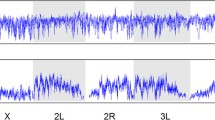
Abstract
Using the data provided by the Drosophila Population Genomics Project, we investigate factors that affect the genetic differentiation between Rwandan and French populations of D. melanogaster. By examining within-population polymorphisms, we show that sites in long introns (especially those >2000 bp) have significantly lower π (nucleotide diversity) and more low-frequency variants (as measured by Tajima's D, minor allele frequencies, and prevalence of variants that are private to one of the two populations) than short introns, suggesting a positive relationship between intron length and selective constraint. A similar analysis of protein-coding polymorphisms shows that 0-fold (degenerate) sites in more conserved genes are under stronger purifying selection than those in less conserved genes. There is limited evidence that selection on codon bias has an effect on differentiation (as measured by FST) at 4-fold (degenerate) sites, and 4-fold sites and sites in 8–30 bp of short introns ⩽65 bp have comparable FST values. Consistent with the expected effect of purifying selection, sites in long introns and 0-fold sites in conserved genes are less differentiated than those in short introns and less conserved genes, respectively. Genes in non-crossover regions (for example, the fourth chromosome) have very high FST values at both 0-fold and 4-fold degenerate sites, which is probably because of the large reduction in within-population diversity caused by tight linkage between many selected sites. Our analyses also reveal subtle statistical properties of FST, which arise when information from multiple single nucleotide polymorphisms is combined and can lead to the masking of important signals of selection.
Similar content being viewed by others
Introduction
Natural populations are often divided into subpopulations. Studying the extent to which different subpopulations are genetically differentiated has been of paramount importance in evolutionary genetics, as it provides a way to examine how different evolutionary forces such as genetic drift, natural selection and migration drive changes in the genome (reviewed in Chapter 7 of Charlesworth and Charlesworth, 2010).
Specifically, insights into fundamental processes such as historical demographic changes, (local) adaptation and speciation can be obtained by comparing patterns of genetic differentiation across different genomic regions (Wu, 2001; Weir and Hill, 2002; Charlesworth et al., 2003; Hey and Machado, 2003; Beaumont, 2005; Holsinger and Weir, 2009). For instance, by scanning for loci that show unusually high levels of differentiation relative to the rest of the genome, we can detect loci that are under diversifying selection, whereby different alleles are favoured in different subpopulations (Beaumont and Nichols, 1996; Beaumont and Balding, 2004; Foll and Gaggiotti, 2008; Excoffier et al., 2009). As another example, in a study comparing African and non-African humans, it was found that the X chromosome was substantially more diverged than the autosomes, over and above the null expectation based on the fact that there are four copies of each autosome for every three copies of the X chromosome, which in turn suggests that dispersal in humans may be sex-biased or that the X chromosome may have experienced repeated selection after the divergence of African and non-African populations (Keinan et al., 2009).
Genetic differentiation between subpopulations is often measured by Wright's FST (Wright, 1951), which is abbreviated as F in this study. F can be defined as the proportion of genetic variation explained by differences in allele frequencies between subpopulations (Charlesworth, 1998; Holsinger and Weir, 2009; Bhatia et al., 2013). F ranges between 0 and 1, which indicate no differentiation and fixed differences between subpopulations, respectively. Various genetic data, for example, single nucleotide polymorphisms (SNPs) and microsatellites, can be used to estimate F, but using statistical procedures that take into account biological properties of the data under consideration (for example, high versus low mutation rate) is vital for acquiring accurate estimates (Weir and Cockerham, 1984; Excoffier et al., 1992; Slatkin, 1995; Nagylaki, 1998; Holsinger and Weir, 2009).
