Abstract
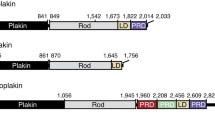
Desmosomes are intercellular junctions in which cadherin cell adhesion molecules are linked to the intermediate filament (IF) system. Desmoplakin is a member of the plakin family of IF-binding proteins. The C-terminal domain of desmoplakin (DPCT) mediates binding to IFs in desmosomes. The DPCT sequence contains three regions, termed A, B and C, consisting of 4.5 copies of a 38-amino acid repeat motif. We demonstrate that these regions form discrete subdomains that bind to IFs and report the crystal structures of domains B and C. In contrast to the elongated structures formed by other kinds of repeat motifs, the plakin repeats form a globular structure with a unique fold. A conserved basic groove found on the domain may represent an IF-binding site.







Similar content being viewed by others
References
Kowalczyk, A.P., Bornslaeger, E.A., Norvell, S.M., Palka, H.L. & Green, K.J. Desmosomes: intercellular adhesive junctions specialized for attachment of intermediate filaments. Int. Rev. Cytol. 185, 237–302 (1999).
Koch, P.J. et al. Identification of desmoglein, a constitutive desmosomal glycoprotein, as a member of the cadherin family of cell adhesion molecules. Eur. J. Cell Biol. 53, 1–12 (1990).
Mechanic, S., Raynor, K., Hill, J.E. & Cowin, P. Desmocollins form a distinct subset of the cadherin family of cell adhesion molecules. Proc. Natl. Acad. Sci. USA 88, 4476–4480 (1991).
Witcher, L.L. et al. Desmosomal cadherin binding domains of plakoglobin. J. Biol. Chem. 271, 10904–10909 (1996).
Hatzfeld, M., Kristjansson, G.I., Plessmann, U. & Weber, K. Band 6 protein, a major constituent of desmosomes from stratified epithelia, is a novel member of the armadillo multigene family. J. Cell Sci. 107, 2259–2270 (1994).
Heid, H.W. et al. Cell type-specific desmosomal plaque proteins of the plakoglobin family: plakophilin 1 (band 6 protein). Differentiation 58, 113–131 (1994).
Wahl, J.D. et al. Plakoglobin domains that define its association with the desmosomal cadherins and the classical cadherins: identification of unique and shared domains. J. Cell Sci. 109, 1143–1154 (1996).
Leung, C.L., Green, K.J. & Liem, R.K.H. Plakins: a family of versatile cytolinker proteins. Trends Cell Biol. 12, 37–45 (2002).
Green, K.J. et al. Structure of the human desmoplakins. Implications for function in the desmosomal plaque. J. Biol. Chem. 265, 2603–2612 (1990).
Kowalczyk, A.P. et al. The amino-terminal domain of desmoplakin binds to plakoglobin and clusters desmosomal cadherin-plakoglobin complexes. J. Cell Biol. 139, 773–784 (1997).
Stappenbeck, T.S. & Green, K.G. The desmoplakin carboxyl terminus coaligns with and specifically disrupts intermediate filament networks when expressed in cultured cells. J. Cell Biol. 116, 1197–1209 (1992).
Kouklis, P.D., Hutton, E. & Fuchs, E. Making a connection: direct binding between keratin intermediate filaments and desmosomal proteins. J. Cell. Biol. 127, 1049–1060 (1994).
Norgett, E.E. et al. Recessive mutation in desmoplakin disrupts desmoplakin-intermediate filament interactions and causes dilated cardiomyopathy, woolly hair and keratoderma. Hum. Mol. Genet. 9, 2761–2766 (2000).
Armstrong, D.K. et al. Haploinsufficiency of desmoplakin causes a striate subtype of palmoplantar keratoderma. Hum. Mol. Genet. 8, 143–148 (1999).
Gallicano, G.I. et al. Desmoplakin is required early in development for assembly of desmosomes and cytoskeletal linkage. J. Cell Biol. 143, 2009–2022 (1998).
