolus, 400 mg/m2; infusion, 2400 mg/m2), leucovorin (200 mg/m2), oxaliplatin (85 mg/m2) and bevacizumab (5 mg/kg) (FOLFOX + Bmab) as NAC, followed by CRT. In addition, 16 patients with cT3N1M0 and an intermediate risk of recurrence received a combination of SRT, followed by four cycles of CapeOx. All patients were histologically diagnosed with adenocarcinoma, and received pre-treatment rectal MRI and chest/abdomen/pelvis CT. All patients had an Eastern Cooperative Oncology Group performance status of 0–2, and received preoperative therapy followed by surgery or non-operative management. The decision for non-operative management was carefully discussed with each patient who achieved clinical complete response (cCR), according to the National Comprehensive Cancer Network (NCCN) guidelines. Patients with previous cancers were excluded. At 4–8 weeks after completing all neoadjuvant, preoperative therapies, all patients, except those receiving SRT, were re-staged using colonoscopy, rectal MRI and chest/abdomen/pelvis CT scan. The median time to surgery in patients receiving SRT was 10 days (IQR, 7–30 days): six patients receiving SRT underwent surgery within 15 days after SRT, but surgery was delayed for about 4 weeks after SRT in three patients. Endoscopic response was classified by mucosal findings, as reported by Chino et al.22 Clinicopathological factors were evaluated by UICC TNM classification. cCR was defined as ycT0N0M0, and pathological complete response (pCR) as ypT0N0M0. Dworak’s criteria were used for tumour regression grade (TRG): TRG 1, minimal regression; TRG 2, moderate regression; TRG 3, near-complete regression; TRG 4, complete regression.23 This study was approved by the Institutional Review Boards of the Japanese Foundation for Cancer Research (Tokyo, Japan). Written informed consent was obtained from all patients.
Blood sampling, cell-free DNA isolation and sequencing
For ctDNA analysis, blood samples were collected before the initiation of preoperative therapy (baseline), after preoperative treatment (post treatment, just before surgery) and at 12 weeks after surgery (post operation). A total of 222 blood samples from 85 patients were collected into EDTA tubes following the manufacturer’s instructions.15 Plasma was extracted by centrifugation at 1600 × g for 10 min at 4 °C, followed by 16,000×g for 10 min at 4 °C to remove cellular debris. cfDNA was extracted from plasma using a MagMAX cfDNA Isolation Kit (Thermo Fisher Scientific), following the manufacturer’s instructions.15 cfDNA quality was checked using Qubit2.0 and 2100 Bioanalyzer (Agilent Technologies). Libraries from the cfDNA were prepared with Oncomine Colon cfDNA Assay (Thermo Fisher Scientific), and checked using Qubit2.0 and 2100 Bioanalyzer.15 Molecular barcoded amplicon-based deep sequencing was used in this assay to reduce the false positives derived from polymerase chain reaction (PCR) errors. The Ion Chef System and Ion 530 Kit-Chef were used for template preparation. Ion 530 chips were sequenced on an Ion S5 system. The six-plex library pool was applied to one Ion 530 chip. The cfDNA panel used in this study covered 14 genes (TP53, KRAS, APC, PIK3CA, FBXW7, NRAS, GNAS, SMAD4, MAP2K1, ERBB2, BRAF, AKT1, CTNNB1 and EGFR) with more than 240 hotspots (SNVs and short indels). The clean reads were mapped to the human reference genome (hg19). Variant caller was used to filter and call mutations in the targeted regions of each gene.24,25 The cut-off value for the mutant allele fraction (MAF) was 0.15%. The average coverage ranged from 20,000 to 50,000. The total MAF of the detected mutant alleles in each patient was used as the metric for ctDNA in this study.
We also measured carcinoembryonic antigen (CEA) in the hospital central laboratory before preoperative therapy, after preoperative treatment and at 12 weeks after surgery.
