Abstract
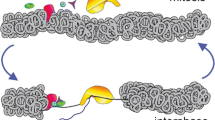
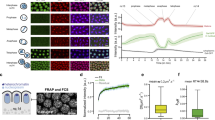
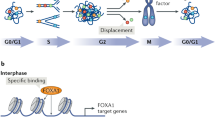
Although transmission of the gene expression program from mother to daughter cells has been suggested to be mediated by gene bookmarking, the precise mechanism by which bookmarking mediates post-mitotic transcriptional re-activation has been unclear. Here, we used a real-time gene expression system to quantitatively demonstrate that transcriptional activation of the same genetic locus occurs with a significantly more rapid kinetics in post-mitotic cells versus interphase cells. RNA polymerase II large subunit (Pol II) and bromodomain protein 4 (BRD4) were recruited to the locus in a different sequential order on interphase initiation versus post-mitotic re-activation resulting from the recognition by BRD4 of increased levels of histone H4 Lys 5 acetylation (H4K5ac) on the previously activated locus. BRD4 accelerated the dynamics of messenger RNA synthesis by de-compacting chromatin and hence facilitating transcriptional re-activation. Using a real-time quantitative approach, we identified differences in the kinetics of transcriptional activation between interphase and post-mitotic cells that are mediated by a chromatin-based epigenetic mechanism.






Similar content being viewed by others
References
Lemon, B. & Tjian, R. Orchestrated response: a symphony of transcription factors for gene control. Genes Dev. 14, 2551–2569 (2000).
Maniatis, T. & Reed, R. An extensive network of coupling among gene expression machines. Nature 416, 499–506 (2002).
Orphanides, G. & Reinberg, D. A unified theory of gene expression. Cell 108, 439–451 (2002).
McNally, J. G., Muller, W. G., Walker, D., Wolford, R. & Hager, G. L. The glucocorticoid receptor: rapid exchange with regulatory sites in living cells. Science 287, 1262–1265 (2000).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Tumbar, T., Sudlow, G. & Belmont, A. S. Large-scale chromatin unfolding and remodeling induced by VP16 acidic activation domain. J. Cell Biol. 145, 1341–1354 (1999).
Yao, J., Munson, K. M., Webb, W. W. & Lis, J. T. Dynamics of heat shock factor association with native gene loci in living cells. Nature 442, 1050–1053 (2006).
Janicki, S. M. et al. From silencing to gene expression: real-time analysis in single cells. Cell 116, 683–698 (2004).
Ben-Ari, Y. et al. The life of an mRNA in space and time. J. Cell Sci. 123, 1761–1774 (2010).
Probst, A. V., Dunleavy, E. & Almouzni, G. Epigenetic inheritance during the cell cycle. Nat. Rev. Mol. Cell Biol. 10, 192–206 (2009).
Sarge, K. D. & Park-Sarge, O. K. Gene bookmarking: keeping the pages open. Trends Biochem. Sci. 30, 605–610 (2005).
Sarge, K. D. & Park-Sarge, O. K. Mitotic bookmarking of formerly active genes: keeping epigenetic memories from fading. Cell Cycle 8, 818–823 (2009).
Zaidi, S. K. et al. Mitotic bookmarking of genes: a novel dimension to epigenetic control. Nat. Rev. Genet. 11, 583–589 (2010).
Rafalska-Metcalf, I. U., Powers, S. L., Joo, L. M., LeRoy, G. & Janicki, S. M. Single cell analysis of transcriptional activation dynamics. PLoS One 5, e10272 (2010).
Metivier, R. et al. Estrogen receptor-α directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell 115, 751–763 (2003).
Darzacq, X. et al. In vivo dynamics of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 14, 796–806 (2007).
Kumaran, R. I. & Spector, D. L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell Biol. 180, 51–65 (2008).
Prasanth, K. V., Sacco-Bubulya, P. A., Prasanth, S. G. & Spector, D. L. Sequential entry of components of the gene expression machinery into daughter nuclei. Mol. Biol. Cell 14, 1043–1057 (2003).
Pines, J. & Rieder, C. L. Re-staging mitosis: a contemporary view of mitotic progression. Nat. Cell Biol. 3, E3–E6 (2001).
