Abstract
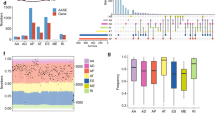
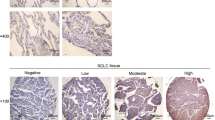
Novel therapeutic strategies for non-small-cell lung cancer (NSCLC) are urgently needed. RNA splicing, orchestrated by the spliceosome, is deregulated in many forms of cancer, including NSCLC. Here, we performed high-throughput screening with a small interfering RNA library targeting all annotated human spliceosome proteins to identify cancer-selective lethal targets in the RNA splicing machinery. Silencing of several spliceosome proteins reduced cell viability in a panel of NSCLC cell lines, but not in non-malignant lung fibroblasts and epithelial cells. Interestingly, the cancer-selective lethal target set comprised all seven Sm proteins that, together with small nuclear RNA, form the core structure of most spliceosome subunits. Interfering with Sm protein expression induced apoptosis in NSCLC cells, but not in non-malignant cells. In silico analysis revealed that Sm proteins are frequently upregulated in NSCLC. For several Sm proteins, increased expression showed a positive correlation with disease severity. Together, our results suggest that the Sm proteins represent particularly useful novel targets for selective treatment of NSCLC.





Similar content being viewed by others
References
Reck M, Heigener DF, Mok T, Soria JC, Rabe KF. Management of non-small-cell lung cancer: recent developments. Lancet. 2013;382:709–19.
Oltean S, Bates DO. Hallmarks of alternative splicing in cancer. Oncogene. 2014;33:5311–8.
David CJ, Manley JL. Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev. 2010;24:2343–64.
Wahl MC, Will CL, Lührmann R. The spliceosome: design principles of a dynamic RNP machine. Cell. 2009;136:701–18.
Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, et al. Alternative isoform regulation in human tissue transcriptomes. Nature. 2008;456:470–6.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E, et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell. 2012;150:1107–20.
Bonnal S, Vigevani L, Valcárcel J. The spliceosome as a target of novel antitumour drugs. Nat Rev Drug Discov. 2012;11:847–59.
Hong DS, Kurzrock R, Naing A, Wheler JJ, Falchook GS, Schiffman JS, et al. A phase I, open-label, single-arm, dose-escalation study of E7107, a precursor messenger RNA (pre-mRNA) spliceosome inhibitor administered intravenously on days 1 and 8 every 21 days to patients with solid tumors. Invest New Drugs. 2014;32:436–44.
Siebring-van Olst E, Blijlevens M, de Menezes RX, van der Meulen-Muileman IH, Smit EF, van Beusechem VW. A genome-wide siRNA screen for regulators of tumor suppressor p53 activity in human non-small cell lung cancer cells identifies components of the RNA splicing machinery as targets for anticancer treatment. Mol Oncol. 2017;11:534–51.
Pao W, Girard N. New driver mutations in non-small-cell lung cancer. Lancet Oncol. 2011;12:175–80.
Forbes SA, Beare D, Boutselakis H, Bamford S, Bindal N, Tate J, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017;45:D777–D783.
Matera AG, Wang Z. A day in the life of the spliceosome. Nat Rev Mol Cell Biol. 2014;15:108–21.
Campbell JD, Alexandrov A, Kim J, Wala J, Berger WH, Sekhar C, et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet.2016;48:607–16.
Quidville V, Alsafadi S, Goubar A, Commo F, Scott V, Pioche-Durieu C, et al. Targeting the deregulated spliceosome core machinery in cancer cells triggers mTOR blockade and autophagy. Cancer Res. 2013;73:2247–58.
Langer W, Sohler F, Leder G, Beckmann G, Seidel H, Gröne J, et al. Exon array analysis using re-defined probe sets results in reliable identification of alternatively spliced genes in non-small cell lung cancer. BMC Genom. 2010;11:676.
Gao Y, Koide K. Chemical perturbation of Mcl-1 pre-mRNA splicing to induce apoptosis in cancer cells. ACS Chem Biol. 2013;8:895–900.
Citterio L, Azzani T, Duga S, Bianchi G. Genomic organization of the human gamma adducin gene. Biochem Biophys Res Commun. 1999;266:110–4.
Dutertre M, Lacroix-Triki M, Driouch K, de la Grange P, Gratadou L, Beck S, et al. Exon-based clustering of murine breast tumor transcriptomes reveals alternative exons whose expression is associated with metastasis. Cancer Res. 2010;70:896–905.
Effenberger KA, Urabe VK, Jurica MS. Modulating splicing with small molecular inhibitors of the spliceosome. Wiley Interdiscip Rev RNA. 2017;8:1–19.
Webb TR, Joyner AS, Potter PM. The development and application of small molecule modulators of SF3b as therapeutic agents for cancer. Drug Discov Today. 2012;18:43–49.
Eskens FALM, Ramos FJ, Burger H, O’Brien JP, Piera A, de Jonge MJA, et al. Phase I pharmacokinetic and pharmacodynamic study of the first-in-class spliceosome inhibitor E7107 in patients with advanced solid tumors. Clin Cancer Res. 2013;19:6296–304.
Pomeranz Krummel DA, Oubridge C, Leung AKW, Li J, Nagai K. Crystal structure of human spliceosomal U1 snRNP at 5.5 resolution. Nature. 2009;458:475–80.
Weber G, Trowitzsch S, Kastner B, Lührmann R, Wahl MC. Functional organization of the Sm core in the crystal structure of human U1 snRNP. EMBO J. 2010;29:4172–84.
Kambach C, Walke S, Young R, Avis JM, de la Fortelle E, Raker VA, et al. Crystal structures of two Sm protein complexes and their implications for the assembly of the spliceosomal snRNPs. Cell. 1999;3:375–87.
Kanehisa M, Susumu G. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000;28:27–30.
Carbon S, et al. AmiGO: Online access to ontology and annotation data. Bioinformatics. 2009;25:288–9.
Croft D, et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 2011;39:691–7.
R Core Team (2014). R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. http://www.R-project.org/.
Bachas C, Hodzic J, van der Mijn JC, Stoepker C, Verheul HWM, Wolthuis RMF, et al. Rscreenorm: normalization of CRISPR and siRNA screen data for more reproducible hit selection. BMC Bioinforma. 2018;19:1–12.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B. 1995;57:289–300.
Benjamini Y, Yekutieli D. The control of the false discovery rate in multiple testing under dependency. Ann Stat. 2001;29:1165–88.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal complementary data sources and analysis options. Sci Signal. 2013;6:1–20.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Arman B, et al. The cBio Cancer Genomics Portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Author contributions
Study design was performed by V.W.v.B., experiments were performed by M.B. and I.H.v.d.M.-M., data analysis was performed by M.B. and R.X.d.M., manuscript was written with input from all the authors by M.B. and V.W.v.B., V.W.v.B. and E.F.S. supervised the project.
Funding
This work was supported by Walter Bruckerhoff Stiftung and Stichting VUmc-CCA.
Code availability
Scripts can be accessed via CodeOcean.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Blijlevens, M., van der Meulen-Muileman, I.H., de Menezes, R.X. et al. High-throughput RNAi screening reveals cancer-selective lethal targets in the RNA spliceosome. Oncogene 38, 4142–4153 (2019). https://doi.org/10.1038/s41388-019-0711-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0711-z
- Springer Nature Limited