Key Points
-
The interplay between transcription factors and epigenetic regulators is crucial for regulating gene-expression programmes during haematopoiesis. Epigenetic regulation — including post-translational modification of histones and DNA methylation — is also linked with upstream signalling pathways and external signals that shape the identity and function of immune cells.
-
Distinctive DNA methylation changes characterize the differentiation of myeloid cells and lymphoid cells from their progenitors. DNA demethylation is more predominant during myeloid differentiation than during lymphoid differentiation.
-
The methylcytosine hydroxylase TET2, which oxidizes 5-methylcytosine, has a major role in the acquisition of myeloid cell identity. This has been demonstrated using transdifferentiation models and is highlighted by the association of TET2 mutations with myeloid malignancies.
-
Epigenetic control has a key role in defining macrophage polarization, connecting external stimuli with the establishment of specific transcriptional programmes.
-
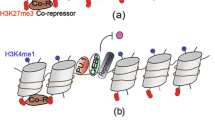
Epigenetic modifications have a key role in the generation of memory-type behaviour in innate immune cells. Following an initial stimulus, the persistence of the trimethylated histone H3K4 at latent enhancers ensures the increased expression of pro-inflammatory genes after restimulation.
Abstract
Myeloid cells are crucial effectors of the innate immune response and important regulators of adaptive immunity. The differentiation and activation of myeloid cells requires the timely regulation of gene expression; this depends on the interplay of a variety of elements, including transcription factors and epigenetic mechanisms. Epigenetic control involves histone modifications and DNA methylation, and is coupled to lineage-specifying transcription factors, upstream signalling pathways and external factors released in the bone marrow, blood and tissue environments. In this Review, we highlight key epigenetic events controlling myeloid cell biology, focusing on those related to myeloid cell differentiation, the acquisition of myeloid identity and innate immune memory.



Similar content being viewed by others
References
De Kleer, I., Willems, F., Lambrecht, B. & Goriely, S. Ontogeny of myeloid cells. Front. Immunol. 5, 423 (2014).
Geissmann, F. et al. Development of monocytes, macrophages, and dendritic cells. Science 327, 656–661 (2010).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Fischle, W., Wang, Y. & Allis, C. D. Histone and chromatin cross-talk. Curr. Opin. Cell Biol. 15, 172–183 (2003).
Turner, B. M. Histone acetylation and an epigenetic code. Bioessays 22, 836–845 (2000).
Hendrich, B. & Bird, A. Identification and characterization of a family of mammalian methyl-CpG binding proteins. Mol. Cell. Biol. 18, 6538–6547 (1998).
Blattler, A. & Farnham, P. J. Cross-talk between site-specific transcription factors and DNA methylation states. J. Biol. Chem. 288, 34287–34294 (2013).
Bird, A., Taggart, M., Frommer, M., Miller, O. J. & Macleod, D. A fraction of the mouse genome that is derived from islands of nonmethylated, CpG-rich DNA. Cell 40, 91–99 (1985).
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Irizarry, R. A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nature Genet. 41, 178–186 (2009).
Maunakea, A. K. et al. Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature 466, 253–257 (2010).
Zilbauer, M. et al. Genome-wide methylation analyses of primary human leukocyte subsets identifies functionally important cell-type-specific hypomethylated regions. Blood 122, e52–e60 (2013).
Schlesinger, F., Smith, A. D., Gingeras, T. R., Hannon, G. J. & Hodges, E. De novo DNA demethylation and noncoding transcription define active intergenic regulatory elements. Genome Res. 23, 1601–1614 (2013).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Goll, M. G. & Bestor, T. H. Eukaryotic cytosine methyltransferases. Annu. Rev. Biochem. 74, 481–514 (2005).
Piccolo, F. M. & Fisher, A. G. Getting rid of DNA methylation. Trends Cell Biol. 24, 136–143 (2014).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009). This article demonstrates that TET1 oxidizes 5meC to 5hmC in vivo , indicating its potential role in epigenetic regulation.
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Dawlaty, M. M. et al. Loss of tet enzymes compromises proper differentiation of embryonic stem cells. Dev. Cell 29, 102–111 (2014).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nature Genet. 46, 17–23 (2014).
Laiosa, C. V., Stadtfeld, M. & Graf, T. Determinants of lymphoid-myeloid lineage diversification. Annu. Rev. Immunol. 24, 705–738 (2006).
Wilting, R. H. et al. Overlapping functions of Hdac1 and Hdac2 in cell cycle regulation and haematopoiesis. EMBO J. 29, 2586–2597 (2010).