Drosophila melanogaster, a classic model organism for population genetics, offers an invaluable system for studying population differentiation. Despite having a worldwide distribution in the present day, it is believed that the species originated in sub-Saharan Africa (David and Capy, 1988; Stephan and Li, 2007). The colonisation of Europe has been suggested to have taken place about 15 000 years ago (David and Capy, 1988; Stephan and Li, 2007; Duchen et al., 2013). The Americas and Australia were colonised much more recently, possibly in the past few hundred years (David and Capy, 1988; Stephan and Li, 2007; Duchen et al., 2013). By studying patterns of genetic differentiation, investigators have obtained evidence that American populations of D. melanogaster may be formed by admixture between African and European flies (Caracristi and Schlotterer, 2003). Multiple attempts have also been made to identify loci with unusually high F, which may have contributed to local adaptation to different habitats (Turner et al., 2008; Yukilevich et al., 2010; Kolaczkowski et al., 2011; Fabian et al., 2012; Langley et al., 2012; Pool et al., 2012; Campo et al., 2013).
These previous studies of D. melanogaster have mainly focused on determining the evolutionary relationship between subpopulations, quantifying the overall level of differentiation, and detecting genomic regions of interest using outlier scans. However, the role of purifying selection in shaping large-scale patterns of differentiation has not been well characterised, although it has been widely accepted that the majority of new mutations that affect fitness will have detrimental effects (Pal et al., 2006; Eyre-Walker and Keightley, 2007). Supporting this view, it has been estimated that only between 1 and 2% of new nonsynonymous mutations in D. melanogaster are (weakly) positively selected, and about 6% are nearly neutral (that is, |Nes|⩽1, where Ne is the effective population size and s the selection coefficient), and the remaining are deleterious (|Nes|>1) (Keightley and Eyre-Walker, 2007; Eyre-Walker and Keightley, 2009; Schneider et al., 2011). Thus, we are interested in testing the following predictions based on population genetic theory of subdivided populations (reviewed in Chapter 7 of Charlesworth and Charlesworth, 2010): (i) purifying selection reduces differentiation between populations at functionally important regions; (ii) the level of reduction is positively correlated with the level of selective constraint. Answering these questions will help us better understand the sources of variation in genetic differentiation across the genome, which is important for example, in interpreting results obtained from genome scans (Beaumont and Nichols, 1996; Beaumont and Balding, 2004; Foll and Gaggiotti, 2008; Excoffier et al., 2009).
We will address the questions raised above by making use of the high-quality whole-genome resequencing data published by the Drosophila Population Genomics Project for one French population and one Rwandan population (Langley et al., 2012; Pool et al., 2012). In addition to protein-coding regions, we investigate introns, as previous studies have shown strong evidence that these genomic regions are under substantial selective constraints, probably as a result of the presence of cis-regulatory elements and noncoding RNA genes (Bergman and Kreitman, 2001; Parsch, 2003; Andolfatto, 2005; Haddrill et al., 2005a; Halligan and Keightley, 2006; Casillas et al., 2007; Roy et al., 2010).
Our study proceeds as follows. First, we present an overview of patterns of genetic variation both within and between populations using data from genomic regions where crossing over occurs (crossover (C) regions). We are interested in understanding whether patterns of differentiation at 4-fold degenerate (hereafter 4-fold) sites are affected by selection on codon usage, and whether 4-fold sites and putatively neutral sites in 8–30 bp regions of introns ⩽65 bp (Halligan and Keightley, 2006; Parsch et al., 2010) are comparable with respect to levels of differentiation. These are intended to identify putatively neutral sites which can be used as a reference in the study of the effects of purifying selection on genetic differentiation. We then examine the relationship between KA (nonsynonymous divergence) and diversity/differentiation patterns at 0-fold degenerate (hereafter 0-fold) sites in protein-coding regions, as well as the relationship between intron length and diversity/differentiation patterns in intronic regions. Finally, we compare non-crossover (NC) regions (for example, the fourth chromosome) and C regions regarding differentiation patterns, study the relative contribution of selection and genetic linkage, and examine the correlation between local recombination rates and F at putatively neutral sites.