Vasioukhin, V., Bowers, E., Bauer, C., Degenstein, L. & Fuchs, E. Desmoplakin is essential in epidermal sheet formation. Nat. Cell Biol. 3, 1076–1085 (2001).
Meng, J.-J., Bornslaeger, E.A., Green, K.J., Steinert, P.M. & Ip, W. Two-hybrid analysis reveals fundamental differences in direct interactions between desmoplakin and cell type-specific intermediate filaments. J. Biol. Chem. 272, 21495–21503 (1997).
Nikolic, B., Nulty, E.M., Mir, B. & Wiche, G. Basic amino acid residue cluster within nuclear targeting sequence motif is essential for cytoplasmic plectin-vimentin network junctions. J. Cell Biol. 134, 1455–1467 (1996).
DiColandrea, T., Karashima, T., Määttä, A. & Watt, F.M. Subcellular distribution of envoplakin and periplakin: insights into their role as precursors of the epidermal cornified envelope. J. Cell Biol. 151, 573–585 (2000).
Gorina, S. & Pavletich, N.P. Structure of the p53 tumor suppressor bound to the ankyrin and SH3 domains of 53BP2. Science 274, 1001–1005 (1996).
Luh, F.Y. et al. Structure of the cyclin-dependent kinase inhibitor p19Ink4d. Nature 389, 999–1003 (1997).
Huber, A.H., Nelson, W.J. & Weis, W.I. Three-dimensional structure of the armadillo repeat region of β-catenin. Cell 90, 871–882 (1997).
Groves, M.R., Hanlon, N., Turowski, P., Hemmings, B.A. & Barford, D. The structure of the protein phosphatase 2A PR65/A subunit reveals the conformation of its 15 tandemly repeated HEAT motifs. Cell 96, 99–110 (1999).
Wall, M.A. et al. The structure of the G protein heterotrimer Giα1β1γ2 . Cell 83, 1047–1058 (1995).
Kobe, B. & Deisenhofer, J. Crystal structure of porcine ribonuclease inhibitor, a protein with leucine-rich repeats. Nature 366, 751–756 (1993).
Yoder, M.D., Keen, N.T. & Jurnak, F. New domain motif: the structure of pectate lyase C, asecreted plant virulence factor. Science 260, 1503–1507 (1993).
Janda, L., Damborsky, J., Rezniczek, G.A. & Wiche, G. Plectin repeats and modules: strategic cysteines and their presumed impact on cytolinker functions. Bioessays 23, 1064–1069 (2001).
Strelkov, S.V. et al. Divide-and-conquer crystallographic approach towards an atomic structure of intermediate filaments. J. Mol. Biol. 306, 773–781 (2001).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Leslie, A.G. Integration of macromolecular diffraction data. Acta Crystallogr. D 55, 1696–1702 (1999).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Terwilliger, T.C. & Berendzen, J. Automated structure solution for MIR and MAD. Acta Crystallogr. D 55, 849–861 (1999).
Brünger, A.T. et al. Crystallography and NMR System (CNS): a new software system for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for the building of protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. 15, 133–138 (1997).
Nicholls, A. GRASP: Graphical Representation and Analysis of Surface Properties (Columbia University, New York; 1992).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
We thank P. Ellis for beamline support. Portions of this research were carried out at the Stanford Synchrotron Radiation Laboratory (SSRL), a national user facility operated by Stanford University on behalf of the U.S. Department of Energy, Office of Basic Energy Sciences. The SSRL Structural Molecular Biology Program is supported by the Department of Energy, Office of Biological and Environmental Research, and by the National Institutes of Health, National Center for Research Resources, Biomedical Technology Program, and the National Institute of General Medical Sciences. H.-J.C. was supported by fellowships from the Korea Science and Engineering Foundation and the American Heart Association. This work was supported by grants to W.I.W. and K.J.G from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Choi, HJ., Park-Snyder, S., Pascoe, L. et al. Structures of two intermediate filament-binding fragments of desmoplakin reveal a unique repeat motif structure. Nat Struct Mol Biol 9, 612–620 (2002). https://doi.org/10.1038/nsb818
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb818
- Springer Nature America, Inc.