Broske, A. M. et al. DNA methylation protects hematopoietic stem cell multipotency from myeloerythroid restriction. Nature Genet. 41, 1207–1215 (2009).
Chen, L. et al. Transcriptional diversity during lineage commitment of human blood progenitors. Science 345, 1251033 (2014).
Kondo, M., Weissman, I. L. & Akashi, K. Identification of clonogenic common lymphoid progenitors in mouse bone marrow. Cell 91, 661–672 (1997).
Wada, T. et al. Expression levels of histone deacetylases determine the cell fate of hematopoietic progenitors. J. Biol. Chem. 284, 30673–30683 (2009).
Trowbridge, J. J., Snow, J. W., Kim, J. & Orkin, S. H. DNA methyltransferase 1 is essential for and uniquely regulates hematopoietic stem and progenitor cells. Cell Stem Cell 5, 442–449 (2009).
Challen, G. A. et al. Dnmt3a and Dnmt3b have overlapping and distinct functions in hematopoietic stem cells. Cell Stem Cell 15, 350–364 (2014).
Sun, D. et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 14, 673–688 (2014).
Boehm, T. Evolution of vertebrate immunity. Curr. Biol. 22, R722–R732 (2012).
Hodges, E. et al. Directional DNA methylation changes and complex intermediate states accompany lineage specificity in the adult hematopoietic compartment. Mol. Cell 44, 17–28 (2011).
Mansson, R. et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity 26, 407–419 (2007).
Ji, H. et al. Comprehensive methylome map of lineage commitment from haematopoietic progenitors. Nature 467, 338–342 (2010). This article represents the first genome-wide analysis of DNA methylation during differentiation, identifying gene-specific hypomethylation in the myeloid lineage and an increase in methylation in the lymphoid lineage.
Mossadegh-Keller, N. et al. M-CSF instructs myeloid lineage fate in single haematopoietic stem cells. Nature 497, 239–243 (2013).
Bocker, M. T. et al. Genome-wide promoter DNA methylation dynamics of human hematopoietic progenitor cells during differentiation and aging. Blood 117, e182–e189 (2011).
Accomando, W. P., Wiencke, J. K., Houseman, E. A., Nelson, H. H. & Kelsey, K. T. Quantitative reconstruction of leukocyte subsets using DNA methylation. Genome Biol. 15, R50 (2014).
Ronnerblad, M. et al. Analysis of the DNA methylome and transcriptome in granulopoiesis reveal timed changes and dynamic enhancer methylation. Blood 123, e79–e89 (2014).
Klug, M. et al. Active DNA demethylation in human postmitotic cells correlates with activating histone modifications, but not transcription levels. Genome Biol. 11, R63 (2010).
Klug, M., Schmidhofer, S., Gebhard, C., Andreesen, R. & Rehli, M. 5-Hydroxymethylcytosine is an essential intermediate of active DNA demethylation processes in primary human monocytes. Genome Biol. 14, R46 (2013).
Zhang, X. et al. DNA methylation dynamics during ex vivo differentiation and maturation of human dendritic cells. Epigenet. Chromatin 7, 21 (2014).
Scott, E. W., Simon, M. C., Anastasi, J. & Singh, H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science 265, 1573–1577 (1994).
Laslo, P. et al. Multilineage transcriptional priming and determination of alternate hematopoietic cell fates. Cell 126, 755–766 (2006).
Feng, R. et al. PU.1 and C/EBPα/β convert fibroblasts into macrophage-like cells. Proc. Natl Acad. Sci. USA 105, 6057–6062 (2008).
Xie, H., Ye, M., Feng, R. & Graf, T. Stepwise reprogramming of B cells into macrophages. Cell 117, 663–676 (2004).
Bussmann, L. H. et al. A robust and highly efficient immune cell reprogramming system. Cell Stem Cell 5, 554–566 (2009).
Barneda-Zahonero, B. et al. HDAC7 is a repressor of myeloid genes whose downregulation is required for transdifferentiation of pre-B cells into macrophages. PLoS Genet. 9, e1003503 (2013).
Kallin, E. M. et al. Tet2 facilitates the derepression of myeloid target genes during CEBPα-induced transdifferentiation of pre-B cells. Mol. Cell 48, 266–276 (2012). This article demonstrates a direct role for TET2 in the 5hmC-mediated activation of myeloid-specific genes during the acquisition of myeloid identity.
Hashimoto, H. et al. Recognition and potential mechanisms for replication and erasure of cytosine hydroxymethylation. Nucleic Acids Res. 40, 4841–4849 (2012).
