Abstract
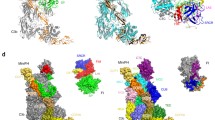
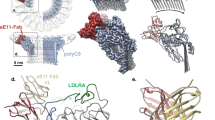
Complement factor H (FH) attenuates C3b molecules tethered by their thioester domains to self surfaces and thereby protects host tissues. Factor H is a cofactor for initial C3b proteolysis that ultimately yields a surface-attached fragment (C3d) corresponding to the thioester domain. We used NMR and X-ray crystallography to study the C3d–FH19–20 complex in atomic detail and identify glycosaminoglycan-binding residues in factor H module 20 of the C3d–FH19–20 complex. Mutagenesis justified the merging of the C3d–FH19–20 structure with an existing C3b–FH1–4 crystal structure. We concatenated the merged structure with the available FH6–8 crystal structure and new SAXS-derived FH1–4, FH8–15 and FH15–19 envelopes. The combined data are consistent with a bent-back factor H molecule that binds through its termini to two sites on one C3b molecule and simultaneously to adjacent polyanionic host-surface markers.






Similar content being viewed by others
References
Ricklin, D., Hajishengallis, G., Yang, K. & Lambris, J.D. Complement: a key system for immune surveillance and homeostasis. Nat. Immunol. 11, 785–797 (2010).
de Córdoba, S.R. & de Jorge, E.G. Translational mini-review series on complement factor H: genetics and disease associations of human complement factor H. Clin. Exp. Immunol. 151, 1–13 (2008).
Janssen, B.J., Christodoulidou, A., McCarthy, A., Lambris, J.D. & Gros, P. Structure of C3b reveals conformational changes that underlie complement activity. Nature 444, 213–216 (2006).
Pangburn, M.K., Schreiber, R.D. & Muller-Eberhard, H.J. Human complement C3b inactivator: isolation, characterization, and demonstration of an absolute requirement for the serum protein beta1H for cleavage of C3b and C4b in solution. J. Exp. Med. 146, 257–270 (1977).
Weiler, J.M., Daha, M.R., Austen, K.F. & Fearon, D.T. Control of the amplification convertase of complement by the plasma protein beta1H. Proc. Natl. Acad. Sci. USA 73, 3268–3272 (1976).
Whaley, K. & Ruddy, S. Modulation of the alternative complement pathways by beta 1 H globulin. J. Exp. Med. 144, 1147–1163 (1976).
Ripoche, J. et al. Partial characterization of human complement factor H by protein and cDNA sequencing: homology with other complement and non-complement proteins. Biosci. Rep. 6, 65–72 (1986).
Meri, S. & Pangburn, M.K. Regulation of alternative pathway complement activation by glycosaminoglycans: specificity of the polyanion binding site on factor H. Biochem. Biophys. Res. Commun. 198, 52–59 (1994).
Pangburn, M.K. Host recognition and target differentiation by factor H, a regulator of the alternative pathway of complement. Immunopharmacology 49, 149–157 (2000).
Gordon, D.L., Kaufman, R.M., Blackmore, T.K., Kwong, J. & Lublin, D.M. Identification of complement regulatory domains in human factor H. J. Immunol. 155, 348–356 (1995).
Pangburn, M.K. Cutting edge: localization of the host recognition functions of complement factor H at the carboxyl-terminal: implications for hemolytic uremic syndrome. J. Immunol. 169, 4702–4706 (2002).
Schmidt, C.Q. et al. A new map of glycosaminoglycan and C3b binding sites on factor H. J. Immunol. 181, 2610–2619 (2008).
Sharma, A.K. & Pangburn, M.K. Identification of three physically and functionally distinct binding sites for C3b in human complement factor H by deletion mutagenesis. Proc. Natl. Acad. Sci. USA 93, 10996–11001 (1996).
Wu, J. et al. Structure of complement fragment C3b-factor H and implications for host protection by complement regulators. Nat. Immunol. 10, 728–733 (2009).
Buddles, M.R., Donne, R.L., Richards, A., Goodship, J. & Goodship, T.H. Complement factor H gene mutation associated with autosomal recessive atypical hemolytic uremic syndrome. Am. J. Hum. Genet. 66, 1721–1722 (2000).
Herbert, A.P., Uhrin, D., Lyon, M., Pangburn, M.K. & Barlow, P.N. Disease-associated sequence variations congregate in a polyanion recognition patch on human factor H revealed in three-dimensional structure. J. Biol. Chem. 281, 16512–16520 (2006).
Pérez-Caballero, D. et al. Clustering of missense mutations in the C-terminal region of factor H in atypical hemolytic uremic syndrome. Am. J. Hum. Genet. 68, 478–484 (2001).
Ferreira, V.P., Herbert, A.P., Hocking, H.G., Barlow, P.N. & Pangburn, M.K. Critical role of the C-terminal domains of factor H in regulating complement activation at cell surfaces. J. Immunol. 177, 6308–6316 (2006).
