Abstract
Inferring demographic history is an important task in population genetics. Many existing inference methods are based on predefined simplified population models, which are more suitable for hypothesis testing than exploratory analysis. We developed a novel model-flexible method called stairway plot, which infers changes in population size over time using SNP frequency spectra. This method is applicable for whole-genome sequences of hundreds of individuals. Using extensive simulation, we demonstrate the usefulness of the method for inferring demographic history, especially recent changes in population size. We apply the method to the whole-genome sequence data of 9 populations from the 1000 Genomes Project and show a pattern of fluctuations in human populations from 10,000 to 200,000 years ago.



Similar content being viewed by others
Change history
14 August 2015
In the version of this article initially published, the authors neglected to acknowledge one of the funding sources for their study. The acknowledgements should have recognized support from Chinese NSF grant 91231120 in addition to the other funding sources listed. The error has been corrected in the HTML and PDF versions of the article.
27 August 2015
A Correction to this paper has been published: https://doi.org/10.1038/ng0915-1099a
References
Keinan, A. & Clark, A.G. Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336, 740–743 (2012).
Schaffner, S.F. et al. Calibrating a coalescent simulation of human genome sequence variation. Genome Res. 15, 1576–1583 (2005).
Gutenkunst, R.N., Hernandez, R.D., Williamson, S.H. & Bustamante, C.D. Inferring the joint demographic history of multiple populations from multidimensional SNP frequency data. PLoS Genet. 5, e1000695 (2009).
Kryukov, G.V., Shpunt, A., Stamatoyannopoulos, J.A. & Sunyaev, S.R. Power of deep, all-exon resequencing for discovery of human trait genes. Proc. Natl. Acad. Sci. USA 106, 3871–3876 (2009).
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl. Acad. Sci. USA 108, 11983–11988 (2011).
Pybus, O.G., Rambaut, A. & Harvey, P.H. An integrated framework for the inference of viral population history from reconstructed genealogies. Genetics 155, 1429–1437 (2000).
Strimmer, K. & Pybus, O.G. Exploring the demographic history of DNA sequences using the generalized skyline plot. Mol. Biol. Evol. 18, 2298–2305 (2001).
Drummond, A.J., Rambaut, A., Shapiro, B. & Pybus, O.G. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol. Biol. Evol. 22, 1185–1192 (2005).
Opgen-Rhein, R., Fahrmeir, L. & Strimmer, K. Inference of demographic history from genealogical trees using reversible jump Markov chain Monte Carlo. BMC Evol. Biol. 5, 6 (2005).
Minin, V.N., Bloomquist, E.W. & Suchard, M.A. Smooth skyride through a rough skyline: Bayesian coalescent-based inference of population dynamics. Mol. Biol. Evol. 25, 1459–1471 (2008).
Heled, J. & Drummond, A.J. Bayesian inference of population size history from multiple loci. BMC Evol. Biol. 8, 289 (2008).
Gill, M.S. et al. Improving Bayesian population dynamics inference: a coalescent-based model for multiple loci. Mol. Biol. Evol. 30, 713–724 (2013).
Ho, S.Y.W. & Shapiro, B. Skyline-plot methods for estimating demographic history from nucleotide sequences. Mol. Ecol. Resour. 11, 423–434 (2011).
Atkinson, Q.D., Gray, R.D. & Drummond, A.J. Bayesian coalescent inference of major human mitochondrial DNA haplogroup expansions in Africa. Proc. Biol. Soc. 276, 367–373 (2009).
Gignoux, C.R., Henn, B.M. & Mountain, J.L. Rapid, global demographic expansions after the origins of agriculture. Proc. Natl. Acad. Sci. USA 108, 6044–6049 (2011).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Sheehan, S., Harris, K. & Song, Y.S. Estimating variable effective population sizes from multiple genomes: a sequentially Markov conditional sampling distribution approach. Genetics 194, 647–662 (2013).
Liu, X., Fu, Y.-X., Maxwell, T.J. & Boerwinkle, E. Estimating population genetic parameters and comparing model goodness-of-fit using DNA sequences with error. Genome Res. 20, 101–109 (2010).