Statistical analysis
Continuous variables are expressed as the median and interquartile range (IQR). Fisher’s exact test was used to compare the proportion of patients with plasma mutations before and after preoperative therapy. Wilcoxon matched-pair signed-rank test was used to evaluate the significance of changes in ctDNA after preoperative therapy. Logistic regression analysis was used to identify significant predictive factors, and to test for the independent contribution of factors in response to preoperative therapy. To assess for potential confounding factors, gender, pre-treatment CEA, distance from the anal verge, clinical T and N factors, NAC with CRT/SRT and change in ctDNA were treated as categorical variables. Furthermore, Cox proportional hazard analysis was used to identify significant prognostic factors, and to test for an independent contribution of these factors to recurrence-free survival. In this case, potential confounding was assessed using gender, postoperative CEA, pathological T and N factors, lymphovascular invasion, response to preoperative therapy (Dworak’s TRG) and NAC with CRT/SRT. Postoperative ctDNA was treated as a categorical variable. Variables with a P value less than 0.2 in the univariate analysis were included in the multivariate analysis. Recurrence-free survival curves were visualised by the Kaplan–Meier method. The differences between the curves were estimated using the log-rank test. Logistic regression and Cox proportional hazard analyses were performed using JMP statistical software. Other analyses were performed using GraphPad Prism7. All P values were two-sided. Values with P < 0.05 were considered to be statistically significant. We used a significance level of 0.0025 (0.05/20 clinical factors) in the association study of clinical factors with ctDNA (Table 1) to adjust for multiple testing by Bonferroni correction.
Results
Clinicopathological characteristics
We prospectively recruited 85 patients with LARC who were receiving preoperative chemotherapy and/or RT. Table 1 shows the characteristics of these patients. At recruitment, the median age was 60 years (IQR, 52–69 years), 65 (76.5%) patients were male and 58 (68.2%) had been diagnosed as clinical stage III (cTanyN1–2). Thirty-nine (45.9%) patients received NAC with CRT/SRT (Table 1). After preoperative therapy, 12 (14.1%) patients achieved cCR (ycT0N0M0) (Table 1). Eight of these 12 patients chose non-operative management (“watch-and-wait” approach) after therapy (Table 1; Supplementary Fig. 1). The remaining 77 (90.6%) patients of the original cohort were treated surgically (Table 1; Supplementary Fig. 1), of which 17 (22.1%) patients were diagnosed as Dworak’s TRG 4 (Fig. 1). Of these 17 patients, four patients had an absence of cancer cells at the primary lesion, but were positive for pathological lymph node metastasis (ypT0N+). The other 13 patients achieved pCR (ypT0N0M0) after preoperative therapy (Fig. 1). The median interval from completing preoperative therapy to surgery was 54 days (IQR, 33–68.5, Table 1).
Gene mutations in 14 genes from samples retrieved from 85 patients with LARC. Grey, no mutation detected; white, analysis not conducted. pCR pathological complete response, cCR clinical complete response, TRG tumour regression grade (Dworak), W&W watch-and-wait, Post-Tx post–preoperative treatment, Post-Ope post operation, MAF the highest mutant allele fraction in each patient.
Detection of somatic mutations in plasma
A total of 222 plasma samples from 85 patients were analysed by amplicon-based deep sequencing. One or more somatic mutations (mutant alleles) were detected in 49 (57.6%) patients at baseline (before preoperative therapy), but only in 19 (22.3%) patients after preoperative treatment, which was a significant reduction (P < 0.0001, Supplementary Fig. 2). No significant associations were observed between baseline or postoperative ctDNA status and any clinicopathological factors (Table 1). In contrast, post-treatment ctDNA detection was associated with post-treatment and postoperative CEA level, preoperative therapy regimens of NAC with CRT/SRT and venous invasion (Table 1). Only post-treatment CEA was significantly associated with ctDNA after Bonferroni correction (P < 0.0001). Mutations in TP53, KRAS and APC genes were detected in 24 (29.3%), 22 (26.8%) and 14 (17.1%) patients at baseline, respectively (Fig. 1 and Supplementary Fig. 3). Mutations in other genes were less common (<10% of patients) (Fig. 1 and Supplementary Figs. 3 and 4). Thirty-two (68.1%) mutations detected at baseline were not detected after preoperative therapy (Fig. 2). TP53 (13 mutations) and APC (11 mutations) were the most frequently mutated genes after preoperative therapy (Fig. 1; Supplementary Fig. 4).