Hanna, J. et al. Direct reprogramming of terminally differentiated mature B lymphocytes to pluripotency. Cell 133, 250–264 (2008).
Di Stefano, B. et al. C/EBPα poises B cells for rapid reprogramming into induced pluripotent stem cells. Nature 506, 235–239 (2014).
Delhommeau, F. et al. Mutation in TET2 in myeloid cancers. N. Engl. J. Med. 360, 2289–2301 (2009).
Li, Z. et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118, 4509–4518 (2011).
Pronier, E. et al. Inhibition of TET2-mediated conversion of 5-methylcytosine to 5-hydroxymethylcytosine disturbs erythroid and granulomonocytic differentiation of human hematopoietic progenitors. Blood 118, 2551–2555 (2011).
Losman, J. A. et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339, 1621–1625 (2013).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011).
Sasaki, M. et al. IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature 488, 656–659 (2012).
Ko, M. et al. Ten-eleven-translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl Acad. Sci. USA 108, 14566–14571 (2011).
de la Rica, L. et al. PU.1 target genes undergo Tet2-coupled demethylation and DNMT3b-mediated methylation in monocyte-to-osteoclast differentiation. Genome Biol. 14, R99 (2014).
Hervouet, E., Vallette, F. M. & Cartron, P. F. Dnmt3/transcription factor interactions as crucial players in targeted DNA methylation. Epigenetics 4, 487–499 (2009).
Lawrence, T. & Natoli, G. Transcriptional regulation of macrophage polarization: enabling diversity with identity. Nature Rev. Immunol. 11, 750–761 (2011).
Mullican, S. E. et al. Histone deacetylase 3 is an epigenomic brake in macrophage alternative activation. Genes Dev. 25, 2480–2488 (2011).
Chen, X. et al. Requirement for the histone deacetylase Hdac3 for the inflammatory gene expression program in macrophages. Proc. Natl Acad. Sci. USA 109, E2865–E2874 (2012).
Ishii, M. et al. Epigenetic regulation of the alternatively activated macrophage phenotype. Blood 114, 3244–3254 (2009).
Satoh, T. et al. The Jmjd3–Irf4 axis regulates M2 macrophage polarization and host responses against helminth infection. Nature Immunol. 11, 936–944 (2010).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007).
Barish, G. D. et al. Bcl-6 and NF-κB cistromes mediate opposing regulation of the innate immune response. Genes Dev. 24, 2760–2765 (2010).
Adelman, K. et al. Immediate mediators of the inflammatory response are poised for gene activation through RNA polymerase II stalling. Proc. Natl Acad. Sci. USA 106, 18207–18212 (2009).
Ghisletti, S. et al. Identification and characterization of enhancers controlling the inflammatory gene expression program in macrophages. Immunity 32, 317–328 (2010).
Kaikkonen, M. U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Austenaa, L. et al. The histone methyltransferase Wbp7 controls macrophage function through GPI glycolipid anchor synthesis. Immunity 36, 572–585 (2012).
Lam, M. T. et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 498, 511–515 (2013).
Medzhitov, R., Schneider, D. S. & Soares, M. P. Disease tolerance as a defense strategy. Science 335, 936–941 (2012).
Medzhitov, R. Origin and physiological roles of inflammation. Nature 454, 428–435 (2008).
Netea, M. G., Quintin, J. & van der Meer, J. W. Trained immunity: a memory for innate host defense. Cell Host Microbe 9, 355–361 (2011).
Quintin, J., Cheng, S. C., van der Meer, J. W. & Netea, M. G. Innate immune memory: towards a better understanding of host defense mechanisms. Curr. Opin. Immunol. 29C, 1–7 (2014).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013). This article identifies latent enhancers as key elements of innate immune memory; these are characterized by being marked with H3K4me3 following an initial stimulus.
Quintin, J. et al. Candida albicans infection affords protection against reinfection via functional reprogramming of monocytes. Cell Host Microbe 12, 223–232 (2012). This article describes the H3K4me3 mark as a mediator of trained immunity in monocytes.
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Cheng, S. C. et al. mTOR- and HIF-1α-mediated aerobic glycolysis as metabolic basis for trained immunity. Science 345, 1250684 (2014).
Adib-Conquy, M. & Cavaillon, J. M. Compensatory anti-inflammatory response syndrome. Thromb. Haemost. 101, 36–47 (2009).
Foster, S. L., Hargreaves, D. C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007). This study characterizes the two types of TLR-induced responses and provides an epigenetic basis for the acquisition of endotoxin tolerance.