Materials and Methods
Data acquisition
To obtain polymorphism and divergence data, we downloaded FASTQ files from the Drosophila Population Genomics Project (http://www.dpgp.org/dpgp2/candidate/) for 17 Rwandan D. melanogaster samples (RG18N, RG19, RG2, RG22, RG24, RG25, RG28, RG3, RG32N, RG33, RG34, RG36, RG38N, RG4N, RG5, RG7 and RG9), which have been estimated to have the lowest estimated levels of admixture with European populations (less than 3%, see Figure 3b of Pool et al., 2012). We also selected seven samples from the French population (FR14, FR151, FR180, FR207, FR217, FR310 and FR361). We will refer to these two samples as RG and FR, respectively. We further masked any regions of the African samples with evidence of admixture from European populations, using the admixture coordinates reported by Pool et al. (2012). Sites with a quality score below 31 (equivalent to a PHRED score of 48, and approximately equivalent to one error per 100 kb; see Pool et al., 2012) were also masked.
From the FASTQ files, we extracted protein-coding regions in C regions, which we abbreviate as CDS-C, using gene annotations from FlyBase release version 5.44 (www.flybase.org) and made FASTA files containing all samples (24 alleles). For each D. melanogaster gene with multiple transcripts, we chose one transcript randomly.
Introns belonging to our chosen transcript were extracted, and were further processed by masking any coding regions that form part of the other transcripts. Only introns occurring in C regions were retained (polymorphism data for introns in NC regions were of significantly lower quality, and were therefore excluded).
Protein-coding regions in NC regions of the D. melanogaster genome, abbreviated as CDS-NC, were obtained from Campos et al. (2012). These data included five unlinked regions: N2 (genes located in heterochromatic regions near the centromere of the second chromosome), N3 (genes located in heterochromatic regions near the centromere of the third chromosome), N4 (the fourth chromosome), NXc (X-linked genes near the centromere) and NXt (X-linked genes near the telomere).
For all CDS-C and CDS-NC, we selected D. yakuba as an out-group to avoid any major influence of ancestral polymorphisms on the estimation of sequence divergence, which can potentially create spurious correlations between sequence divergence and recombination (for example, Cutter and Choi, 2010). One-to-one orthologous D. yakuba sequences were obtained from FlyBase (available at ftp://ftp.flybase.net/releases/FB2012_02/precomputed_files/genes/gene_orthologs_fb_2012_02.tsv.gz). We then performed amino-acid sequence alignments using MAFFT (Katoh et al., 2002). These amino-acid sequence alignments were translated back to nucleotides using custom scripts in PERL to produce in-frame coding sequence alignments that included the 24 D. melanogaster alleles and the D. yakuba reference.
For introns, we used D. simulans as an out-group because we considered D. yakuba too distant for producing reliable alignments, because of the increased prevalence of indels in non-coding regions. We obtained orthologous intronic D. simulans sequences from Hu et al. (2013), which was based on an updated D. simulans genome assembly and careful alignment procedures to preserve gene structures (http://genomics.princeton.edu/AndolfattoLab/w501_genome_files/alnMSY.tar.gz).
Recombination rate for the midpoint of all alignments was obtained using the Drosophila melanogaster Recombination Rate Calculator v2.3 (Fiston-Lavier et al., 2010) and the high-resolution genetic map published recently by Comeron et al. (2012).
Sequence analysis
For CDS-C and CDS-NC, we calculated KA and KS (the numbers of nonsynonymous and synonymous substitutions per nonsynonymous and synonymous site, respectively) using the kaks() function from the seqinr package (Charif and Lobry, 2007) in R (http://www.r-project.org/), which implements the method of Li (1993). For introns, we calculated divergence (K) to the D. simulans reference using the dist.dna() function in the ape package of R (Paradis et al., 2004), with the ‘K80’ method (Kimura, 1980). For conducting analyses using polymorphism data in the two D. melanogaster samples, we split CDS-C and CDS-NC into 0-fold degenerate sites and 4-fold degenerate sites by analysing the alignments codon by codon. A codon column was retained if the following requirements were met: (i) data from all individuals were available; (ii) it had at most one SNP. These were to avoid uncertainty of the order of mutations in codons with multiple SNPs. We retained 7235 autosomal and 1150 X-linked CDS alignments for which we had both more than 10 bp of 0-fold sites and more than 10 bp of 4-fold sites.