Association between response to preoperative therapy and ctDNA response
Patients were classified into two groups in this study: responders (n = 21) and non-responders (n = 64). Responders were patients who achieved pCR after preoperative therapy or who were managed by the watch-and-wait approach for more than 12 months after achieving cCR (ycT0N0M0). Non-responders were patients who did not achieve pCR after preoperative therapy. No significant association was observed between baseline, post-treatment and postoperative ctDNA status and the rate of responders (Table 1).
To examine the change in MAFs in response to preoperative treatment, we used data from 51 patients (12 responders and 39 non-responders) for whom mutation(s) were detected before and/or after preoperative treatment (Fig. 2). MAF after preoperative treatment (median, 0%; IQR, 0–0.27%) was significantly lower than that at baseline (median, 0.49%; IQR, 0.23–1.22%; P = 0.0003), as shown in Fig. 2. Of the 12 responders, all but one patient showed a decrease in MAFs after preoperative treatment (Fig. 2b). Post-treatment ctDNA was detected in this one responder (8.3%), who had no detectable ctDNA at baseline. In comparison, 7 of the 39 non-responders showed an increase in MAF (Fig. 2c). Post-treatment ctDNA was detected in the three non-responders (7.7%) for whom there was also no detectable ctDNA at baseline. We performed a univariate logistic regression analysis to identify which factors were associated with response to preoperative therapy. We found a significant association between response and change in ctDNA (≥80% vs < 80%, P = 0.015; OR, 8.5; 95% CI, 1.4–163, Table 2). Preoperative therapy regimens of NAC with CRT/SRT showed a trend towards an increased response to preoperative therapy (P = 0.1178; OR, 2.9; 95% CI, 0.8–12.3, Table 2). In the multivariate analysis, the change in ctDNA still remained an independent predictor for response to preoperative therapy after adjustment (P = 0.0276; adjusted OR, 7.4; 95% CI, 1.2–144 for the change in ctDNA, Table 2).
Association between postoperative ctDNA and clinical outcome
We next investigated the clinical significance of postoperative ctDNA as a prognostic marker of clinical outcome after radical operation in 59 patients with LARC (Supplementary Fig. 1). In the univariate Cox proportional hazard analysis, postoperative CEA, ypT factor, lymphovascular invasion and postoperative ctDNA were considered to be significantly associated with recurrence-free survival after surgery (Table 3). In the multivariate analysis, postoperative CEA and ctDNA levels still remained independent prognostic factors of postoperative recurrence after adjusting for six parameters used in the univariate analysis (P = 0.0105; adjusted HR, 6.9; 95% CI, 1.6–29 for postoperative CEA and P = 0.0127; adjusted HR, 7.7; 95% CI, 1.6–42 for postoperative ctDNA, Table 3). Kaplan–Meier estimates indicated significantly different recurrence-free survival for patients with higher postoperative CEA (≥5 ng/ml) and higher postoperative ctDNA (≥0.5%) (Log-rank P = 7.5 × 10–7 for CEA, Log-rank P = 1.7 × 10–17 for ctDNA, Fig. 3). Furthermore, a combined analysis of postoperative CEA and ctDNA revealed cumulative effects on recurrence-free survival (Log-rank P = 1.0 × 10–16). The adjusted HR for risk of recurrence computed for patients carrying risk factors increased from 4.2-fold (either higher CEA or ctDNA) to 33.9-fold (both of them) compared with those without any risk factors (Fig. 3).
Discussion
This study represents the first association between response to preoperative therapy and the results of serial ctDNA analysis for patients with LARC. In this study, we show that the change in ctDNA measured in consecutive samples could act as an indicator of response to preoperative therapy (pCR) in patients with LARC. Through clinical and genetic analyses, we show that postoperative ctDNA and CEA were significantly associated with recurrence-free survival in 59 Japanese patients with LARC after preoperative therapy and radical operation. Furthermore, a combined analysis of postoperative ctDNA with postoperative CEA levels revealed that the number of risk factors (0, 1 or 2 factors) has a cumulative effect on the rate of recurrence-free survival in patients with LARC. These findings may help physicians to select the best treatment strategy for patients with LARC after preoperative therapy (i.e., surgery vs the watch-and-wait approach) and choose an optimal adjuvant therapy based on the results of the postoperative ctDNA and CEA levels.
