Abstract
Up-regulation of Uridine-cytidine kinase 2 (UCK2), a rate-limiting enzyme of the pyrimidine salvage pathway, has been suggested in HCC, but the detailed molecular mechanisms and therapic role of UCK2 remain elusive. Bioinformatic analyses revealed that UCK2 might be a key up-regulated metabolic gene in HCCs. The expressional pattern and prognostic value of UCK2 were further examined in a large number of clinical samples. Functional assays based on site-directed mutagenesis showed that UCK2 promoted cell proliferation in a metabolic manner, but non-catalytically facilitates HCC metastasis. Mechanistically, in response to EGF, UCK2 interacted with EGFR to block EGF-induced EGFR ubiquitination and degradation, which resulted in elevated EGFR-AKT pathway activation and metastasis enhancement in HCCs. Concurrent pharmacological targeting on UCK2 and EGFR showed synergistic effects on HCC treatment. This study disclosed the non-metabolic role of UCK2 and suggested the therapeutic potential of concurrent blocking the metabolic and non-metabolic roles of UCK2 in HCC treatment.
Similar content being viewed by others
Introduction
China reports over half of the new cases and deaths of liver cancer10. Hepatocellular carcinoma (HCC), which accounts for 85–90% of primary liver cancers, affects more than 700,000 patients every year17. Despite the multiple curative treatments available29, the prognosis is still poor in HCC patients mainly due to the high recurrence rate of HCC18, which is closely related to tumor invasion and metastasis. Therefore, identifying novel targets for effective treatment of HCC metastasis is urgently required.
Epidermal growth factor receptor (EGFR) is a transmembrane receptor tyrosine kinase, whose overexpression and activation have been frequently reported in many types of cancers25. Overexpression of EGFR has been found in 60–85% of HCCs and significantly correlated with advanced stage, intrahepatic and extrahepatic metastasis, poor differentiation and tumor recurrence2,16,42. Although genomic EGFR amplification and mutations have been reported to contribute to EGFR overexpression and activation in various types of cancers, including glioblastoma14, oligodendroglioma11, non-small-cell lung cancer9 and gastric carcinomas32, such genomic alterations of EGFR are barely reported in HCCs8,13, suggesting the existence of other mechanisms leading to the high frequency of EGFR overexpression and activation in HCCs.
Uridine-cytidine kinase (UCK) is a type of rate-limiting enzymes in the salvage pathway of pyrimidine-nucleotide biosynthesis. By mono-phosphorylation, UCK respectively converts cytidine and uridine to cytidine 5′-monophosphate and uridine 5′-monophosphate4. There are two UCK genes in humans, UCK1 and UCK238. Despite the similarity of their catalytic functions, the catalytic efficacy of UCK2 in uridine and cytosine is 15–20 times higher than that of UCK138, suggesting that UCK2 mainly contributes to the salvage pyrimidine-nucleotide biosynthesis. UCK2 up-regulation has been detected in several types of cancers including pancreatic tumor, colorectal cancer, neuroblastoma, breast cancer, and HCCs28,37,40. Some recent studies have demonstrated the correlation of UCK2 with poor prognosis of HCCs40, but the underlying mechanism by which UCK2 contributes to HCC progression and the metabolic/non-metabolic roles of UCK2 in HCCs are largely undetermined.
In present study, we demonstrate that UCK2 is highly expressed in HCC tissues and serves as an independent predictor of poor prognosis, negatively correlating with overall survival (OS), recurrence-free survival (RFS) and early-RFS (within 2 years). Functionally, both in vitro and in vivo assays showed that UCK2 promotes proliferation and metastasis of HCC cells. Interestingly, the kinase activity of UCK2 is required for UCK2-mediated proliferation but not for UCK2-induced metastasis in HCCs. Mechanistically, UCK2 interacts with EGFR to inhibit EGF-induced EGFR ubiquitination and degradation, resulting in increased EGFR stability and sustained activation of the EGFR-AKT pathway. Finally, we demonstrate the synergistic potential of concurrent pharmacological targeting on UCK2 and EGFR in HCC treatment in both xenograft and patient-derived xenograft (PDX) models.
Materials and methods
Patients and samples
Fresh human HCC tumor tissues and matched adjacent non-tumor liver tissues were collected from patients who underwent surgical resection at the Department of Transplantation and Hepatic Surgery, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, between 2013 and 2017 for qRT-PCR (n = 120), western blot (n = 32), and immunohistochemistry (IHC) (n = 40) analyses. For clinical analyses, two independent TMA cohorts of HCC patients were retrospectively adopted. TMA cohort 1 included 153 HCCs collected between Feb 2004 and Aug 2008 from the Department of Transplantation and Hepatic Surgery, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, and TMA cohort 2 included 307 HCCs collected between June 2007 and June 2012 from the Shanghai Eastern Hepatobiliary Surgery Hospital. Written informed consent was obtained from each patient, and the studies were approved by Shanghai Jiao tong University School of Medicine, Renji Hospital Ethics Committee.
Establishment of patient-derived HCC cell lines
To establish the patient-derived HCC cell lines (PDC), fresh tumors were minced into ~1 mm3 fragments prior to enzymatic digestion using 1 mg/ml Collagenase type IV (17104019, Thermo Fisher, MA, USA) in DMEM/F12 (11320-033, Gibco, MA, USA), at 37 °C for 3 h. Then, cells were washed and re-suspended in phosphate buffered saline (PBS) (14190235, Thermo Fisher, MA, USA) for three cycles. The final cell suspensions were filtered with 70 µm cell-strainers (352350, Falcon, CA, USA) and centrifuged at 600 g for 5 min at 4 °C. The cell pellets were re-suspended in DMEM (12430-054, Gibco, MA, USA), supplemented with 10% (v/v) fetal bovine serum (FBS) (10099, Gibco, MA, USA), and 1% penicillin-streptomycin (15070063, Gibco, MA, USA) and cultured.
Cell lines
The normal hepatocyte LO2 cells, as well as the commercially available HCC cell lines Huh-7, HepG2, HCC-LM3, SMMC-7721 and MHCC-97L, were purchased from the Shanghai Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Home-made patient-derived HCC cell lines PDC-26#, PDC-14#, PDC-12#, PDC-9#, PDC-23#, PDC-10# and PDC-11# were established as described above. All of these cells were cultured in Dulbecco’s modified eagle medium (DMEM) supplemented with 10% (v/v) FBS and 1% penicillin-streptomycin at 37 °C in a humidified incubator under 5% CO2 condition.
