Abstract
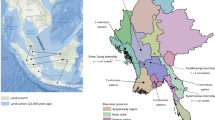
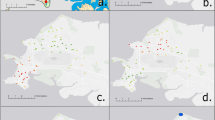
The environmental bacterium Burkholderia pseudomallei causes an estimated 165,000 cases of human melioidosis per year worldwide and is also classified as a biothreat agent. We used whole genome sequences of 469 B. pseudomallei isolates from 30 countries collected over 79 years to explore its geographic transmission. Our data point to Australia as an early reservoir, with transmission to Southeast Asia followed by onward transmission to South Asia and East Asia. Repeated reintroductions were observed within the Malay Peninsula and between countries bordered by the Mekong River. Our data support an African origin of the Central and South American isolates with introduction of B. pseudomallei into the Americas between 1650 and 1850, providing a temporal link with the slave trade. We also identified geographically distinct genes/variants in Australasian or Southeast Asian isolates alone, with virulence-associated genes being among those over-represented. This provides a potential explanation for clinical manifestations of melioidosis that are geographically restricted.



Similar content being viewed by others
References
Limmathurotsakul, D. et al. Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat. Microbiol. 1, 15008 (2016).
Nandi, T. et al. Burkholderia pseudomallei sequencing identifies genomic clades with distinct recombination, accessory, and epigenetic profiles. Genome Res. 25, 129–141 (2015).
Johnson, S. L. et al. Whole-genome sequences of 80 environmental and clinical isolates of Burkholderia pseudomallei. Genome Announc. 3, e01282-14 (2015).
Holden, M. T. G. et al. Genomic plasticity of the causative agent of melioidosis, Burkholderia pseudomallei. Proc. Natl Acad. Sci. USA 101, 14240–14245 (2004).
Price, E. P. et al. Unprecedented melioidosis cases in Northern Australia caused by an Asian Burkholderia pseudomallei strain identified by using large-scale comparative genomics. Appl. Environ. Microbiol. 82, 954–963 (2015).
Pearson, T. et al. Phylogeographic reconstruction of a bacterial species with high levels of lateral gene transfer. BMC Biol. 7, 78 (2009).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Dale, J. et al. Epidemiological tracking and population assignment of the non-clonal bacterium, Burkholderia pseudomallei. PLoS Negl. Trop. Dis. 5, e1381 (2011).
Gee, J. E., Allender, C. J., Tuanyok, A., Elrod, M. G. & Hoffmaster, A. R. Burkholderia pseudomallei type G in western hemisphere. Emerg. Infect. Dis. 20, 682–684 (2014).
Sarovich, D. S. et al. Phylogenomic analysis reveals an Asian origin for African Burkholderia pseudomallei and further supports melioidosis endemicity in Africa. mSphere 1, e00089-15 (2016).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Kolchin, P. American Slavery: 1619–1877 (Penguin, 1995).
Thomas, H. The Slave Trade, The Story of the Atlantic Slave Trade: 1440–1870 (Simon & Schuster, 1997).
Nguyen, T. D. The Mekong River and the Struggle for Indochina: Water, War and Peace (Praeger, 1999).
Liu, J. H., Lawrence, B., Ward, C. & Abraham, S. Social representations of history in Malaysia and Singapore: on the relationship between national and ethnic identity. Asian J. Soc. Psychol. 5, 3–20 (2002).
Currie, B. J. Melioidosis: evolving concepts in epidemiology, pathogenesis, and treatment. Semin. Respir. Crit. Care Med. 36, 111–125 (2015).
Lees, J. A. et al. Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes. Nat. Commun. 7, 12797 (2016).
Sarovich, D. S. et al. Variable virulence factors in Burkholderia pseudomallei (melioidosis) associated with human disease. PLoS ONE 9, e91682 (2014).
Tuanyok, A. et al. A horizontal gene transfer event defines two distinct groups within Burkholderia pseudomallei that have dissimilar geographic distributions. J. Bacteriol. 189, 9044–9049 (2007).
French, C. T. et al. Dissection of the Burkholderia intracellular life cycle using a photothermal nanoblade. Proc. Natl Acad. Sci. USA 108, 12095–12100 (2011).
Benanti, E. L., Nguyen, C. M. & Welch, M. D. Virulent Burkholderia species mimic host actin polymerases to drive actin-based motility. Cell 161, 348–360 (2015).
Tuanyok, A. et al. Genomic islands from five strains of Burkholderia pseudomallei. BMC Genomics 9, 566 (2008).
Willcocks, S. J., Denman, C. C., Atkins, H. S. & Wren, B. W. Intracellular replication of the well-armed pathogen Burkholderia pseudomallei. Curr. Opin. Microbiol. 29, 94–103 (2016).