We split introns into short (⩽65 bp long) and long (>65 bp long) classes, and further trimmed short introns to retain positions 8–30 from the 5′ end (hereafter the SI sites), to retain sites under the least amount of selective constraint (Halligan and Keightley, 2006; Parsch et al., 2010). This left us with 7483 autosomal and 752 X-linked short introns, and 8851 autosomal and 1869 X-linked long introns. To keep the sample size the same as the CDS data, only intronic sites where data from all individuals were available were retained. This requirement appears to be conservative with respect to detecting SNPs (note also that regions within 3 bp of an indel were also masked by Drosophila Population Genomics Project; see Pool et al., 2012 and http://www.dpgp.org/dpgp2/DPGP2.html). For instance, π (nucleotide diversity) estimated using data from SI sites that fulfilled the above criterion was 0.0145 in the RG sample, whereas the estimate increased to 0.0164 when we retained, within the same genomic regions, all sites that had data from at least two individuals. However, as we will show in the Results, there is no detectable difference between the SI and 4-fold sites in terms of skewness of allele frequency spectrum (as measured by Tajima’s D), average minor allele frequency (MAF), and FST. Our conservative data filtering procedure is unlikely to bias our analysis of population differentiation.
Because of the complete linkage between the CDS-NC genes located within a NC region, each non-recombining region of the genome effectively represents a single locus. Therefore all genes within a single NC region were concatenated and analysed as a single gene. In total, we have three autosomal NC regions, which are N2, N3 and N4, and two X-linked NC regions, which are NXc and NXt. These loci were kept intact in the permutation tests used to compare values of summary statistics calculated using data from NC and C regions.
Nucleotide diversity (π), Tajima’s D (Tajima, 1989) and relative Tajima’s D, (Schaeffer, 2002) were calculated using nuc.div() and a modified version of the tajima.test() function, both from the pegas package in R (Paradis, 2010). Because conclusions drawn from Tajima's D and relative Tajima's D are identical, only the former is presented. Permutation tests were carried out to assess whether these statistics were different between different types of sites. For instance, to investigate whether values of Tajima’s D at 4-fold sites and SI sites were comparable, 10 000 pseudosamples were generated by randomly shuffling both the 4-fold and SI sites in the data (SNPs from a single locus were shuffled as a unit), such that in each pseudosample there were similar numbers of SNPs in the two ‘site classes’ as in the real data.
To assess the effects of selection on codon bias on population differentiation at 4-fold sites, we calculated frequency of optimal codons (Fop) with CodonW (Peden, 1999), using the built-in table of optimal codons for D. melanogaster.
Measuring population differentiation
Levels of differentiation between the two populations of D. melanogaster were measured by Wright's FST, which is abbreviated as F. We used the definition of Weir and Cockerham (1984), which can be expressed as

where πB is the expected divergence between a pair of alleles sampled from two different populations, and πS is the expected within-population diversity (see also Charlesworth, 1998; Keinan et al., 2007). Previous investigations (Maruki et al., 2012; Jakobsson et al., 2013) of the dependence of F on MAF were based on a different definition of F put forward by Nei (1973). We therefore derive the maximum value of Weir and Cockerham's definition of F as a function of MAF.
Consider a population divided into two subpopulations. We examine a biallelic locus. The frequency of one of the two alleles in the k-th subpopulation is referred to as pk (k=1 or 2). It can be shown that πB=p1(1−p2)+p2(1−p1) and πS=p1(1−p1)+p2(1−p2) (Charlesworth, 1998). Substituting these into Equation (1), F can be rewritten as

where δ=|p1−p2| and σ=p1+p2. Without loss of generality, we assume that 1⩽σ⩽2. Three properties are of use: (i) MAF=1–σ/2; (ii) 0⩽2σ–σ2⩽1 for 1⩽σ⩽2; (iii) 0⩽δ⩽(2–σ)2. Rearranging Equation (2), we deduce that

Because MAF=1–σ/2, the above inequality is equivalent to max(F)⩽2 MAF. In Supplementary Figure S1, we display the differences between the upper bounds of F derived here and that obtained in previous studies using Nei’s F (Maruki et al., 2012; Jakobsson et al., 2013). It can be seen that F can only assume a very restrictive range of values when MAF is small.