Shav-Tal, Y. et al. Dynamics of single mRNPs in nuclei of living cells. Science 304, 1797–1800 (2004).
Yang, Z., He, N. & Zhou, Q. Brd4 recruits P-TEFb to chromosomes at late mitosis to promote G1 gene expression and cell cycle progression. Mol. Cell Biol. 28, 967–976 (2008).
Valls, E., Sanchez-Molina, S. & Martinez-Balbas, M. A. Role of histone modifications in marking and activating genes through mitosis. J. Biol. Chem. 280, 42592–42600 (2005).
Prescott, D. M. & Bender, M. A. Synthesis of RNA and protein during mitosis in mammalian tissue culture cells. Exp. Cell Res. 26, 260–268 (1962).
Muramoto, T., Muller, I., Thomas, G., Melvin, A. & Chubb, J. R. Methylation of H3K4 is required for inheritance of active transcriptional states. Curr. Biol. 20, 397–406 (2010).
Dey, A., Nishiyama, A., Karpova, T., McNally, J. & Ozato, K. Brd4 marks select genes on mitotic chromatin and directs postmitotic transcription. Mol. Biol. Cell 20, 4899–4909 (2009).
Jeppesen, P. Histone acetylation: a possible mechanism for the inheritance of cell memory at mitosis. Bioessays 19, 67–74 (1997).
Zippo, A. et al. Histone crosstalk between H3S10ph and H4K16ac generates a histone code that mediates transcription elongation. Cell 138, 1122–1136 (2009).
MacDonald, V. E. & Howe, L. J. Histone acetylation: where to go and how to get there. Epigenetics 4, 139–143 (2009).
Edmunds, J. W., Mahadevan, L. C. & Clayton, A. L. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J. 27, 406–420 (2008).
LeRoy, G., Rickards, B. & Flint, S. J. The double bromodomain proteins Brd2 and Brd3 couple histone acetylation to transcription. Mol. Cell 30, 51–60 (2008).
Kouskouti, A. & Talianidis, I. Histone modifications defining active genes persist after transcriptional and mitotic inactivation. EMBO J. 24, 347–357 (2005).
Liu, Y. et al. Structural basis and binding properties of the second bromodomain of Brd4 with acetylated histone tails. Biochemistry 47, 6403–6417 (2008).
Yang, Z. et al. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol. Cell 19, 535–545 (2005).
Jang, M. K. et al. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol. Cell 19, 523–534 (2005).
Masotti, A. et al. Comparison of different commercially available cationic liposome-DNA lipoplexes: Parameters influencing toxicity and transfection efficiency. Colloids Surf. B Biointerfaces 68, 136–144 (2009).
Andrews, J. M., Newbound, G. C. & Lairmore, M. D. Transcriptional modulation of viral reporter gene constructs following induction of the cellular stress response. Nucleic Acids Res. 25, 1082–1084 (1997).
Ip, J. Y. et al. Global impact of RNA polymerase II elongation inhibition on alternative splicing regulation. Genome Res. 21, 390–401 (2011).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Dey, A., Chitsaz, F., Abbasi, A., Misteli, T. & Ozato, K. The double bromodomain protein Brd4 binds to acetylated chromatin during interphase and mitosis. Proc. Natl Acad. Sci. USA 100, 8758–8763 (2003).
Larsen, A. & Weintraub, H. An altered DNA conformation detected by S1 nuclease occurs at specific regions in active chick globin chromatin. Cell 29, 609–622 (1982).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Marushige, K. Activation of chromatin by acetylation of histone side chains. Proc. Natl Acad. Sci. USA 73, 3937–3941 (1976).
Eskeland, R., Freyer, E., Leeb, M., Wutz, A. & Bickmore, W. A. Histone acetylation and the maintenance of chromatin compaction by polycomb repressive complexes. Cold Spring Harb. Symp. Quant. Biol. 75, 71–78 (2010).
Eskeland, R. et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 38, 452–464 (2010).
Mochizuki, K. et al. The bromodomain protein Brd4 stimulates G1 gene transcription and promotes progression to S phase. J. Biol. Chem. 283, 9040–9048 (2008).
Blobel, G. A. et al. A reconfigured pattern of MLL occupancy within mitotic chromatin promotes rapid transcriptional reactivation following mitotic exit. Mol. Cell 36, 970–983 (2009).