Phospho-antibody array
Huh-7 cells with stable overexpression of UCK2WT or UCK2D62A, and control cells were gathered. The cell lysates were then obtained and applied to a Cancer Signaling Phospho-Antibody Array (PCS248, Full Moon Biosystems, CA, USA). The array experiment was performed by Wayen Biotechnologies (Shanghai, China) Inc. according to the manufacturer’s protocol. The array contained 269 site-specific and phospho-specific antibodies against total 93 proteins. Each specific site or protein contains six technical replicates. The slide was scanned on a GenePix 4000B scanner (Axon Instruments, CA, USA), and the images were analyzed with GenePix Pro 6.0. The fluorescence intensity of each array spot was quantified, and the mean value was calculated. The following equation was used to calculate a phosphorylation signal intensity (Phos-In): Phos-In = phospho A/unphospho A
where phospho A represented the signal from phosphorylated A protein, and unphospho A represented the signal from unphosphorylated A protein in the experimental samples.
Proximity ligation assay
Proximity ligation assay was performed using the Duolink In Situ Red Starter Kit Mouse/Rabbit (Sigma-Aldrich, MO, USA). Briefly, cells were cultured on chamber slides, fixed with paraformaldehyde for 3 min. After washing, cells were permeabilized with 0.1% Triton X-100 in PBS for 5 min, blocked with PLA blocking buffer for 1 h and incubated with primary antibodies for overnight at 4 °C. Following three washes with PBS the cells were incubated with anti-rabbit-PLUS (Sigma-Aldrich; DUO92002 for UCK2 antibody) and anti-mouse-MINUS (Sigma-Aldrich; DUO92004 for EGFR antibody) PLA probes and subjected to ligation and amplification reaction using Duolink In Situ Detection Reagents Red (Sigma-Aldrich; DUO92008) according to the manufacturer’s protocol. The cells were mounted with DAPI and visualized with Leica DM IL LED-Inverted fluorescence microscope.
Dose matrix experiments
For dose matrix experiments, cells were seeded in 96-well plates at 5 × 103 cells per well. After 24 h, cells were treated with ECyd (dose range of 0–27 nM) and Gefitinib (dose range of 0–25 μM) in a 7 × 7 matrix. Cells were cultured with inhibitors for 72 h and cell viability was determined using CCK-8. Each treatment was done in triplicate wells. The following equation was used to calculate the Bliss Score: Bliss Score = AB − [(A + B) − A × B]. At a given dose, AB represented the fractional growth inhibitions induced by combined treatment with A and B, while A and B represented the fractional growth inhibitions induced by separate usage of A or B.
Generation of PDXs
Tumor samples were obtained from patients post surgery after obtaining informed patient consent in accordance to the ethical review committee of the WHO Collaborating Center for Research in Human Production (authorized by Shanghai Municipal Government). Tumors were minced into ~1 mm3 fragments and suspended in a mixture of 5% Matrigel (356234, Corning, NY, USA) in DMEM/F12. The tumor fragment mixtures were then implanted subcutaneously into the left and right flanks of 5–7 weeks old NSG mice (Biocytogen, Beijing, China), using 18-gauge needles. Tumors were excised and passaged when they reached 1.5 cm3. For tumor passage, tissues were cut into small fragment of 1 mm3 prior to resuspension in 20% Matrigel/DMEM/F12 mix, before subcutaneous inoculation of tumor fragments into 5–7 weeks old BALB/C nude mice.
Statistical analysis
Statistical analyses were performed with SPSS version 18.0 software. Graphical representations were performed with GraphPad Prism 6 software. Differentially expressed genes were obtained using online analysis tool GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/). The chi-square test or Fisher’s exact test were used to compare qualitative variables, and the Student’s t test or the Mann–Whitney test were used to compare continuous variables. Kaplan–Meier analysis was used to assess survival. Log-rank tests were used to compare survival of patients between subgroups. Multivariate analyses were performed by multivariate Cox proportional hazard regression model. Data were presented as mean ± SD. A difference was defined as significant at p < 0.05.
Results
Identification of UCK2 as a key up-regulated metabolic gene in HCCs
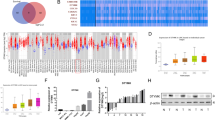
GSE14520, one HCC dataset in Gene Expression Omnibus (GEO) database that contains largest number of samples and detailed clinical prognostic information, was adopted to screen out dysregulated metabolic genes (MDEGs) in HCCs. Firstly, 2772 dysregulated genes [p value < 0.05, fold change (FC) > 1.5 or <−1.5] were identified by GEO2R, 62.2% of which were up-regulated and 37.8% down-regulated in HCC tissues (Fig. S1A). Meanwhile, 1151 genes assigned to metabolic pathways in the Kyoto Encyclopedia of Genes and Genomes database were gathered as metabolic genes (Fig. S1B). Finally, 155 up-regulated metabolic genes (uMDEGs) and 287 down-regulated metabolic genes were obtained by the overlapping analysis of these two gene sets (Fig. S1C). Heatmap assays according to these dysregulated metabolic genes apparently separated tumor and non-tumor tissues (Fig. 1A). Here, we are more interested in uMDEGs in HCCs. The top ten uMDEGs were listed according to the p values (Fig. 1B). Prognostic analyses showed that UCK2 was the most effective uMDEG to differentiate the OS, RFS and early-RFS (within 2 years) of HCC patients in GSE14520 (Fig. S1D–S1F).
a Heatmap of differentially expressed metabolic genes in GSE14520. b The top ten up-regulated metabolic genes in GSE14520 according to the p values. c qRT-PCR analyses of UCK2 mRNA levels in 120 pairs of HCC tumor and adjacent non-tumor tissues. d UCK2 protein levels in 32 pairs of HCC tumor and adjacent non-tumor tissues detected by western blotting. e UCK2 protein levels in 40 pairs of HCC tumor and adjacent non-tumor tissues detected by IHC assays. f–g Kaplan–Meier analyses of overall survival (OS), recurrence-free survival (RFS) and early-RFS (within 2 years) of HCC patients in correlation with high or low UCK2 levels in TMA Cohort 1 (f) and TMA Cohort 2 (g). The absolute number of patients at risk is listed below each curve.