Numerous studies have sought to identify clinically useful predictors of pCR after preoperative radiation therapy and/or chemotherapy for patients with LARC.4,8,9,10,11,20,26,27 Several parameters have been reported as possible predictors of the response to preoperative CRT in patients with LARC, including CEA levels before CRT, the distance of the tumour from the anal verge, tumour size, clinical lymph node metastasis and the interval between CRT and surgery.4,8,9,11 Habr-Gama et al. suggested that a strict definition of the clinical and endoscopic findings of patients with cCR after preoperative CRT could be used to select for patients who could be managed with a watch-and-wait strategy.28 Others, however, have reported that endoscopic evaluation for the response to preoperative CRT has low sensitivity to detect pCR.12,14,22,28 Although studies have reported an association between clinical outcomes and cfDNA or ctDNA among patients receiving CRT for LARC,17,29,30 no study has reported the positive correlation between pCR and cfDNA or ctDNA after CRT. Intensified regimens, such as NAC with CRT, might have had an impact on the response to preoperative therapy. Indeed, in the univariate analysis, NAC with CRT/SRT showed a trend towards an increased response to preoperative therapy. However, the multivariate logistic regression analysis showed only change in ctDNA as an independent predictor for treatment response (Table 2). The present study is the first to identify a change in ctDNA as a promising predictor of response to preoperative chemotherapy and/or radiation therapy in patients with LARC. Although the positive and negative predictive values were 33.3% (11 responders among 33 patients with the change in ctDNA ≥ 80%) and 94.4% (17 non-responders among 18 patients with the change in ctDNA < 80%), respectively, the positive predictive value increased to 54.5% (6 responders among 11 patients with endoscopic CR and a change in ctDNA ≥ 80%) when the change in ctDNA data was combined with endoscopic findings (Supplementary Fig. 5). Hence, a combinatorial analysis of the change in ctDNA with clinical factors, including endoscopic findings, might help to identify and select for patients who do not need immediate surgical management; further validation studies are needed to verify the clinical utility of this proposal.
In our study, the number of patients with mutations decreased after preoperative therapy (Supplementary Figs. 3 and 4), and this reduction was noted for all genes, presumably because tumour shrinkage after preoperative therapy reduced the amount of ctDNA available for analysis. The number of patients with TP53 mutations—the most frequently detected in this study—was significantly lower after preoperative therapy amongst responders (P = 0.026) but not amongst non-responders (Supplementary Table 1). TP53 is one of the best-studied tumour-suppressor genes31—referred to as the guardian of the genome—and represents a key regulator of cellular growth control.32 p53 plays a critical role in regulating DNA repair and apoptosis in response to radiation, and TP53 mutations are reported to decrease radiation-induced apoptosis in several types of cancers.33,34 These lines of evidence suggest that chemoradiation therapy provides a selective pressure for the expansion of TP53-mutant cells in residual tumours; further analysis using a larger number of patients and comparing the mutational status of tissues and plasma is required to verify the above hypothesis.35
There were several limitations in this study. First, because the number of recruited patients we assessed was small and the follow-up period was short, there were too few events to correct for potential confounding factors in the multivariate analyses. Second, the preoperative therapy regimen was not completely standardised, and patients underwent different approaches, based on different risk profiles. Third, tumour biopsy sequencing was not performed in our study. However, mutations in cfDNA, which are not detected in tumour biopsies, may comprise a subset of alterations that reflect ongoing tumour evolution and heterogeneity not captured in the small and usually anatomically constrained biopsy. Moreover, the proportion of patients with positive ctDNA at baseline was low, and the frequencies of mutated genes in plasma ctDNA samples from patients with LARC in our study were inconsistent with those measured from DNA tissue samples recorded in The Cancer Genome Atlas database.36,37 For example, the frequency of APC mutations in colorectal cancer tissue samples is reported to be ~80%,36,38 whereas we found a mutation frequency of only 17.1%. The differences in the frequencies could be partially due to an insufficient coverage of mutation detection for the APC gene in our ctDNA study. Further technical improvement in the gene panel would increase the detection accuracy among patients with LARC.
