Abstract
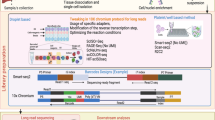
Here, we describe a method of systematic PCR screening with multiround sample pooling for the isolation of rare PCR-positive samples. As an example, we have applied this protocol to the recovery of gene-targeted clones in human somatic cells comprising only 0.02–0.17% of cells transduced with targeting vectors. Initially, cells infected with targeting vectors are seeded and grown in fourteen 96-well tissue culture plates. Samples are then collected from these plates and subjected to two rounds of pooling to yield twelve 'superpools' used for an initial PCR. After identifying PCR-positive samples, de-pooling is carried out with successive rounds of PCR screening, using samples of decreasing complexity. Single-cell cloning is subsequently performed to isolate gene-targeted clones. The entire protocol can be completed in 4–8 weeks depending on the proliferative capacity of the cell line.







Similar content being viewed by others
References
Busch, M.P. et al. Evaluation of screened blood donations for human immunodeficiency virus type 1 infection by culture and DNA amplification of pooled cells. N. Engl. J. Med. 325, 1–5 (1991).
Roth, W.K., Weber, M. & Seifried, E. Feasibility and efficacy of routine PCR screening of blood donations for hepatitis C virus, hepatitis B virus, and HIV-1 in a blood-bank setting. Lancet 353, 359–363 (1999).
Quinn, T.C. et al. Feasibility of pooling sera for HIV-1 viral RNA to diagnose acute primary HIV-1 infection and estimate HIV incidence. AIDS 14, 2751–2757 (2000).
Bergen, A., Wang, C.Y., Nakhai, B. & Goldman, D. Mass allele detection (MAD) of rare 5-HT1A structural variants with allele-specific amplification and electrochemiluminescent detection. Hum. Mutat. 7, 135–143 (1996).
Till, B.J. et al. High-throughput discovery of rare human nucleotide polymorphisms by Ecotilling. Nucleic Acids Res. 34, e99 (2006).
Heard, E., Davies, B., Feo, S. & Fried, M. An improved method for the screening of YAC libraries. Nucleic Acids Res. 17, 5861 (1989).
Green, E.D. & Olson, M.V. Systematic screening of yeast artificial-chromosome libraries by use of the polymerase chain reaction. Proc. Natl. Acad. Sci. USA 87, 1213–1217 (1990).
Cai, L. et al. Construction and characterization of a bovine bacterial artificial chromosome library. Genomics 29, 413–425 (1995).
Kim, U.J. et al. Construction and characterization of a human bacterial artificial chromosome library. Genomics 34, 213–218 (1996).
McAlinden, T.P. & Krawetz, S.A. A practical method to screen libraries of cloned DNA. Anal. Biochem. 218, 237–238 (1994).
Bloem, L.J. & Yu, L. A time-saving method for screening cDNA or genomic libraries. Nucleic Acids Res. 18, 2830 (1990).
Israel, D.I. A PCR-based method for high stringency screening of DNA libraries. Nucleic Acids Res. 21, 2627–2631 (1993).
McCallum, C.M., Comai, L., Greene, E.A. & Henikoff, S. Targeted screening for induced mutations. Nat. Biotechnol. 18, 455–457 (2000).
Stepanova, A.N. & Alonso, J.M. PCR-based screening for insertional mutants. Methods Mol. Biol. 323, 163–172 (2006).
Lesa, G.M. Isolation of Caenorhabditis elegans gene knockouts by PCR screening of chemically mutagenized libraries. Nat. Protoc. 1, 2231–2240 (2006).
Brown, A.C., Lerner, C.P., Graber, J.H., Shafferand, D.J. & Roopenian, D.C. Pooling and PCR as a method to combat low frequency gene targeting in mouse embryonic stem cells. Cytotechnology 51, 81–88 (2006).
Campbell, T.N. & Choy, F.Y. Approaches to library screening. J. Mol. Microbiol. Biotechnol. 4, 551–554 (2002).