Consistent with the analyzed results and previous reports40, UCK2 up-regulation in HCCs was confirmed in our great number of clinical samples (Fig. 1C–E). Furthermore, the prognostic value of UCK2 in HCCs was confirmed in two independent HCC TMA cohorts (Fig. S1G and Table S1). Clinical association analyses revealed that increased UCK2 levels were significantly associated several aggressive clinicopathological features (Table S2). Moreover, Kaplan–Meier analyses showed that UCK2 negatively correlates with OS, RFS and early-RFS in HCC patients (Fig. 1F, G), which was in line with the analyzed results of GSE14520. Multivariate Cox Regression showed that UCK2 as an independent prognostic predictor for OS, RFS, and early-RFS (Table S3, 4). Furthermore, UCK2 also separated favorable and poor prognosis on OS, RFS and early-RFS in HCC patients at the same TNM stage (Fig. S1H–J). These findings indicate that UCK2 might be a key metabolic gene, which is commonly up-regulated and associated with poor prognosis in HCCs.
UCK2 promotes HCC proliferation, migration and invasion in vitro
UCK2 expression was first examined in 13 cell lines including one normal hepatocyte cell lines (LO2), five commercial available HCC cell lines (Huh-7, HepG2, HCC-LM3, SMMC-7721, and MHCC-97L) and seven home-made primary HCC cell lines (PDC-26#, PDC-14#, PDC-12#, PDC-9#, PDC-23#, PDC-10#, and PDC-11#). Enhanced UCK2 expression was observed in the 12 HCC cell lines compared with LO2 cells (Fig. S2A). However, UCK1 did not exhibit such expression differences (Fig. S2B). Furthermore, HCC cells were chosen for loss- or gain-of-function studies due to their high or low endogenous UCK2 levels. UCK2 knockdown or overexpression was confirmed (Fig. S2C, F). CCK8, colony formation, and Edu incorporation assays showed reduced proliferation in HCC cells with UCK2 knockdown (Figs. 2A, S2D, E) and enhanced proliferation with UCK2 overexpression (Figs. 2D, S2G, H). In addition, transwell-migration and -invasion assays showed that UCK2 knockdown or overexpression respectively impaired or promoted migration and invasion abilities of HCC cells (Fig. 2B-C and E–F). These findings indicate that UCK2 has an oncogenic role in promoting HCC proliferation, migration, and invasion in vitro.
a CCK-8 assays were performed to test the effects of UCK2 knockdown on the proliferation of MHCC-97L and PDC-11# cells in vitro. b–c Transwell-migration (b) and -invasion (c) assays were performed to test the effects of UCK2 knockdown on the migration and invasion abilities of MHCC-97L and PDC-11# cells in vitro. d CCK-8 assays were performed to test the effects of UCK2 overexpression on the proliferation of Huh-7 and PDC-26# cells in vitro. e–f Transwell-migration (e) and -invasion (f) assays were performed to test the effects of UCK2 overexpression on the migration and invasion abilities of Huh-7 and PDC-26# cells in vitro. Scale bar = 100 μm. *, p < 0.05; **, p < 0.01; ***, p < 0.001. Error bars indicate means ± SD.
UCK2-mediated metastasis enhancement in HCCs is independent on its catalytic activity
To investigate whether the oncogenic role of UCK2 is dependent on its catalytic activity, we replaced UCK2 Asp62, a conserved amino acid residue which was found to be essential for UCK2’s catalytic activity30,33, with Ala (UCK2D62A), and stable cell lines were respectively established (Fig. S3A). Neither UCK2WT nor UCK2D62A had a significant effect on UCK1 expression (Fig. S3B). The catalytic activity loss of UCK2D62A was examined by cytotoxicity assays by using the cytotoxic pyrimidine analogues 5-Fluorouridine (5-FUrd) and 3′-C-ethynylcytidine (ECyd)37. Unlike UCK2WT, UCK2D62A failed to sensitize Huh-7 and PDC-26# cells to 5-FUrd and ECyd (Fig. S3C, D), suggesting the deprived catalytic activity in UCK2D62A.
Correspondingly, CCK8, colony formation and Edu incorporation assays demonstrated that, compared with UCK2WT, UCK2D62A lost the growth advantages (Figs. 3A and S3E–F). Interestingly, both UCK2WT and UCK2D62A showed comparable migration and invasion enhancement (Fig. 3B, C), suggesting that UCK2 may non-catalytically promotes HCC metastasis. In line with our in vitro findings, the in vivo xenograft model showed that UCK2 knockdown significantly inhibited tumor growth (Fig. S3G), whereas UCK2WT but not UCK2D62A greatly accelerated tumor growth (Fig. 3D). Meanwhile, the lung metastasis model showed that UCK2 knockdown impaired lung metastasis (Fig. S3H), whereas both UCK2WT and UCK2D62A greatly increased lung metastasis although the size of metastatic foci were smaller in UCK2D62A group (Fig. 3E). Similar results were observed in intrahepatic metastastic models (Figs. 3F and S3I). Therefore, these findings indicate that UCK2-mediated metastasis enhancement is independent on its catalytic activity.
a CCK-8 assays were performed to test the effects of UCK2WT or UCK2D62A overexpression on the proliferation of Huh-7 and PDC-26# cells in vitro. b–c Transwell-migration (b) and -invasion (c) assays were performed to test the effects of UCK2WT or UCK2D62A overexpression on the migration and invasion abilities of Huh-7 and PDC-26# cells in vitro. d Tumor growth curves (left), end-point tumor photography (middle) and tumor weight (right) in Huh-7 cells with either UCK2WT or UCK2D62A overexpression. e Representative images (left) and statistic results (right) of lung sections with metastasis foci in Huh-7 cells with either UCK2WT or UCK2D62A overexpression in a pulmonary metastatic model. f Representative images (left) and statistic results (right) of intrahepatic metastasis foci in Huh-7 cells with either UCK2WT or UCK2D62A overexpression in a orthotopic implanted intrahepatic metastastic model. Scale bar = 100 μm. **, p < 0.01; n.s no significance. Error bars indicate means ± SD.