Nielsen, R. Estimation of population parameters and recombination rates from single nucleotide polymorphisms. Genetics 154, 931–942 (2000).
Hudson, R.R. Two-locus sampling distributions and their application. Genetics 159, 1805–1817 (2001).
Boitard, S., Schlötterer, C., Nolte, V., Pandey, R.V. & Futschik, A. Detecting selective sweeps from pooled next-generation sequencing samples. Mol. Biol. Evol. 29, 2177–2186 (2012).
Gayral, P. et al. Reference-free population genomics from next-generation transcriptome data and the vertebrate-invertebrate gap. PLoS Genet. 9, e1003457 (2013).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Palo, J.U., Ulmanen, I., Lukka, M., Ellonen, P. & Sajantila, A. Genetic markers and population history: Finland revisited. Eur. J. Hum. Genet. 17, 1336–1346 (2009).
Hudson, R.R. Generating samples under a Wright-Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Chen, G.K., Marjoram, P. & Wall, J.D. Fast and flexible simulation of DNA sequence data. Genome Res. 19, 136–142 (2009).
Kong, A. et al. Rate of de novo mutations and the importance of father's age to disease risk. Nature 488, 471–475 (2012).
Campbell, C.D. et al. Estimating the human mutation rate using autozygosity in a founder population. Nat. Genet. 44, 1277–1281 (2012).
Conrad, D.F. et al. Variation in genome-wide mutation rates within and between human families. Nat. Genet. 43, 712–714 (2011).
Scally, A. & Durbin, R. Revising the human mutation rate: implications for understanding human evolution. Nat. Rev. Genet. 13, 745–753 (2012).
Boyko, A.R. et al. Assessing the evolutionary impact of amino acid mutations in the human genome. PLoS Genet. 4, e1000083 (2008).
Pruitt, K.D., Tatusova, T., Brown, G.R. & Maglott, D.R. NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy. Nucleic Acids Res. 40, D130–D135 (2012).
Lachance, J. et al. Evolutionary history and adaptation from high-coverage whole-genome sequences of diverse African hunter-gatherers. Cell 150, 457–469 (2012).
Matsumura, S. & Forster, P. Generation time and effective population size in Polar Eskimos. Proc. Biol. Soc. 275, 1501–1508 (2008).
Langergraber, K.E. et al. Generation times in wild chimpanzees and gorillas suggest earlier divergence times in great ape and human evolution. Proc. Natl. Acad. Sci. USA 109, 15716–15721 (2012).
Awadalla, P. et al. Direct measure of the de novo mutation rate in autism and schizophrenia cohorts. Am. J. Hum. Genet. 87, 316–324 (2010).
Fenner, J.N. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am. J. Phys. Anthropol. 128, 415–423 (2005).
Michaelson, J.J. et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 151, 1431–1442 (2012).
Garrigan, D. & Hammer, M.F. Reconstructing human origins in the genomic era. Nat. Rev. Genet. 7, 669–680 (2006).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014).
Polanski, A. & Kimmel, M. New explicit expressions for relative frequencies of single-nucleotide polymorphisms with application to statistical inference on population growth. Genetics 165, 427–436 (2003).
Fu, Y.X. Statistical properties of segregating sites. Theor. Popul. Biol. 48, 172–197 (1995).
Pedersen, M.E.H. Tuning and Simplifying Heuristical Optimization. PhD thesis, Univ. Southampton (2010).
Storn, R. & Price, K. Differential evolution—a simple and efficient heuristic for global optimization over continuous spaces. J. Glob. Optim. 11, 341–359 (1997).
Davidon, W.C. Variable metric method for minimization. SIAM J. Optim. 1, 1–17 (1991).
Kong, A. et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 467, 1099–1103 (2010).