Chen, Y. et al. Characterization and analysis of the Burkholderia pseudomallei BsaN virulence regulon. BMC Microbiol. 14, 206 (2014).
Bast, A. et al. Caspase-1-dependent and -independent cell death pathways in Burkholderia pseudomallei infection of macrophages. PLoS Pathogens 10, e1003986 (2014).
Wallace, A. R. On the physical geography of the Malay archipelago. J. R. Geogr. Soc. Lond. 7, 205–212 (1863).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Chewapreecha, C. et al. Dense genomic sampling identifies highways of pneumococcal recombination. Nat. Genet. 46, 305–309 (2014).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Nandi, T. et al. A genomic survey of positive selection in Burkholderia pseudomallei provides insights into the evolution of accidental virulence. PLoS Pathogens 6, e1000845 (2010).
Spring-Pearson, S. M. et al. Pangenome analysis of Burkholderia pseudomallei: genome evolution preserves gene order despite high recombination rates. PLoS ONE 10, e0140274 (2015).
Price, M. N., Dehal, P. S. & Arkin, A. P. Fasttree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Harris, S. R. et al. Evolution of MRSA during hospital transmission and intercontinental spread. Science 327, 469–474 (2010).
Page, A. J. et al. SNP-sites: rapid efficient extraction of SNPs from multiFASTA alignments. Microb. Genomics http://dx.doi.org/10.1099/mgen.0.000056 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and postanalysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Corander, J., Marttinen, P., Sirén, J. & Tang, J. Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinformatics 9, 539 (2008).
Cheng, L., Connor, T. R., Siren, J., Aanensen, D. M. & Corander, J. Hierarchical and spatially explicit clustering of DNA sequences with BAPS software. Mol. Biol. Evol. 30, 1224–1228 (2013).
Croucher, N. J. et al. Population genomics of post-vaccine changes in pneumococcal epidemiology. Nat. Genet. 45, 656–663 (2013).
Assefa, S., Keane, T. M., Otto, T. D., Newbold, C. & Berriman, M. ABACAS: algorithm-based automatic contiguation of assembled sequences. Bioinformatics 25, 1968–1969 (2009).
Carver, T. J. et al. ACT: the artemis comparison tool. Bioinformatics 21, 3422–3423 (2005).
Harris, S. R. et al. Genome specialization and decay of the strangles pathogen, Streptococcus equi, is driven by persistent infection. Genome Res. 25, 1360–1371 (2015).
Rambaut, A., Suchard, M. A., Xie, D. & Drummond, A. J. Tracer v.1.6 (2014), http://beast.bio.ed.ac.uk/Tracer
Ho, S. Y. W. & Duchene, S. Molecular-clock methods for estimating evolutionary rates and timescales. Mol. Ecol. 23, 5947–5965 (2014).
Murray, G. G. R. et al. The effect of genetic structure on molecular dating and tests for temporal signal. Methods Ecol. Evol. 7, 80–89 (2016).
Lieberman, T. D. et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nat. Genet. 43, 1275–1280 (2011).
Mathers, A. J. et al. Klebsiella pneumoniae carbapenemase (KPC)-producing K. pneumoniae at a single institution: insights into endemicity from whole-genome sequencing. Antimicrob. Agents Chemother. 59, 1656–1663 (2015).
Young, B. C. et al. Evolutionary dynamics of Staphylococcus aureus during progression from carriage to disease. Proc. Natl Acad. Sci. USA 109, 4550–4555 (2012).
Croucher, N. J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Revell, L. J. Phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Bollback, J. P. SIMMAP: stochastic character mapping of discrete traits on phylogenies. BMC Bioinformatics 7, 88 (2006).
Laabei, M. et al. Predicting the virulence of MRSA from its genome sequence. Genome Res. 24, 839–849 (2014).
Chewapreecha, C. et al. Comprehensive identification of single nucleotide polymorphisms associated with β-lactam resistance within pneumococcal mosaic genes. PLoS Genet. 10, e1004547 (2014).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Ooi, W. F. et al. The condition-dependent transcriptional landscape of Burkholderia pseudomallei. PLoS Genet. 9, e1003795 (2013).