The estimator of F proposed by Hudson et al. (1992) (see also Keinan et al., 2007; Bhatia et al., 2013) was employed:

where  and
and  are estimates of πB and πS obtained from data. Equation (4) can be calculated using information from a single SNP. To combine information from multiple SNPs, the following two methods were used (Weir and Cockerham, 1984; Bhatia et al., 2013):
are estimates of πB and πS obtained from data. Equation (4) can be calculated using information from a single SNP. To combine information from multiple SNPs, the following two methods were used (Weir and Cockerham, 1984; Bhatia et al., 2013):

and

where S is the number of SNPs, and  and
and  are values of the terms defined in Equation (4) obtained using data from the i-th SNP.
are values of the terms defined in Equation (4) obtained using data from the i-th SNP.
It should be noted that FU gives equal weight to all SNPs, whereas FW gives more weight to SNPs with higher expected levels of polymorphism. In other words, FU is expected to be more sensitive to the presence of SNPs with low MAFs, but FW is dominated by SNPs that are on average more polymorphic. To see this more explicitly, assume that we are combining information from two SNPs (that is, S=2). We add a subscript j to the symbols defined above to signify the locus under consideration, so that we have pjk, σj and δj=|pj1−pj2|. We further assume that 2>σ1⩾σ2⩾1. Note that σj are regarded as parameters (for example, a SNP under strong selective constraints is expected to have a larger σ (that is, a smaller MAF) than a neutral SNP). Some straightforward algebra leads to the following results: (i) max[F(σ1)]⩽max[FU]⩽max[F(σ2)]; (ii) max[F(σ1)]⩽max[FW]⩽max[F(σ2)], where max[F(σj)]=2−σj (these results hold when S>2; proof not shown). To see the differential sensitivities to SNPs with small MAFs, we define Δ1(U)=max[FU]−max[F(σ1)], Δ2(U)=max[F(σ2)]−max[FU], Δ1(W)=max[FW]−max[F(σ1)], and Δ2(W)=max[F(σ2)]−max[FW]. Using Equation (3), we show that Δ1(U)/Δ2(U)=1, but Δ1(W)/Δ2(W)=(2−σ2)/(2−σ1)⩾1. Thus, the behaviour of FW is more akin to that of the more polymorphic SNP (that is, max[FW] is closer to max[F(σ2)] than to max[F(σ1)]). As we will see later, this property of FW can lead to the masking of important signatures of evolution when SNPs with different properties are combined.
Results
Genome-wide polymorphism patterns in crossover regions
Table 1 presents summaries of polymorphism patterns for autosomal (A) and X-linked (X) loci situated in genomic regions where crossing-over occurs (C regions). For ease of presentation, we will refer to nucleotide diversity, π, calculated using 0-fold sites, 4-fold sites and SI sites (positions 8–30 from the 5′ end of short introns ⩽65 bp) as π0, π4 and πSI, respectively; a similar notational convention will be used for other statistics. For both A and X, and in both the Rwandan (RG) and French (FR) samples, π0, Tajima's D0 (Tajima 1989), and MAF0 are significantly smaller than the corresponding estimates obtained from 4-fold and SI sites (Ppermutation<0.001 in all cases), consistent with the well-known fact that most nonsynonymous mutations are deleterious (Pal et al., 2006; Eyre-Walker and Keightley, 2007), and are therefore kept at low frequencies in the population by purifying selection (Kimura, 1983). Previous studies have suggested that SI sites may be neutrally evolving (Halligan and Keightley, 2006; Parsch et al., 2010). In our data set, πSI seems to be somewhat smaller than π4, which may be due to the stringent data filtering procedure we employed (see Materials and Methods), or the higher GC content at 4-fold sites compared to intronic sites, which in turn is expected to result in an increased mutation rate in 4-fold sites (Singh et al., 2005; Keightley et al., 2009). There is, however, no statistically discernible difference with respect to either MAF or Tajima’s D between 4-fold and SI sites (Table 1; Ppermutation>0.1 for both A and X).