AKT activation is required for the non-catalytic role of UCK2 in promoting HCC metastasis
To investigate the mechanism by which UCK2 non-catalytically promotes HCC metastasis, a cancer signaling phospho-antibody array was performed (Fig. S4A). Compared with the vector control, 14 and 10 phospho-candidates with significant phosphorylation intensity (Phos-Ins) alterations (FC ≥ 1.5 or ≤−1.5) were respectively selected from Huh-7 cells with either UCK2WT or UCK2D62A overexpression (Fig. 4A and Table S5). Phosphorylation at the Ser473 of AKT1 was the most elevated one in the eight common altered phospho-sites between UCK2WT and UCK2D62A groups (Fig. 4B and Table S5). Furthermore, increased or decreased phosphorylation at AKT1 on Ser473 was confirmed in HCC cells with UCK2WT/UCK2D62A overexpression or UCK2 knockdown, respectively (Fig. 4C, D). To determine whether AKT1 activation was required for UCK2WT/UCK2D62A-mediated metastasis enhancement. MK-2206 (an AKT inhibitor) (Fig. S4B) or AKT1 knockdown (Fig. S4C) significantly inhibited UCK2WT/UCK2D62A-induced migration (Fig. 4E, G) and invasion (Fig. 4F, H) in vitro. Moreover, AKT1 activation, by either SC79 (an activator of AKT) (Fig. S4D) or pmyr-AKT (a constitutively active AKT) (Fig. S4E), greatly rescued the inhibition on migration and invasion induced by UCK2 knockdown (Fig. 4I–L). These findings indicate that UCK2 can non-catalytically induce AKT1 activation, which is required for the non-catalytic role of UCK2 in promoting HCC metastasis.
a Eight common phosphorylation changes collected from Huh-7 cells with UCK2WT or UCK2D62A overexpression were presented in a Venn diagram. b Details of the eight common phosphorylation signal changes. c–d Confirmation of UCK2-induced AKT activation by western blot in Huh-7 and PDC-26# cells with UCK2WT/UCK2D62A overexpression (c) and in MHCC-97L and PDC-11# cells with UCK2 knockdown (d). e–f Transwell-migration (e) or -invasion (f) assays in Huh-7 cells with UCK2WT/UCK2D62A overexpression and MK-2206-mediated AKT inhibition. g–h Transwell-migration (g) or -invasion (h) assays in Huh-7 cells with UCK2WT/UCK2D62A overexpression and siRNA-mediated AKT downregulation. i–j Transwell-migration (i) or -invasion (j) assays in MHCC-97L cells with endogenous UCK2 knockdown and treatment of an AKT1 activator, SC79. k–l Transwell-migration (k) or -invasion (l) assays in MHCC-97L cells with endogenous UCK2 knockdown and overexpression of constantly activated AKT1, pmyr-AKT1. Scale bar = 100 μm. **, p < 0.01. Error bars indicate means ± SD.
UCK2 non-catalytically induces AKT activation through enhanced EGFR pathway activity
Receptor Tyrosine Kinases (RTKs) are a family of proteins whose activation can activate oncogenic AKT23. To further investigate the mechanism by which UCK2 non-catalytically induces AKT1 activation, we performed a second-round scrutiny focusing on phosphorylation changes in RTKs in our phospho-antibody array data. UCK2WT/UCK2D62A-induced phosphorylation elevation was only observed in EGFR but not in other RTKs such as HER2, IGF1R, VEGFR2 and so on (Table S6). Interestingly, in addition to inducing EGFR phosphorylation, UCK2WT/UCK2D62A parallel increased EGFR protein levels, which results in comparable Phos-Ins of EGFR in cells transfected with UCK2WT, UCK2D62A or control vectors (Table S6), explaining why we did not find significant Phos-In alterations in EGFR (Fig. 4B and Table S5). In addition, UCK2-mediated EGFR activation was confirmed by western blotting (Fig. 5A, B). Consistent with our array data, UCK2 had no effect on HGF-induced AKT activation but greatly affected EGF-induced AKT activation (Fig. S5A–D). Interestingly, the phosphorylation levels of other downstream targets of EGFR pathway (p-p38, p-ERK1/2 and p-JNK1/2) did not appear to change significantly when UCK2 was overexpressed or interfered (Fig. S5E–F), which was in line with our array data. Moreover, UCK1 loss or overexpression did not affect EGFR signaling, including changes in Akt phosphorylation (Fig. S5G–H). These findings suggest that UCK2 may non-catalytically induce AKT activation through activating EGFR pathway.
a–b Confirmation of UCK2 non-catalytically inducing EGFR activation by western blot to probe total and phosphorylated EGFR levels [p-EGFR(Y992), p-EGFR(Y1045), p-EGFR(Y1068) and p-EGFR(Y1110)] in Huh-7 and PDC-26# cells with UCKWT/UCK2D62A overexpression (a) or in MHCC-97L and PDC-11# cells with endogenous UCK2 knockdown (b). c–d Cycloheximide (CHX) chase assays to evaluate the role of UCK2 on EGF-induced EGFR degradation in Huh-7 cells with UCKWT/UCK2D62A overexpression (c) or in MHCC-97L cells with endogenous UCK2 knockdown (d). e–f Proteasomal degradation was inhibited by MG-132, and EGFR protein levels were determined in Huh-7 cells with UCKWT/UCK2D62A overexpression (e) or in MHCC-97L cells with endogenous UCK2 knockdown (f). g–h Detection of EGFR ubiquitination levels in Huh-7 cells with UCK2WT/UCK2D62A overexpression (g) or in MHCC-97L cells with endogenous UCK2 knockdown (h) in the context of with or without EGF treatment.
UCK2 non-catalytically stabilizes EGFR proteins by blocking EGF-induced EGFR ubiquitination
Given that UCK2 non-catalytically increases EGFR levels and activates EGFR-AKT pathway, we first examined whether UCK2 regulates EGFR transcriptionally. qRT-PCR assays indicated that either UCK2WT/UCK2D62A overexpression or UCK2 knockdown had no effect on EGFR transcription (Fig. S5I, J), suggesting that UCK2 may non-catalytically affect EGFR protein stability. Cycloheximide (CHX) chase assays showed that, in response to EGF treatment, UCK2WT/UCK2D62A overexpression or UCK2 knockdown greatly retarded or accelerated EGFR degradation, respectively (Fig. 5C, D). Moreover, these differences disappeared when protein degradation was inhibited by proteasome inhibitor MG-132, confirming the non-catalytical effect of UCK2 on EGFR degradation (Fig. 5E, F). Ubiquitination of EGFR is critical for EGF-induced EGFR degradation via either proteasomal or lysosomal degradation pathways12,27, we investigated whether UCK2 non-catalytically affects EGF-induced EGFR ubiquitination. As shown in Fig. 5G, H, UCK2WT/UCK2D62A overexpression greatly blocked EGF-induced EGFR ubiquitination whereas UCK2 knockdown significantly enhanced this process. These findings suggest that UCK2 non-catalytically stabilizes EGFR proteins by inhibiting EGF-induced EGFR ubiquitination.