Kanno, T. et al. Selective recognition of acetylated histones by bromodomain proteins visualized in living cells. Mol. Cell 13, 33–43 (2004).
Umehara, T. et al. Structural basis for acetylated histone H4 recognition by the human BRD2 bromodomain. J. Biol. Chem. 285, 7610–7618 (2010).
Umehara, T. et al. Structural implications for K5/K12-di-acetylated histone H4 recognition by the second bromodomain of BRD2. FEBS Lett. 584, 3901–3908 (2010).
Bisgrove, D. A., Mahmoudi, T., Henklein, P. & Verdin, E. Conserved P-TEFb-interacting domain of BRD4 inhibits HIV transcription. Proc. Natl Acad. Sci. USA 104, 13690–13695 (2007).
Wu, S. Y. & Chiang, C. M. The double bromodomain-containing chromatin adaptor Brd4 and transcriptional regulation. J. Biol. Chem. 282, 13141–13145 (2007).
Chen, Y. C., Kappel, C., Beaudouin, J., Eils, R. & Spector, D. L. Live cell dynamics of promyelocytic leukemia nuclear bodies upon entry into and exit from mitosis. Mol. Biol. Cell 19, 3147–3162 (2008).
Linkert, M. et al. Metadata matters: access to image data in the real world. J. Cell Biol. 189, 777–782 (2010).
Mao, Y. S. et al. Essential and unique roles of PIP5K-γ and −α in Fcγ receptor-mediated phagocytosis. J. Cell Biol. 184, 281–296 (2009).
Steger, D. J. et al. DOT1L/KMT4 recruitment and H3K79 methylation are ubiquitously coupled with gene transcription in mammalian cells. Mol. Cell Biol. 28, 2825–2839 (2008).
Acknowledgements
We thank R. Singer (Albert Einstein College of Medicine, USA) for the MS2 clone, K. Ozato (NIH, USA) for the BRD4 clone, E. Verdin (UCSF, USA) for the BRD4-Δ1; Δ2; Δ1Δ2 constructs and C. D. Allis (The Rockfeller University, USA) for TH4 antibody. We greatly appreciate the gift of JQ1 from J. Bradner (Dana-Farber Cancer Institute, USA) and C. Vakoc. We thank C. Vakoc, A. A. Chakraborty and W. P. Tansey (Vanderbilt University, USA) for helpful comments and suggestions on ChIP assays. We also would like to thank Y. (S.) Mao, B. Zhang, M. S. Bodnar and M. R. Hübner, as well as other members of the Spector laboratory, for discussions and comments throughout the course of this work. This work was supported by a grant from NIH/NIGMS 42694, 42694-2OS1 (ARRA Supplement) to D.L.S.
Author information
Authors and Affiliations
Contributions
R.Z., T.N. and D.L.S. designed the research; R.Z. and T.N. carried out the experiments and analysed data; Y.F. wrote the MatLab software for quantitative analysis and also analysed data; Z.L. carried out the super-resolution structured illumination microscopy and locus size analysis. R.Z. and D.L.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2217 kb)
Supplementary Movie 1
Supplementary Information (MPG 383 kb)
Supplementary Movie 2
Supplementary Information (MPG 2468 kb)
Rights and permissions
About this article
Cite this article
Zhao, R., Nakamura, T., Fu, Y. et al. Gene bookmarking accelerates the kinetics of post-mitotic transcriptional re-activation. Nat Cell Biol 13, 1295–1304 (2011). https://doi.org/10.1038/ncb2341
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2341
- Springer Nature Limited
This article is cited by
-
Mitotic bookmarking redundancy by nuclear receptors in pluripotent cells
Nature Structural & Molecular Biology (2024)
-
CRISPR-dCas13-tracing reveals transcriptional memory and limited mRNA export in developing zebrafish embryos
Genome Biology (2023)
-
Release of Histone H3K4-reading transcription factors from chromosomes in mitosis is independent of adjacent H3 phosphorylation
Nature Communications (2023)
-
Lipid droplet-associated lncRNA LIPTER preserves cardiac lipid metabolism
Nature Cell Biology (2023)
-
BRD2 compartmentalizes the accessible genome
Nature Genetics (2022)