The FR sample has a lower level of diversity than RG for all three types of sites (Table 1), reflecting a loss of genetic variation induced by population bottlenecks which are believed to have occurred as the species migrated out of Africa (Haddrill et al., 2005b; Li and Stephan, 2006; Thornton and Andolfatto, 2006; Hutter et al., 2007; Duchen et al., 2013). The difference in π0 between the two populations is somewhat smaller than those observed for π4 and πSI (for example, on A, π0(FR)/π0(RG)=0.83 versus π4(FR)/π4(RG)=0.77). This is probably because more 0-fold sites are under strong selective constraint, so that variants at these sites behave almost deterministically, and are therefore less sensitive to demographic changes (for example, Zeng, 2013).
To inspect overall patterns of genetic differentiation between the RG and FR populations, we calculated FST (abbreviated here as F; see Equation (1) in Materials and Methods), as defined by Weir and Cockerham (1984), using the estimator of Hudson et al. (1992). Two approaches were employed to combine information over multiple SNPs: un-weighted mean F (Equation (5)) and weighted mean F (Equation (6)), which will be referred to as FU and FW, respectively. Because most nonsynonymous mutations are likely to be deleterious, it is expected that levels of population differentiation at these selectively constrained sites should be lower than those at less constrained sites (for example, 4-fold sites) (Barreiro et al., 2008; Maruki et al., 2012). Surprisingly, values of  , estimated using either the autosomal or X-linked data, are not statistically different from those of either
, estimated using either the autosomal or X-linked data, are not statistically different from those of either  or
or  (Table 1; Ppermutation>0.1 in all cases). There is also no detectable difference between
(Table 1; Ppermutation>0.1 in all cases). There is also no detectable difference between  and
and  (Ppermutation>0.1 for both A and X). In contrast,
(Ppermutation>0.1 for both A and X). In contrast,  was found to be significantly smaller than both
was found to be significantly smaller than both  and
and  (Ppermutation<0.001 for both A and X), whereas the differences between
(Ppermutation<0.001 for both A and X), whereas the differences between  and
and  remain non-significant (Ppermutation>0.1 for both A and X). The patterns obtained from FU are therefore more compatible with the a priori expectation that 0-fold sites are on average more constrained than 4-fold and SI sites. We will investigate causes for the lack of difference between
remain non-significant (Ppermutation>0.1 for both A and X). The patterns obtained from FU are therefore more compatible with the a priori expectation that 0-fold sites are on average more constrained than 4-fold and SI sites. We will investigate causes for the lack of difference between  and either
and either  or
or  in a later section.
in a later section.
Several differences between A and X are of note (Table 1). Firstly, consistent with previous reports (Caracristi and Schlotterer, 2003; Hutter et al., 2007; Charlesworth, 2012b; Pool et al., 2012; Campos et al., 2013), the X:A ratio in diversity at putatively neutral sites (that is, 4-fold and SI sites) is about 1 in the RG population (π4(X)/π4(A)=1.08 and πSI(X)/πSI(A)=1.10), higher than the null expectation of 3/4. Secondly, the reduction in diversity in FR is more pronounced for X than A for all three types of sites (for example, π4(FR)/π4(RG)=0.41 and 0.77 for X and A, respectively), as reported in previous investigations (Caracristi and Schlotterer, 2003; Hutter et al., 2007). Finally, the extent of population differentiation at both 4-fold and SI sites, as measured by either FU or FW, is significantly higher on the X than on A (Ppermutation<0.001 for all comparisons). This is probably largely driven by the greater reduction in diversity on the X in non-African populations, as values of Dxy, the mean number of nucleotide substitutions between sequences taken from different subpopulations (Nei and Miller, 1990), are comparable between A and X in this study: Dxy,4=1.65 and 1.64%, and Dxy,SI=1.51 and 1.58%. A systematic examination of possible causes of the apparent differences between A and X is beyond the scope of this study; the interested reader can refer to previous studies of this topic (Charlesworth, 2001; Pool and Nielsen, 2007; Singh et al., 2007; Pool and Nielsen, 2008; Yukilevich et al., 2010; Charlesworth, 2012b; Campos et al., 2013). In what follows, results obtained from A and X will be presented separately.