Physical interaction of UCK2-EGFR is required for the non-catalytic activities of UCK2
A previous EGFR interactome study has suggested a possible EGFR-UCK2 interaction34. Here, physical UCK2-EGFR interaction was confirmed by reciprocal IP in Huh-7 cells with UCK2WT/UCK2D62A overexpression and in MHCC-97L cells with high endogenous UCK2 expression (Fig. 6A, B). Such a UCK2-EGFR interaction was exclusively detectable in the presence of EGF (Fig. S6A, B). Moreover, the proximity ligation assay confirmed the endogenous interaction of UCK2 and EGFR in presence of EGF (Fig. S6C). To investigate whether UCK2-EGFR interaction is required for the non-catalytic activities of UCK2, we constructed a series of truncated UCK2 mutants (Fig. S6D, E). Reciprocal IP showed that the mutant without N-terminal half (UCK2Δ1-124) failed to interact with EGFR (Figs. 6C and S6F). CHX chase assays showed that UCK2FL or UCK2Δ125-261 significantly delayed EGF-induced EGFR degradation, whereas UCK2Δ1-124 failed to do so (Fig. 6D). Furthermore, unlike UCK2FL and UCK2Δ125-261 which significantly blocked EGF-induced EGFR ubiquitination, UCK2Δ1-124 had no effect on this process (Fig. 6E). Correspondingly, EGF treatment greatly enhanced EGFR-AKT activation in Huh-7 cells overexpressing UCK2FL or UCK2Δ125-261 but not in cells overexpressing UCK2Δ1-124 (Fig. 6F). In line with this notion, compared with UCK2Δ125-261, UCK2Δ1-124 failed to enhance in vitro migration/invasion and in vivo lung metastasis (Fig. 6G, H), although both UCK2Δ125-261 and UCK2Δ1-124 showed no effect on HCC proliferation (Fig. 6I, J). Therefore, these findings indicate that the UCK2-EGFR interaction is critical for the non-catalytic roles of UCK2 in HCC.
a–b Reciprocal IP assays of EGFR and UCK2 in Huh-7 cells with UCK2WT/UCK2D62A overexpression (a) or in MHCC-97L cells with high endogenous UCK2 expression (b). c IP assays of EGFR and UCK2 in Huh-7 cells overexpressing indicated UCK2 full length and mutants. d Cycloheximide (CHX) chase assays to examine EGFR protein stability in response to EGF in Huh-7 cells overexpressing indicated UCK2 full length and mutants. e After 18 h of serum starvation, EGFR ubiquitination levels were detected in Huh-7 cells overexpressing indicated UCK2 full length and mutants. f After 18 h of serum starvation, EGF-induced EGFR-AKT activation was detected of in Huh-7 cells overexpressing indicated UCK2 full length and mutants with or without EGF treatment. g Transwell-migration and -invasion assays in Huh-7 cells overexpressing indicated UCK2 full length and mutants. h Representative images (left half) and statistic results (right half) of lung sections with metastasis foci in Huh-7 cells overexpressing indicated UCK2 full length and mutants in a pulmonary metastatic model. i–j CCK-8 (i) and EdU incorporation (j) assays were performed to test the effects of UCK2FL, UCK2Δ125-261, or UCK2Δ1-124 on the proliferation of Huh-7 cells in vitro. Scale bar = 100 μm. **, p < 0.01. Error bars indicate means ± SD.
Concurrent pharmacological targeting UCK2 and EGFR leads to synergistic suppression of HCC progression
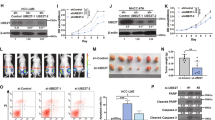
Since the metabolic and non-metabolic roles of UCK2, we speculated synergistical inhibition on HCC might be achieved by concurrent pharmacologic targeting UCK2 and EGFR. To test this point, we first examined whether UCK2 would affect the efficacy of cytotoxic pyrimidine analogues (5-Furd or ECyd) and EGFR inhibitors (Erlotinib or Gefitinib) in HCCs. UCK2 knockdown induced significant resistance on cytotoxic pyrimidine analogues (fold change of IC50: 2.88-fold for 5-FUrd and 3.27-fold for ECyd; Fig. S7A) and EGFR inhibitors (fold change of IC50: 2.61-fold for Erlotinib and 2.89-fold for Gefitinib; Fig. S7B), whereas overexpression greatly sensitized to them (fold change of IC50: 0.27-fold for 5-FUrd, 0.31-fold for ECyd, 0.35-fold for Erlotinib and 0.34-fold for Gefitinib; Fig. S7C, D), suggesting the synergistic potential on targeting both UCK2 and EGFR. Then, dose matrix experiments were performed in combination of ECyd and Gefitinib, and synergistic effects were analyzed using the Bliss independent model41. Combined treatment of ECyd and Gefitinib was more effective in MHCC-97L (high endogenous UCK2 expression) than in Huh-7 cells (low endogenous UCK2 expression) (Fig. S7E–G). Consistently, UCK2 knockdown abrogated such synergistic effects (Fig. 7A, B), while overexpression enhanced them (Fig. 7C, D). More importantly, the xenograft mouse models further validated the synergistic effects of the combined ECyd and Gefitinib treatment in inhibiting tumor growth in MHCC-97L (Fig. 7E, F), and in Huh-7 cells overexpressing UCK2 (Fig. 7G, H). Body weights of mice had not significantly changed between different treatment groups (Fig. S7H, I). These data indicate that concurrent pharmacological targeting UCK2 and EGFR has synergistic effect on inhibition of HCC progression.
a–d A bliss independent model to evaluate the synergistic cytotoxic effects of concurrent targeting EGFR by Gefitinib and UCK2 by ECyd in MHCC-97L cells with endogenous UCK2 knockdown (a–b) or in Huh-7 cells with UCK2 overexpression (c–d). e–h Tumor volume, tumor inhibitory rate, end-point tumor photography and tumor weight were presented in the xenograft mouse models to evaluate the synergistic effects of concurrent targeting EGFR by Gefitinib and UCK2 by ECyd on the inhibition of tumor growth in MHCC-97L cells with or without endogenous UCK2 knockdown (e–f) or in Huh-7 cells with or without UCK2 overexpression (g–h). *, p < 0.05; **, p < 0.01; ***, p < 0.001; n.s no significance. Error bars indicate means ± SD.