Winsor, G. L. et al. The Burkholderia genome database: facilitating flexible queries and comparative analyses. Bioinformatics 24, 2803–2804 (2008).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2008); http://www.R-project.org
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Acknowledgements
The authors acknowledge the Wellcome Trust Sanger Institute library construction, sequence and core informatics teams and E. Blane for their technical support. The authors thank T. Nandi and P. Tan at Genome Institute of Singapore, and E. Price and D. Sarovich at Menzies School of Health Research, Australia, for providing access to publically available WGS data. The authors thank the following people who provided isolates or DNA: N. Day, MORU, Faculty of Tropical Medicine, Mahidol University; P. Newton, M. Vongsouvath, M. Mayway, V. Davong, O. Lattana, C. Moore, S. Rattanavong and the directors and staff of Mahosot Hospital, Vientiane, Lao PDR; V. Kumar, Ankor Hospital for Children, Siem Reap, Cambodia; J. Campbell, Oxford University Clinical Research Unit, Ho Chi Minh City, Vietnam; H. Suk Wai, Ocean Park Corporation, Hong Kong SAR, China; C. Kham and T. Phe, Sihanouk Hospital Centre of Hope, Phnom Penh, Cambodia; J.W. Wiersinga, Academic Medical Center (AMC), Amsterdam, the Netherlands; J. Jacobs, ITM, Antwerp, Belgium; J.E. Russell, National Collection of Type Cultures, UK; T. Pitt, NHS Blood and Transplant, UK; D. Godoy, Imperial College, UK; S. Emonet, Geneva University Hospitals, Switzerland; S. Morpeth, Middlemore Hospital, New Zealand and J. Gee, CDC, USA. C.C. is a Sir Henry Wellcome post-doctoral Fellow (grant no. 107376/Z/15/Z). J.C., M.V. and Z.Y. were supported by the COIN Centre of Excellence and Z.Y. through an HIIT post-doctoral fellowship. A.E.M. is supported by a Biotechnology and Biological Sciences Research Council grant (no. BB/M014088/1). B.G.S. was supported by Wellcome Trust grant no. WT089472. D.A.B.D. and R.P. are supported by Wellcome Trust grants 106698/Z/14 and B9R00760. D.L. and V.W. are supported by Wellcome Trust grant 089275/Z/09/Z. M.M. and B.J.C. are supported by the Australian National Health and Medical Research Council through project grants 1046812 and 1098337. This publication presents independent research supported by the Health Innovation Challenge Fund (WT098600, HICF-T5-342), a parallel funding partnership between the Department of Health and the Wellcome Trust. The views expressed in this publication are those of the author(s) and not necessarily those of the Department of Health or the Wellcome Trust. This project was also funded by a grant awarded to the Wellcome Trust Sanger Institute (no. 098051).
Author information
Authors and Affiliations
Contributions
A.T., B.D.S., S.L.H., C.B., M.M., V.W., D.L., R.P., B.G.S., P.K., D.A.B.D. and B.J.C. collected and provided the samples for the study. C.C. designed and performed the analyses. M.T.G.H., S.R.H., A.E.M., J.C., J.P. and G.D. designed and contributed materials and analysis tools. M.V., N.V., Z.Y., and J.C. performed the kmer-based analyses in the first draft. C.C. performed the kmer-based analysis in the revised draft. Z.Y. and J.C. performed cluster analyses. S.J.P. was responsible for management of the study. S.J.P. and C.C. wrote the paper with input from all authors. All authors approved the manuscript prior to submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–12, Supplementary Note, Supplementary References. (PDF 13377 kb)
Supplementary Data 1
Epidemiological data. (XLSX 77 kb)
Supplementary Data 2
Frequency, association scores and direction of association for significant kmers identified in Australasian and Southeast Asian GWAS analyses. (TXT 8024 kb)
Supplementary Data 3
Raw kmer sequences. (TXT 8583 kb)
Supplementary Data 4
Coding sequences found in region-specific loci; summary data of all annotated coding sequences located in 468 Australasia- and 14 Southeast Asia-specific loci. (TXT 199 kb)
Supplementary Data 5
COG, GO and pathway terms with significant kmer enrichment. (XLSX 88 kb)
Rights and permissions
About this article
Cite this article
Chewapreecha, C., Holden, M., Vehkala, M. et al. Global and regional dissemination and evolution of Burkholderia pseudomallei. Nat Microbiol 2, 16263 (2017). https://doi.org/10.1038/nmicrobiol.2016.263
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.263
- Springer Nature Limited
This article is cited by
-
Burkholderia pseudomallei and melioidosis
Nature Reviews Microbiology (2024)
-
Multidrug resistance plasmids underlie clonal expansions and international spread of Salmonella enterica serotype 1,4,[5],12:i:- ST34 in Southeast Asia
Communications Biology (2023)
-
Genotyping of Burkholderia pseudomallei Isolated From Patients in South-Western Coastal Region of India
Current Microbiology (2022)
-
Human gut-derived B. longum subsp. longum strains protect against aging in a d-galactose-induced aging mouse model
Microbiome (2021)
-
A multi-country study using MALDI-TOF mass spectrometry for rapid identification of Burkholderia pseudomallei
BMC Microbiology (2021)