Jokiranta, T.S. et al. Binding of complement factor H to endothelial cells is mediated by the carboxy-terminal glycosaminoglycan binding site. Am. J. Pathol. 167, 1173–1181 (2005).
Blackmore, T.K. et al. Identification of the second heparin-binding domain in human complement factor H. J. Immunol. 160, 3342–3348 (1998).
Blackmore, T.K., Sadlon, T.A., Ward, H.M., Lublin, D.M. & Gordon, D.L. Identification of a heparin binding domain in the seventh short consensus repeat of complement factor H. J. Immunol. 157, 5422–5427 (1996).
Hageman, G.S. et al. A common haplotype in the complement regulatory gene factor H (HF1/CFH) predisposes individuals to age-related macular degeneration. Proc. Natl. Acad. Sci. USA 102, 7227–7232 (2005).
Klein, R.J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Kuehn, B.M. Gene discovery provides clues to cause of age-related macular degeneration. J. Am. Med. Assoc. 293, 1841–1845 (2005).
Zareparsi, S. et al. Strong association of the Y402H variant in complement factor H at 1q32 with susceptibility to age-related macular degeneration. Am. J. Hum. Genet. 77, 149–153 (2005).
Wiesmann, C. et al. Structure of C3b in complex with CRIg gives insights into regulation of complement activation. Nature 444, 217–220 (2006).
Lian, L.Y., Barsukov, I.L., Sutcliffe, M.J., Sze, K.H. & Roberts, G.C. Protein-ligand interactions: exchange processes and determination of ligand conformation and protein-ligand contacts. Methods Enzymol. 239, 657–700 (1994).
Matsuo, H. et al. Identification by NMR spectroscopy of residues at contact surfaces in large, slowly exchanging macromolecular complexes. J. Am. Chem. Soc. 121, 9903–9904 (1999).
Daughdrill, G.W. et al. Chemical shift changes provide evidence for overlapping single-stranded DNA- and XPA-binding sites on the 70 kDa subunit of human replication protein A. Nucleic Acids Res. 31, 4176–4183 (2003).
McAlister, M.S.B. et al. NMR analysis of interacting soluble forms of the cell-cell recognition molecules CD2 and CD48. Biochemistry 35, 5982–5991 (1996).
Ferreira, V.P. et al. The binding of factor H to a complex of physiological polyanions and C3b on cells is impaired in atypical hemolytic uremic syndrome. J. Immunol. 182, 7009–7018 (2009).
Lehtinen, M.J., Rops, A.L., Isenman, D.E., van der Vlag, J. & Jokiranta, T.S. Mutations of factor H impair regulation of surface-bound C3b by three mechanisms in atypical hemolytic uremic syndrome. J. Biol. Chem. 284, 15650–15658 (2009).
Morikis, D. & Lambris, J.D. The electrostatic nature of C3d-complement receptor 2 association. J. Immunol. 172, 7537–7547 (2004).
Ricklin, D., Ricklin-Lichtsteiner, S.K., Markiewski, M.M., Geisbrecht, B.V. & Lambris, J.D. Cutting edge: members of the Staphylococcus aureus extracellular fibrinogen-binding protein family inhibit the interaction of C3d with complement receptor 2. J. Immunol. 181, 7463–7467 (2008).
Bhattacharjee, A., Lehtinen, M.J., Kajander, T., Goldman, A. & Jokiranta, T.S. Both domain 19 and domain 20 of factor H are involved in binding to complement C3b and C3d. Mol. Immunol. 47, 1686–1691 (2010).
Prosser, B.E. et al. Structural basis for complement factor H linked age-related macular degeneration. J. Exp. Med. 204, 2277–2283 (2007).
Schmidt, C.Q. et al. The central portion of factor H (modules 10–15) is compact and contains a structurally deviant CCP module. J. Mol. Biol. 395, 105–122 (2010).
Barlow, P.N. et al. Solution structure of a pair of complement modules by nuclear magnetic resonance. J. Mol. Biol. 232, 268–284 (1993).
Barlow, P.N. et al. Solution structure of the fifth repeat of factor H: a second example of the complement control protein module. Biochemistry 31, 3626–3634 (1992).
Szakonyi, G. et al. Structure of complement receptor 2 in complex with its C3d ligand. Science 292, 1725–1728 (2001).
Hammel, M. et al. A structural basis for complement inhibition by Staphylococcus aureus. Nat. Immunol. 8, 430–437 (2007).
Chen, H. et al. Allosteric inhibition of complement function by a staphylococcal immune evasion protein. Proc. Natl. Acad. Sci. USA 107, 17621–17626 (2010).
Jokiranta, T.S. et al. Structure of complement factor H carboxyl-terminus reveals molecular basis of atypical haemolytic uremic syndrome. EMBO J. 25, 1784–1794 (2006).