Preclinical evaluation of the synergistic suppression in patient-derived HCC cells and xenografts
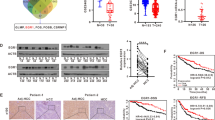
Patient-derived cells (PDC) and xenograft (PDX) models were used to predict the clinical responses to such a combined treatment in HCC patients6,15. In line with commercial available HCC cells, UCK2 knockdown induced significant resistance on cytotoxic pyrimidine analogues and EGFR inhibitors in PDC-11# cells (Fig. S8A, B), whereas overexpression UCK2 in PDC-26# cells greatly sensitized to them (Fig. S8C, D). Moreover, dose matrix experiments showed that the combined treatment was more effective in PDC-11# (high endogenous UCK2 expression) than in PDC-26# cells (low endogenous UCK2 expression) (Fig. S8E–G). UCK2 knockdown in PDC-11# cells abrogated such synergistic effects (Fig. 8A, B), whereas overexpression enhanced them in PDC-26# cells (Fig. 8C, D). Furthermore, in vivo combined ECyd and Gefitinib treatment showed synergistic effects in 3 out of 4 PDX tumors (Fig. 8E–H). More importantly, high UCK2 and EGFR protein and EGFR activation levels were observed in all the three responsive PDX tumors by western blot (Fig. 8I). Meanwhile, IHC assays confirmed high UCK2 and EGFR protein levels in all the three responsive PDX tumors (Fig. 8J). In addition, either single or combined drug treatment did not cause significant body weight changes (Fig. S8H). These findings preclinically confirmed the pharmacological efficacy and safety of simultaneous targeting the metabolic and non-metabolic functions of UCK2 in HCC.
a–d A bliss independent model to evaluate the synergistic cytotoxic effects of concurrent targeting EGFR by Gefitinib and UCK2 by ECyd in PDC-11# cells with endogenous UCK2 knockdown (a–b) or in PDC-26# cells with UCK2 overexpression (c–d). e–h Tumor volume, tumor inhibitory rate, end-point tumor photography and tumor weight were presented in four established PDX HCC tumors (e, PDX-1#; f, PDX-2#; g, PDX-3#; h, PDX-4#) in response to treatments of vehicle, ECyd, Gefitinib, or combined ECyd and Gefitinib. i Protein levels of EGFR, UCK2 and phosphorylated EGFR and AKT were detected in the four PDX HCC tumors by western blot. j Protein levels of EGFR and UCK2 were detected in the four PDX HCC tumors by IHC assays. *, p < 0.05; **, p < 0.01; ***, p < 0.001; n.s no significance. Error bars indicate means ± SD.
Discussion
In present study, we confirm up-regulated UCK2 serves as an independent risk factor of poor prognosis in HCCs. UCK2 plays a pro-oncogenic role by promoting tumor cell proliferation and metastasis. Interestingly, UCK2 catalytically promotes tumor cell proliferation but non-catalytically enhances tumor cell metastasis. Mechanistically, UCK2 can non-catalytically activate EGFR-AKT pathway by interacting with EGFR to inhibit EGF-induced EGFR ubiquitination and degradation, resulting in enhanced HCC metastasis. Pharmaceutically, we demonstrate the synergistic effects in treatment of HCCs by concurrently targeting UCK2 and EGFR in both xenograft and PDX models.
Despite the early observation of aerobic glycolysis by Otto Warburg (the ‘Warburg effect’) in the 1960’s, cancer associated metabolic reprogramming has been long neglected due to simple interpretation of this alteration as a passive response to fulfill the high energy and nutrient demands of tumor cells39. Recently, some studies have demonstrated the existence of onco-metabolites to activate oncogenic signalings to potentiate malignant transformation, suggesting metabolism reprogramming as a driven force of cancer39. Previous metabolism investigations mainly focus on the role of metabolic reprogramming in promoting tumor cell proliferation20, leading to the generation of most metabolism-based anticancer drugs targeting cancer cell proliferation. Recently, the pro-metastasis role of metabolism reprogramming has been revealed. For example, elevated methylglyoxal (MG) levels induced by aerobic glycolysis enhanced the growth and metastasis in breast cancer cells by increasing nuclear accumulation of YAP;26 and elevated free fatty acid levels and uptake contribute to epithelial-mesenchymal transition induction in HCC cells by activating Wnt and TGF-β signaling24. Given that metastasis is a less frequent phenotype in normal cells compared with proliferation, targeting metastasis may serve as a more selective strategy in developing novel metabolism-based anticancer drugs. In the current study, we demonstrate that UCK2 promotes both proliferation and metastasis in HCCs, suggesting that UCK2 is a potential metabolism-based therapeutic target for HCC treatment.
In the context of metabolism reprogramming, many metabolic enzymes, involved in glycolysis19, lipogenesis31, amino acid biosynthesis3 and nucleotide biosynthesis35, are up-regulated to support anabolic growth of cancer cells. To further complicate the matter, some recent studies even demonstrate the non-metabolic roles of some key metabolic enzymes, such as PKM2 and PGK1, in promoting tumor progression22. One kind of interesting non-metabolic role comes from the kinase activity of those metabolism enzymes to phosphorylate non-metabolite substrates, such as proteins. For examples, PKM2 has been reported to function as a kinase of multiple proteins including histone H3, STAT3 and SREBPs to promote tumor progression; and PGK1 phosphorylates Beclin1 and pyruvate dehydrogenase kinase isozyme 1 to support tumor development22. In the current study, we demonstrate that UCK2 also has a non-metabolic role in promoting HCC development. Interestingly, this non-metabolic role of UCK2 does not require its kinase activity, because kinase-dead UCK2 (UCK2D62A) remains functional in promoting HCC metastasis. This finding expands our understanding of the non-metabolic roles performed by metabolic enzymes.