In conclusion, we show that serial ctDNA analysis is applicable for the prediction of treatment response among patients with LARC who undergo preoperative therapy. Our study also shows that postoperative ctDNA may offer a strong indicator of clinical outcome after radical operation for patients with LARC. Moreover, we demonstrate a cumulative effect of combining postoperative ctDNA with postoperative CEA levels as prognostic markers of recurrence-free survival among patients with LARC who are treated surgically. Our findings provide new insight into precision medicine for patients with LARC. To improve the quality of life of patients with LARC, future studies should integrate ctDNA testing of hundreds of cancer-related genes with large amounts of clinical data to improve patient selection and management.
References
Cedermark, B., Dahlberg, M., Glimelius, B., Pahlman, L., Rutqvist, L. E. & Wilking, N. Improved survival with preoperative radiotherapy in resectable rectal cancer. N. Engl. J. Med. 336, 980–987 (1997).
Kapiteijn, E., Marijnen, C. A., Nagtegaal, I. D., Putter, H., Steup, W. H., Wiggers, T. et al. Preoperative radiotherapy combined with total mesorectal excision for resectable rectal cancer. N. Engl. J. Med. 345, 638–646 (2001).
Peeters, K. C., Marijnen, C. A., Nagtegaal, I. D., Kranenbarg, E. K., Putter, H., Wiggers, T. et al. The TME trial after a median follow-up of 6 years: increased local control but no survival benefit in irradiated patients with resectable rectal carcinoma. Ann. Surg. 246, 693–701 (2007).
Das, P., Skibber, J. M., Rodriguez-Bigas, M. A., Feig, B. W., Chang, G. J., Wolff, R. A. et al. Predictors of tumor response and downstaging in patients who receive preoperative chemoradiation for rectal cancer. Cancer 109, 1750–1755 (2007).
Silberfein, E. J., Kattepogu, K. M., Hu, C. Y., Skibber, J. M., Rodriguez-Bigas, M. A., Feig, B. et al. Long-term survival and recurrence outcomes following surgery for distal rectal cancer. Ann. Surg. Oncol. 17, 2863–2869 (2010).
Smith, K. D., Tan, D., Das, P., Chang, G. J., Kattepogu, K., Feig, B. W. et al. Clinical significance of acellular mucin in rectal adenocarcinoma patients with a pathologic complete response to preoperative chemoradiation. Ann. Surg. 251, 261–264 (2010).
Habr-Gama, A., Perez, R. O., Nadalin, W., Sabbaga, J., Ribeiro, U., Silva e Sousa, A. H. et al. Operative versus nonoperative treatment for stage 0 distal rectal cancer following chemoradiation therapy. Ann Surg. 240, 711–718 (2004).
Restivo, A., Zorcolo, L., Cocco, I. M., Manunza, R., Margiani, C., Marongiu, L. et al. Elevated CEA levels and low distance of the tumor from the anal verge are predictors of incomplete response to chemoradiation in patients with rectal cancer. Ann. Surg. Oncol. 20, 864–871 (2013).
Garland, M. L., Vather, R., Bunkley, N., Pearse, M. & Bissett, I. P. Clinical tumour size and nodal status predict pathologic complete response following neoadjuvant chemoradiotherapy for rectal cancer. Int. J. Colorectal Dis. 29, 301–307 (2014).
Russo, A. L., Ryan, D. P., Borger, D. R., Wo, J. Y., Szymonifka, J., Liang, W. Y. et al. Mutational and clinical predictors of pathologic complete response in the treatment of locally advanced rectal cancer. J. Gastrointest. Cancer 45, 34–39 (2014).
Bitterman, D. S., Resende Salgado, L., Moore, H. G., Sanfilippo, N. J., Gu, P., Hatzaras, I. et al. Predictors of complete response and disease recurrence following chemoradiation for rectal cancer. Front. Oncol. 5, 286 (2015).
Smith, F. M., Wiland, H., Mace, A., Pai, R. K. & Kalady, M. F. Clinical criteria underestimate complete pathological response in rectal cancer treated with neoadjuvant chemoradiotherapy. Dis. Colon Rectum 57, 311–315 (2014).
van den Broek, J. J., van der Wolf, F. S., Lahaye, M. J., Heijnen, L. A., Meischl, C., Heitbrink, M. A. et al. Accuracy of MRI in restaging locally advanced rectal cancer after preoperative chemoradiation. Dis. Colon Rectum 60, 274–283 (2017).