Yang, X. et al. Epigenetic regulation of macrophage polarization by DNA methyltransferase 3b. Mol. Endocrinol. 28, 565–574 (2014).
Sica, A. & Mantovani, A. Macrophage plasticity and polarization: in vivo veritas. J. Clin. Invest. 122, 787–795 (2012).
Shih, A. H., Abdel-Wahab, O., Patel, J. P. & Levine, R. L. The role of mutations in epigenetic regulators in myeloid malignancies. Nature Rev. Cancer 12, 599–612 (2012).
Quintas-Cardama, A., Santos, F. P. & Garcia-Manero, G. Therapy with azanucleosides for myelodysplastic syndromes. Nature Rev. Clin. Oncol. 7, 433–444 (2010).
Patel, J. P. et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N. Engl. J. Med. 366, 1079–1089 (2012).
Xia, M. et al. Histone methyltransferase Ash1l suppresses interleukin-6 production and inflammatory autoimmune diseases by inducing the ubiquitin-editing enzyme A20. Immunity 39, 470–481 (2013).
Moore, K. J., Sheedy, F. J. & Fisher, E. A. Macrophages in atherosclerosis: a dynamic balance. Nature Rev. Immunol. 13, 709–721 (2013).
Hoeksema, M. A. et al. Targeting macrophage histone deacetylase 3 stabilizes atherosclerotic lesions. EMBO Mol. Med. 6, 1124–1132 (2014).
Cao, Q. et al. Histone deacetylase 9 represses cholesterol efflux and alternatively activated macrophages in atherosclerosis development. Arterioscler Thromb. Vasc. Biol. 34, 1871–1879 (2014).
Kruidenier, L. et al. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488, 404–408 (2012).
Rodríguez-Ubreva, J. et al. Pre-B cell to macrophage transdifferentiation without significant promoter DNA methylation changes. Nucleic Acids Res. 40, 1954–1968 (2012).
Acknowledgements
This work was supported by grant SAF2011-29635 from the Spanish Ministry of Science and Innovation, grant CIVP16A1834 from the Fundación Ramón Areces and grant Precisesads 115565–3 of the Innovative Medicines Initiative (IMI) Programme.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Histones
-
A family of basic proteins found in the nuclei of eukaryotic cells that package and organize DNA into repetitive structural units named nucleosomes.
- Shores
-
Regions that are located within 2 kb of CpG islands and are usually methylation hotspots.
- Shelves
-
Regions that flank CpG island shores and are located 2–4 kb from CpG islands.
- Open sea
-
CpG sites that are located outside of the CpG island context.
- Imprinted genes
-
Genes for which the expression status is determined by the parent that contributed them.
- Methyl-CpG binding proteins
-
A family of proteins characterized by the presence of the methyl-CpG binding domain, several of which are integral components of or are involved in the recruitment of different histone deacetylase complexes and chromatin remodelling factors of different nuclear complexes that have a role in gene expression.
- Yamanaka factors
-
A set of transcription factors that are highly expressed in embryonic stem cells. Their overexpression induces pluripotency in somatic cells.
- 'M1' or 'classical' activation
-
Lipopolysaccharide- or interferon-γ-mediated stimulation of macrophages leading to a pro-inflammatory phenotype.
- 'M2' or 'alternative' activation
-
Interleukin-4-dependent stimulation of macrophages that leads to an anti-inflammatory phenotype.
- Enhancer RNAs
-
(eRNAs). A class of relatively short non-coding RNA molecules (50–2000 nucleotides in length) that are transcribed from the DNA sequence of enhancer regions.
Rights and permissions
About this article
Cite this article
Álvarez-Errico, D., Vento-Tormo, R., Sieweke, M. et al. Epigenetic control of myeloid cell differentiation, identity and function. Nat Rev Immunol 15, 7–17 (2015). https://doi.org/10.1038/nri3777
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3777
- Springer Nature Limited
This article is cited by
-
Correlation of DNA methylation of DNMT3A and TET2 with oral squamous cell carcinoma
Discover Oncology (2024)
-
Targeting the myeloid microenvironment in neuroblastoma
Journal of Experimental & Clinical Cancer Research (2023)
-
Investigating the effects of maltreatment and acute stress on the concordance of blood and DNA methylation methods of estimating immune cell proportions
Clinical Epigenetics (2023)
-
DNA Methylation is Involved in Sex Determination in Spinach
Biochemical Genetics (2023)
-
PCBP1 is associated with rheumatoid arthritis by affecting RNA products of genes involved in immune response in Th1 cells
Scientific Reports (2022)