Genetic studies have disclosed the high correlation of genomic EGFR amplification/mutations with EGFR overexpression/activation in many malignancies9,11,14,32, but EGFR amplification/mutations are rare in HCCs8,13, despite the high EGFR overexpression reported in 60–85% of HCCs2,16. We demonstrate that UCK2 interacts with EGFR to impair EGF-induced EGFR ubiquitination, resulting in stabilization of EGFR. Notably, our finding is a bit discrepant with a previous EGFR interactome study showing that such an EGFR-UCK2 interaction was in an EGF-independent manner. Such an EGF-independent EGFR-UCK2 interaction may result from the possible autonomous activation of EGFR due to their overexpression of EGFR in HEK293 cells5,34. We demonstrate that the presence of EGF is required for EGFR-UCK2 interaction, suggesting that EGF-induced EGFR modification or internalization may be required for EGFR-UCK2 interaction.
Although pyrimidine biosynthesis has been identified as one of ideal targets for cancer treatment due to its frequent up-regulation in many malignancies7, several clinical trials on the inhibitors antagonizing key enzymes of de novo pyrimidine biosynthesis have been failed partly because of the activation and up-regulation of the salvage pyrimidine biosynthesis21. For example, compared with normal colon, the activity elevation of UCK (in salvage pathway) was much higher than that of orotidine 5′-phosphate (OMP) decarboxylase (in de novo pathway)1. As a rate-limiting enzyme of salvage pyrimidine biosynthesis, UCK2 overexpression has been reported in many types of cancers including HCCs40, but the biological function of UCK2 in HCCs is underexplored. In addition to our finding of UCK2-mediated HCC proliferation, one most interesting finding of this study is that we unexpectedly found that UCK2 can non-catalytically promote HCC metastasis. These findings suggest that UCK2 may have divergent pro-tumor functions either dependent or independent on its catalytic activity. This may partially explain the failure of a clinical trial of nucleoside 3′-C-ethynylcytidine (TAS-106), a cytidine analogue subjected to UCK2 phosphorylation, in treatment of head and neck cancer and nasopharyngeal cancer36.
In the current study, we demonstrate that UCK2 catalytically promotes HCC proliferation and non-catalytically enhances HCC metastasis by activating the EGFR-AKT pathway, which inspired us to design a combined drug treatment to simultaneously block the catalytic dependent and independent functions of UCK2. Concurrent pharmacological targeting on UCK2 by ECyd and EGFR by Gefitinib showed synergistic effects in treatment of HCCs, suggesting that the non-metabolic roles of targeted metabolic enzymes may need to be considered in metabolism-based therapy for better efficacy.
References
Ahmed, N. K. Enzymes of the de novo and salvage pathways for pyrimidine biosynthesis in normal colon, colon carcinoma, and xenografts. Cancer 54, 1370–1373 (1984).
Altimari, A. et al. Investigation of ErbB1 and ErbB2 expression for therapeutic targeting in primary liver tumours. Dig. Liver Dis. 35, 332–338 (2003).
Amelio, I., Cutruzzola, F., Antonov, A., Agostini, M. & Melino, G. Serine and glycine metabolism in cancer. Trends Biochem. Sci. 39, 191–198 (2014).
Appleby, T. C. et al. Structure of human uridine-cytidine kinase 2 determined by SIRAS using a rotating-anode X-ray generator and a single samarium derivative. Acta Crystallogr. Sect. D., Biol. Crystallogr. 61, 278–284 (2005).
Arteaga, C. L. & Engelman, J. A. ERBB receptors: from oncogene discovery to basic science to mechanism-based cancer therapeutics. Cancer Cell 25, 282–303 (2014).
Bissig-Choisat, B. et al. Novel patient-derived xenograft and cell line models for therapeutic testing of pediatric liver cancer. J. Hepatol. 65, 325–333 (2016).
Bokkerink, J. P. et al. Sequence-, time- and dose-dependent synergism of methotrexate and 6-mercaptopurine in malignant human T-lymphoblasts. Biochem. Pharmacol. 35, 3549–3555 (1986).
Buckley, A. F., Burgart, L. J., Sahai, V. & Kakar, S. Epidermal growth factor receptor expression and gene copy number in conventional hepatocellular carcinoma. Am. J. Clin. Pathol. 129, 245–251 (2008).
Cappuzzo, F. et al. Epidermal growth factor receptor gene and protein and gefitinib sensitivity in non-small-cell lung cancer. J. Natl Cancer Inst. 97, 643–655 (2005).
Chen, W. et al. Cancer statistics in China, 2015. CA: Cancer J. Clin. 66, 115–132 (2016).
Fallon, K. B. et al. Prognostic value of 1p, 19q, 9p, 10q, and EGFR-FISH analyses in recurrent oligodendrogliomas. J. Neuropathol. Exp. Neurol. 63, 314–322 (2004).
Feng, F. Y. et al. Role of epidermal growth factor receptor degradation in gemcitabine-mediated cytotoxicity. Oncogene 26, 3431–3439 (2007).
Galluzzi, L., Kepp, O., Vander Heiden, M. G. & Kroemer, G. Metabolic targets for cancer therapy. Nat. Rev. Drug Discov. 12, 829–846 (2013).
Hatanpaa, K. J., Burma, S., Zhao, D. & Habib, A. A. Epidermal growth factor receptor in glioma: signal transduction, neuropathology, imaging, and radioresistance. Neoplasia 12, 675–684 (2010).
Hidalgo, M. et al. Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov. 4, 998–1013 (2014).
Ito, Y. et al. Expression and clinical significance of erb-B receptor family in hepatocellular carcinoma. Br. J. Cancer 84, 1377–1383 (2001).
Lafaro, K. J., Demirjian, A. N. & Pawlik, T. M. Epidemiology of hepatocellular carcinoma. Surgical Oncol. Clin. North Am. 24, 1–17 (2015).
Lee, S. C., Tan, H. T. & Chung, M. C. Prognostic biomarkers for prediction of recurrence of hepatocellular carcinoma: current status and future prospects. World J. Gastroenterol. 20, 3112–3124 (2014).
Li, X. B., Gu, J. D. & Zhou, Q. H. Review of aerobic glycolysis and its key enzymes - new targets for lung cancer therapy. Thorac. Cancer 6, 17–24 (2015).
Llovet, J. M. & Bruix, J. Molecular targeted therapies in hepatocellular carcinoma. Hepatology 48, 1312–1327 (2008).