Liu, S., Zhong, G. X., Zhou, W. X., Xue, H. D., Pan, W. D., Xu, L. et al. Can endorectal ultrasound, MRI, and mucosa integrity accurately predict the complete response for mid-low rectal cancer after preoperative chemoradiation? A prospective observational study from a single medical center. Dis. Colon Rectum 61, 903–910 (2018).
Osumi, H., Shinozaki, E., Takeda, Y., Wakatsuki, T., Ichimura, T., Saiura, A. et al. Clinical relevance of circulating tumor DNA assessed through deep sequencing in patients with metastatic colorectal cancer. Cancer Med. 8, 408–417 (2019).
Husain, H. & Velculescu, V. E. Cancer DNA in the circulation: the liquid biopsy. J. Am. Med. Assoc. 318, 1272–1274 (2017).
Tie, J., Kinde, I., Wang, Y., Wong, H. L., Roebert, J., Christie, M. et al. Circulating tumor DNA as an early marker of therapeutic response in patients with metastatic colorectal cancer. Ann. Oncol. 26, 1715–1722 (2015).
Nakamura, Y. & Yoshino, T. Clinical utility of analyzing circulating tumor DNA in patients with metastatic colorectal cancer. Oncologist 23, 1310–1318 (2018).
Thomsen, C. B., Hansen, T. F., Andersen, R. F., Lindebjerg, J., Jensen, L. H. & Jakobsen, A. Monitoring the effect of first line treatment in RAS/RAF mutated metastatic colorectal cancer by serial analysis of tumor specific DNA in plasma. J. Exp. Clin. Cancer Res. 37, 55 (2018).
Schou, J. V., Larsen, F. O., Sorensen, B. S., Abrantes, R., Boysen, A. K., Johansen, J. S. et al. Circulating cell-free DNA as predictor of treatment failure after neoadjuvant chemo-radiotherapy before surgery in patients with locally advanced rectal cancer. Ann. Oncol. 29, 610–615 (2018).
Habr-Gama, A., Sabbaga, J., Gama-Rodrigues, J., Sao Juliao, G. P., Proscurshim, I., Bailao Aguilar, P. et al. Watch and wait approach following extended neoadjuvant chemoradiation for distal rectal cancer: are we getting closer to anal cancer management? Dis. Colon Rectum 56, 1109–1117 (2013).
Chino, A., Konishi, T., Ogura, A., Kawachi, H., Osumi, H., Yoshio, T. et al. Endoscopic criteria to evaluate tumor response of rectal cancer to neoadjuvant chemoradiotherapy using magnifying chromoendoscopy. Eur. J. Surg. Oncol. 44, 1247–1253 (2018).
Dworak, O., Keilholz, L. & Hoffmann, A. Pathological features of rectal cancer after preoperative radiochemotherapy. Int. J. Colorectal Dis. 12, 19–23 (1997).
Shin, S., Kim, Y., Chul Oh, S., Yu, N., Lee, S. T., Rak Choi, J. et al. Validation and optimization of the ion torrent S5 XL sequencer and oncomine workflow for BRCA1 and BRCA2 genetic testing. Oncotarget 8, 34858–34866 (2017).
Xu, C. A review of somatic single nucleotide variant calling algorithms for next-generation sequencing data. Comput Struct. Biotechnol. J. 16, 15–24 (2018).
Wasserberg, N. Interval to surgery after neoadjuvant treatment for colorectal cancer. World J. Gastroenterol. 20, 4256–4262 (2014).
Chow, O. S., Kuk, D., Keskin, M., Smith, J. J., Camacho, N., Pelossof, R. et al. KRAS and combined KRAS/TP53 mutations in locally advanced rectal cancer are independently associated with decreased response to neoadjuvant therapy. Ann. Surg. Oncol. 23, 2548–2555 (2016).
Habr-Gama, A., Perez, R. O., Wynn, G., Marks, J., Kessler, H. & Gama-Rodrigues, J. Complete clinical response after neoadjuvant chemoradiation therapy for distal rectal cancer: characterization of clinical and endoscopic findings for standardization. Dis. Colon Rectum 53, 1692–1698 (2010).
Tie, J., Wang, Y., Tomasetti, C., Li, L., Springer, S., Kinde, I. et al. Circulating tumor DNA analysis detects minimal residual disease and predicts recurrence in patients with stage II colon cancer. Sci. Transl. Med. 8, 346ra92 (2016).