Saunders, R.E. et al. The interactive Factor H-atypical hemolytic uremic syndrome mutation database and website: update and integration of membrane cofactor protein and Factor I mutations with structural models. Hum. Mutat. 28, 222–234 (2007).
Miller, E.C., Roumenina, L., Fremeaux-Bacchi, V. & Atkinson, J.P. Functional characterization of mutations in complement C3 that predispose to aHUS. Mol. Immunol. 47, 2291 (2010).
Frémeaux-Bacchi, V. et al. Mutations in complement C3 predispose to development of atypical hemolytic uremic syndrome. Blood 112, 4948–4952 (2008).
Aslam, M. & Perkins, S.J. Folded-back solution structure of monomeric factor H of human complement by synchrotron X-ray and neutron scattering, analytical ultracentrifugation and constrained molecular modelling. J. Mol. Biol. 309, 1117–1138 (2001).
Fernando, A.N. et al. Associative and structural properties of the region of complement factor H encompassing the Tyr402His disease-related polymorphism and its interactions with heparin. J. Mol. Biol. 368, 564–581 (2007).
Oppermann, M. et al. The C-terminus of complement regulator Factor H mediates target recognition: evidence for a compact conformation of the native protein. Clin. Exp. Immunol. 144, 342–352 (2006).
Okemefuna, A.I. et al. The regulatory SCR-1/5 and cell surface-binding SCR-16/20 fragments of factor H reveal partially folded-back solution structures and different self-associative properties. J. Mol. Biol. 375, 80–101 (2008).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Young, K.A., Chen, X.S., Holers, V.M. & Hannan, J.P. Isolating the Epstein-Barr virus gp350/220 binding site on complement receptor type 2 (CR2/CD21). J. Biol. Chem. 282, 36614–36625 (2007).
Shaw, C.D. et al. Delineation of the complement receptor type 2–C3d complex by site-directed mutagenesis and molecular docking. J. Mol. Biol. 404, 697–710 (2010).
Leslie, A.G.W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 ESF-EAMCB Newslett. Protein Crystallogr. 26, (1992).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Nagar, B., Jones, R.G., Diefenbach, R.J., Isenman, D.E. & Rini, J.M. X-ray crystal structure of C3d: a C3 fragment and ligand for complement receptor 2. Science 280, 1277–1281 (1998).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Acknowledgements
Use of the Protein Production Facilities at the University of Edinburgh was supported by The Wellcome Trust, the Scottish University Life Sciences Alliance and the UK Biotechnology and Biological Sciences Research Council. C.Q.S. and M.G. were supported by Wellcome Trust grant 081179, A.P.H. was supported by Wellcome Trust grant 078780/Z/05/Z (to D.U. and P.N.B.), B.S.B. was supported by an EastChem PhD studentship and a University of Edinburgh campaign small project grant and C.M.J. was supported by EC FP6 (Marie Curie EST Fellowship, contract MEST-CT-2005-020744).
Author information
Authors and Affiliations
Contributions
M.G., C.Q.S., D.K. and A.P.H. engineered and produced factor H proteins; J.P.H. and D.G. engineered and produced C3d proteins; C.M.J. generated dp8 heparin fragments. J.P.H. and D.G. crystallized the C3d–FH complex; H.P.M. and J.P.H. collected crystallographic data; H.P.M. determined and refined the structure; B.S.B. produced 15N-labeled factor H and carried out NMR studies of the FH–C3d and FH–dp8 interactions; B.S.B. and A.P.H. carried out the hemolysis assay; J.P.H. carried out the C3d-dp8 competition ELISA; C.Q.S. and M.G. carried out the SPR measurements; H.D.T.M. and D.I.S. carried out the SAXS analysis; D.U., J.P.H. and P.N.B. conceived and supervised the project; H.P.M., C.Q.S., M.G., B.S.B., D.U., J.P.H. and P.N.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–3 and Supplementary Methods (PDF 2975 kb)
Rights and permissions
About this article
Cite this article
Morgan, H., Schmidt, C., Guariento, M. et al. Structural basis for engagement by complement factor H of C3b on a self surface. Nat Struct Mol Biol 18, 463–470 (2011). https://doi.org/10.1038/nsmb.2018
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2018
- Springer Nature America, Inc.
This article is cited by
-
Gliosis-dependent expression of complement factor H truncated variants attenuates retinal neurodegeneration following ischemic injury
Journal of Neuroinflammation (2024)
-
Invariant surface glycoprotein 65 of Trypanosoma brucei is a complement C3 receptor
Nature Communications (2022)
-
The influences of α-hemolytic Streptococcus on class switching and complement activation of human tonsillar cells in IgA nephropathy
Immunologic Research (2022)
-
A receptor for the complement regulator factor H increases transmission of trypanosomes to tsetse flies
Nature Communications (2020)
-
Immunological features and functional analysis of anti-CFH autoantibodies in patients with atypical hemolytic uremic syndrome
Pediatric Nephrology (2019)