Loffler, M., Fairbanks, L. D., Zameitat, E., Marinaki, A. M. & Simmonds, H. A. Pyrimidine pathways in health and disease. Trends Mol. Med. 11, 430–437 (2005).
Lu, S. & Wang, Y. Nonmetabolic functions of metabolic enzymes in cancer development. Cancer Commun. 38, 63 (2018).
Moritz, A. et al. Akt-RSK-S6 kinase signaling networks activated by oncogenic receptor tyrosine kinases. Sci. Signal. 3, ra64 (2010).
Nath, A., Li, I., Roberts, L. R. & Chan, C. Elevated free fatty acid uptake via CD36 promotes epithelial-mesenchymal transition in hepatocellular carcinoma. Sci. Rep. 5, 14752 (2015).
Nicholson, R. I., Gee, J. M. & Harper, M. E. EGFR and cancer prognosis. Eur. J. Cancer 37(Suppl 4), S9–S15 (2001).
Nokin M. J. et al. Methylglyoxal, a glycolysis side-product, induces Hsp90 glycation and YAP-mediated tumor growth and metastasis. eLife 5, 19375 (2016).
Oksvold, M. P. et al. Serine mutations that abrogate ligand-induced ubiquitination and internalization of the EGF receptor do not affect c-Cbl association with the receptor. Oncogene 22, 8509–8518 (2003).
Shimamoto, Y. et al. Sensitivity of human cancer cells to the new anticancer ribo-nucleoside TAS-106 is correlated with expression of uridine-cytidine kinase 2. Jpn. J. Cancer Res.: Gann 93, 825–833 (2002).
Singal, A. G., Pillai, A. & Tiro, J. Early detection, curative treatment, and survival rates for hepatocellular carcinoma surveillance in patients with cirrhosis: a meta-analysis. PLoS Med. 11, e1001624 (2014).
Suzuki, N. N., Koizumi, K., Fukushima, M., Matsuda, A. & Inagaki, F. Structural basis for the specificity, catalysis, and regulation of human uridine-cytidine kinase. Structure 12, 751–764 (2004).
Swinnen, J. V., Brusselmans, K. & Verhoeven, G. Increased lipogenesis in cancer cells: new players, novel targets. Curr. Opin. Clin. Nutr. Metab. Care 9, 358–365 (2006).
Takehana, T. et al. Expression of epidermal growth factor receptor in gastric carcinomas. Clin. Gastroenterol. Hepatol. 1, 438–445 (2003).
Tomoike, F., Nakagawa, N., Kuramitsu, S. & Masui, R. Structural and Biochemical Studies on the Reaction Mechanism of Uridine-Cytidine Kinase. Protein J. 34, 411–420 (2015).
Tong, J., Taylor, P. & Moran, M. F. Proteomic analysis of the epidermal growth factor receptor (EGFR) interactome and post-translational modifications associated with receptor endocytosis in response to EGF and stress. Mol. Cell. Proteom. 13, 1644–1658 (2014).
Tong, X., Zhao, F. & Thompson, C. B. The molecular determinants of de novo nucleotide biosynthesis in cancer cells. Curr. Opin. Genet. Dev. 19, 32–37 (2009).
Tsao, A. et al. Phase II study of TAS-106 in patients with platinum-failure recurrent or metastatic head and neck cancer and nasopharyngeal cancer. Cancer Med. 2, 351–359 (2013).
van Kuilenburg, A. B. & Meinsma, R. The pivotal role of uridine-cytidine kinases in pyrimidine metabolism and activation of cytotoxic nucleoside analogues in neuroblastoma. Biochimica et. Biophysica Acta 1862, 1504–1512 (2016).
Van Rompay, A. R., Norda, A., Linden, K., Johansson, M. & Karlsson, A. Phosphorylation of uridine and cytidine nucleoside analogs by two human uridine-cytidine kinases. Mol. Pharmacol. 59, 1181–1186 (2001).
Ward, P. S. & Thompson, C. B. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21, 297–308 (2012).
Yu, S. et al. UCK2 upregulation might serve as an indicator of unfavorable prognosis of hepatocellular carcinoma. IUBMB Life 71, 105–112 (2019).
Zhao, W. et al. A New Bliss Independence Model to Analyze Drug Combination Data. J. Biomol. Screen. 19, 817–821 (2014).
Zhao, Y. N. et al. [Expression and significance of EGF mRNA and EGFR mRNA in hepatocellular carcinoma]. Ai zheng = Aizheng = Chin. J. Cancer 23, 762–766 (2004).
Acknowledgements
Thanks to Prof. Liwei Dong for his support in construction of PDX models. This work was supported by the National Key Sci-Tech Special Project of China (2018ZX10723204-006-004) to X.K. The National Natural Science Foundation of China (81873582 and 81670562 to X.K., 81670598 to Q.X., 31870905 and 31671453 to H.W.), and Shanghai Municipal Education Commission—Gaofeng Clinical Medicine Grant Support (20171911) to X.K., the National Key Research and Development Program of China (2017YFC0908100) to Q.X., the Scientific Program of Shanghai Municipal Health Commission (201940352) to H.W.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cai, J., Sun, X., Guo, H. et al. Non-metabolic role of UCK2 links EGFR-AKT pathway activation to metastasis enhancement in hepatocellular carcinoma. Oncogenesis 9, 103 (2020). https://doi.org/10.1038/s41389-020-00287-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-020-00287-7
- Springer Nature Limited
This article is cited by
-
UCK2 promotes intrahepatic cholangiocarcinoma progression and desensitizes cisplatin treatment by PI3K/AKT/mTOR/autophagic axis
Cell Death Discovery (2024)
-
Immune- and metabolism-related gene signature analysis uncovers the prognostic and immune microenvironments of hepatocellular carcinoma
Journal of Cancer Research and Clinical Oncology (2024)
-
M7G methylated core genes (METTL1 and WDR4) and associated RNA risk signatures are associated with prognosis and immune escape in HCC
BMC Medical Genomics (2023)
-
Targeting uridine–cytidine kinase 2 induced cell cycle arrest through dual mechanism and could improve the immune response of hepatocellular carcinoma
Cellular & Molecular Biology Letters (2022)
-
UBE2T-mediated Akt ubiquitination and Akt/β-catenin activation promotes hepatocellular carcinoma development by increasing pyrimidine metabolism
Cell Death & Disease (2022)