Kim, H.S. & Smithies, O. Recombinant fragment assay for gene targeting based on the polymerase chain reaction. Nucleic Acids Res. 16, 8887–8903 (1988).
Konishi, H. et al. Knock-in of mutant K-ras in non-tumorigenic human epithelial cells as a new model for studying K-ras mediated transformation. Cancer Res. 67, 8460–8467 (2007).
Eppig, J.T. & Strivens, M. Finding a mouse: the International Mouse Strain Resource (IMSR). Trends Genet. 15, 81–82 (1999).
Buerstedde, J.M. & Takeda, S. Increased ratio of targeted to random integration after transfection of chicken B cell lines. Cell 67, 179–188 (1991).
Shirasawa, S., Furuse, M., Yokoyama, N. & Sasazuki, T. Altered growth of human colon cancer cell lines disrupted at activated Ki-ras. Science 260, 85–88 (1993).
Park, B.H., Vogelstein, B. & Kinzler, K.W. Genetic disruption of PPARδ decreases the tumorigenicity of human colon cancer cells. Proc. Natl. Acad. Sci. USA 98, 2598–2603 (2001).
Rhee, I. et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416, 552–556 (2002).
Cummins, J.M. et al. Tumour suppression: disruption of HAUSP gene stabilizes p53. Nature 428, 487 (2004).
Dang, D.T. et al. Hypoxia-inducible factor-1α promotes nonhypoxia-mediated proliferation in colon cancer cells and xenografts. Cancer Res. 66, 1684–1936 (2006).
Hiyama, T. et al. Haploinsufficiency of the Mus81-Eme1 endonuclease activates the intra-S-phase and G2/M checkpoints and promotes rereplication in human cells. Nucleic Acids Res. 34, 880–892 (2006).
Hyndman, D.L. & Mitsuhashi, M. PCR primer design. Methods Mol. Biol. 226, 81–88 (2003).
Kohli, M., Rago, C., Lengauer, C., Kinzler, K.W. & Vogelstein, B. Facile methods for generating human somatic cell gene knockouts using recombinant adeno-associated viruses. Nucleic Acids Res. 32, e3 (2004).
Acknowledgements
This work was supported by The Flight Attendant Medical Research Institute (FAMRI), NIH/NCI CA109274, The Elsa Pardee Foundation, the Summer Running Fund and the Avon Foundation. H.K. is a recipient of a Young Clinical Scientist Award from FAMRI and was also supported by the YASUDA Medical Research Foundation and the Kanzawa Medical Research Foundation. J.L. receives support from the American Society of Clinical Oncology's Young Investigator Award and is a recipient of a Young Clinical Scientist Award from FAMRI. J.P. Garay is a recipient of a Research Supplement to Promote Diversity in Health-Related Research, A.M.A. is supported by an NIH Institutional Training Grant T32CA67751 and J.P. Gustin is a recipient of a Department of Defense Breast Cancer Research Program Predoctoral Fellowship Award W81XWH-06-1-0325.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Video 1
Demonstration of Sample pooling round 1 (generation of 84 Pools) and round 2 (generation of 12 Superpools) procedures (MPG 19402 kb)
Rights and permissions
About this article
Cite this article
Konishi, H., Lauring, J., Garay, J. et al. A PCR-based high-throughput screen with multiround sample pooling: application to somatic cell gene targeting. Nat Protoc 2, 2865–2874 (2007). https://doi.org/10.1038/nprot.2007.409
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.409
- Springer Nature Limited
This article is cited by
-
PIK3CA mutations and TP53 alterations cooperate to increase cancerous phenotypes and tumor heterogeneity
Breast Cancer Research and Treatment (2017)
-
Multiscale Cues Drive Collective Cell Migration
Scientific Reports (2016)
-
Engineering targeted chromosomal amplifications in human breast epithelial cells
Breast Cancer Research and Treatment (2015)
-
Knock in of the AKT1 E17K mutation in human breast epithelial cells does not recapitulate oncogenic PIK3CA mutations
Oncogene (2010)