Tie, J., Cohen, J. D., Wang, Y., Li, L., Christie, M., Simons, K. et al. Serial circulating tumour DNA analysis during multimodality treatment of locally advanced rectal cancer: a prospective biomarker study. Gut 68, 663–671 (2019).
Naccarati, A., Polakova, V., Pardini, B., Vodickova, L., Hemminki, K., Kumar, R. et al. Mutations and polymorphisms in TP53 gene–an overview on the role in colorectal cancer. Mutagenesis 27, 211–218 (2012).
Lane, D. P. Cancer. p53, guardian of the genome. Nature 358, 15–16 (1992).
Merritt, A. J., Potten, C. S., Kemp, C. J., Hickman, J. A., Balmain, A., Lane, D. P. et al. The role of p53 in spontaneous and radiation-induced apoptosis in the gastrointestinal tract of normal and p53-deficient mice. Cancer Res. 54, 614–617 (1994).
Spitz, F. R., Nguyen, D., Skibber, J. M., Meyn, R. E., Cristiano, R. J. & Roth, J. A. Adenoviral-mediated wild-type p53 gene expression sensitizes colorectal cancer cells to ionizing radiation. Clin. Cancer Res. 2, 1665–1671 (1996).
Sakai, K., Kazama, S., Nagai, Y., Murono, K., Tanaka, T., Ishihara, S. et al. Chemoradiation provides a physiological selective pressure that increases the expansion of aberrant TP53 tumor variants in residual rectal cancerous regions. Oncotarget 5, 9641–9649 (2014).
The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Yaeger, R., Chatila, W. K., Lipsyc, M. D., Hechtman, J. F., Cercek, A., Sanchez-Vega, F. et al. Clinical sequencing defines the genomic landscape of metastatic colorectal cancer. Cancer Cell 33, 125–136 (2018).
Sjoblom, T., Jones, S., Wood, L. D., Parsons, D. W., Lin, J., Barber, T. D. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Acknowledgements
We express our heartfelt gratitude to all study participants. We thank Ms. Aya Imai, Mr. Yuki Sano and Ms. Marie Muramatsu for technical assistance. We thank Rebecca Jackson for her editorial support. We thank all other members and staff for their contributions to sample collection and the completion of our study.
Author information
Authors and Affiliations
Contributions
S.M., T.A. and H.Z. designed the study and wrote the paper. T.A., T.S., Y.F. and M.U. contributed to sample collection. S.M., T.N. and H.Z. contributed to the biological analysis. All authors critically reviewed and approved the final version of the paper.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Institutional Review Boards of the Japanese Foundation for Cancer Research (Tokyo, Japan) and performed in accordance with the Declaration of Helsinki. The reference number was 2013-1003. Written informed consent was obtained from all patients.
Consent to publish
Not applicable.
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
Competing interests
The authors declare no competing interests.
Funding information
This work was supported in part by the Japan Society for the Promotion of Science (JSPS) KAKEN, JSPS KAKENHI (18K08635 and 18K08664), and Grant from Daiwa Securities Health Foundation.
Additional information
Note This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Murahashi, S., Akiyoshi, T., Sano, T. et al. Serial circulating tumour DNA analysis for locally advanced rectal cancer treated with preoperative therapy: prediction of pathological response and postoperative recurrence. Br J Cancer 123, 803–810 (2020). https://doi.org/10.1038/s41416-020-0941-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-020-0941-4
- Springer Nature Limited
This article is cited by
-
Total neoadjuvant therapy in oesophageal and gastro-oesophageal junctional adenocarcinoma
British Journal of Cancer (2024)
-
ctDNA responds to neoadjuvant treatment in locally advanced rectal cancer
Journal of Cancer Research and Clinical Oncology (2024)
-
Assessing circulating tumour DNA (ctDNA) as a prognostic biomarker in locally advanced rectal cancer: a systematic review and meta-analysis
International Journal of Colorectal Disease (2024)
-
The clinical application of ctDNA to predict response to neoadjuvant chemoradiotherapy in patients with locally-advanced rectal cancer
Biomarker Research (2023)
-
Practical recommendations for using ctDNA in clinical decision making
Nature (2023)