Acknowledgements
The authors acknowledge the Texas Advanced Computing Center (TACC; https://www.tacc.utexas.edu/) at the University of Texas at Austin for providing high-performance computing resources that have contributed to the research results reported within this paper. The authors acknowledge the 1000 Genomes Project Consortium for generating and providing the DNA sequence resources used in this study. The authors thank S. Barton for copyediting the manuscript. The authors thank S. Xu, Y. Guan, T.J. Maxwell, F. Yu and E. Boerwinkle for helpful discussions and encouragement. This study was supported by US National Institutes of Health grants 1U01HG005728 and 2U54HG003273 and by Chinese NSF grant 91231120.
Author information
Authors and Affiliations
Contributions
X.L. designed the study, developed the method, conducted the analyses and wrote the manuscript. Y.-X.F. provided critical advice on methodology development and result interpretation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
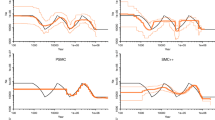
Supplementary Figure 1 Comparing the inferred histories of the stairway plot and the PSMC method using simulated samples based on five models.
(a) PSMC “sim-1” model. (b) PSMC “sim-2” model. (c) PSMC “sim-3” model. (d) PSMC “sim-YH” model. (e) Population 1 of the population split model. (f) Population 2 of the population split model. We assumed a mutation rate of 1.2 × 10–8 mutations per base pair per generation and a generation time of 24 years. Thin black lines, true models. Thick orange lines, medians of the inferred histories of the stairway plot; thin orange lines, 2.5 and 97.5 percentiles of the inferred histories of the stairway plot. Thick green lines, medians of the inferred histories of the PSMC method; thin green lines, 2.5 and 97.5 percentiles of the inferred histories of the PSMC method. n is number of simulated sequences, and L is the length of the simulated sequences. Dotted lines in e and f indicate the time point at which population 1 and population 2 split.
Supplementary Figure 2 Effects of sequence length, sample size and recombination rate on the inferences of the stairway plot.
We assumed a mutation rate of 1.2 × 10–8 mutations per base pair per generation and a generation time of 24 years. Sequences were simulated on the basis of the complex model I (see the Supplementary Note for details) assuming different sequence lengths (top row, a–c), sample sizes (middle row, a,d,e) or recombination rates (bottom row, a,f,g). Thin black lines, true models. Thick orange lines, medians of the inferred histories of the stairway plot; thin orange lines, 2.5 and 97.5 percentiles of the inferred histories of the stairway plot. N is the number of simulated sequences, L is the length of the simulated sequences, r is the recombination rate per site and t is the mutation rate per site.
Supplementary Figure 3 Simulation experiments on the unobserved bottlenecks of the 1000 Genomes Project populations.
(a) YRI model. (b) CEU-LWK hybrid model. (c) FIN-LWK hybrid model. (d) FIN-CEU-LWK hybrid model. Thin black lines, true models. Thick orange lines, medians of the inferred histories of the stairway plot; thin orange lines, 2.5 and 97.5 percentiles of the inferred histories of the stairway plot.
Supplementary Figure 4 Stairway plot estimations (medians) of the nine 1000 Genomes Project populations.
The bottlenecks after 200,000 years ago are likely artificial bottlenecks.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Note. (PDF 1448 kb)
Source data
Rights and permissions
About this article
Cite this article
Liu, X., Fu, YX. Exploring population size changes using SNP frequency spectra. Nat Genet 47, 555–559 (2015). https://doi.org/10.1038/ng.3254
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3254
- Springer Nature America, Inc.
This article is cited by
-
Which contributes more to the relict flora distribution pattern in East Asia, geographical processes or climate change? New evidence from the phylogeography of Rehderodendron kwangtungense
BMC Plant Biology (2024)
-
Past volcanic activity predisposes an endemic threatened seabird to negative anthropogenic impacts
Scientific Reports (2024)
-
Population genomics reveals how 5 ka of human occupancy led the Lima leaf-toed gecko (Phyllodactylus sentosus) to the brink of extinction
Scientific Reports (2023)
-
Domestication of the Amazonian fruit tree cupuaçu may have stretched over the past 8000 years
Communications Earth & Environment (2023)
-
Species-specific traits mediate avian demographic responses under past climate change
Nature Ecology & Evolution (2023)