Limited evidence for selection on codon usage bias affecting patterns of population differentiation at 4-fold degenerate sites
To investigate whether selection on codon usage bias affects differentiation patterns at 4-fold sites, we first examined the relationship between  and Fop, as the latter is well known to be correlated with the intensity of selection on codon usage bias (reviewed in Hershberg and Petrov, 2008; Zeng and Charlesworth, 2009). Considering the large variance of the F estimators and the dearth of SNPs in individual genes, we grouped the genes into equal-sized bins with similar numbers of SNPs at 4-fold sites. As shown in Supplementary Figure S2A, Fop and
and Fop, as the latter is well known to be correlated with the intensity of selection on codon usage bias (reviewed in Hershberg and Petrov, 2008; Zeng and Charlesworth, 2009). Considering the large variance of the F estimators and the dearth of SNPs in individual genes, we grouped the genes into equal-sized bins with similar numbers of SNPs at 4-fold sites. As shown in Supplementary Figure S2A, Fop and  are not correlated on A (Kendall's τ=−0.01, P>0.1). On the X, some evidence for a weak negative correlation was obtained (Supplementary Figure S2B), but it is not statistically significant (Kendall's τ=−0.6, P=0.13). When
are not correlated on A (Kendall's τ=−0.01, P>0.1). On the X, some evidence for a weak negative correlation was obtained (Supplementary Figure S2B), but it is not statistically significant (Kendall's τ=−0.6, P=0.13). When  was considered, no correlation was found on either A or X (Supplementary Figures S2E and F). To investigate this further, for the genes within each bin on the X, we tested whether
was considered, no correlation was found on either A or X (Supplementary Figures S2E and F). To investigate this further, for the genes within each bin on the X, we tested whether  differed from
differed from  statistically. Amongst the six bins, no evidence of a significant difference was found for the first four bins, whereas the differences were marginally significant for the last two bins with highest Fop (Ppermutation=0.04 and 0.05, respectively). Similarly, we did not detect any correlation between KS and either
statistically. Amongst the six bins, no evidence of a significant difference was found for the first four bins, whereas the differences were marginally significant for the last two bins with highest Fop (Ppermutation=0.04 and 0.05, respectively). Similarly, we did not detect any correlation between KS and either  or
or  (Supplementary Figure S2).
(Supplementary Figure S2).
Overall, there is limited evidence that selection on codon usage bias is strong enough to substantially alter patterns of genetic differentiation at 4-fold sites. Considering that 4-fold and SI sites in C regions are comparable with respect to both MAF and F, in what follows, we will use population differentiation patterns obtained from the two types of site as neutral standards, and will refer to them as putatively neutral sites.
Evolutionarily conserved genes are under stronger purifying selection and have reduced F at 0-fold degenerate sites
Genes in C regions were divided into equal-sized bins (with similar numbers of SNPs) based on their KA values between D. melanogaster and D. yakuba. We inspected polymorphism patterns in the RG sample as a function of KA; a qualitatively identical set of results were obtained using the FR sample (Supplementary Figure S3). On both A and X, KA was found to be significantly positively correlated with both π0 (Figures 1a and b; A: Kendall's τ=0.989 and P<0.001; X: Kendall's τ=1 and P=0.009) and Tajima's D0 (Figures 1c and d; A: Kendall's τ=0.884, P<0.001; X: Kendall's τ=0.867 and P=0.024). No statistically significant relationship was found when comparing KA with Tajima's D4 (Figures 1c and d; Kendall's τ=−0.2 and −0.333, P>0.1, for X and A), although there is a negative correlation between KA and π4 on A (Figure 1a; Kendall's τ=−0.6, P<0.001) (see also Andolfatto, 2007; Haddrill et al., 2011). In particular, on both A and X, π0 and Tajima's D0 approach π4 and Tajima’s D4, respectively, as KA increases. In contrast, values of π4 and Tajima's D4, regardless of the KA bin from which they were obtained, remain similar to the values of πSI and Tajima’s DSI. These results suggest that 0-fold sites are under stronger constraints than 4-fold and SI sites, and that 0-fold sites in genes with smaller KA are, on average, under stronger purifying selection. We obtained the same results when we used the D. simulans genome as an out-group (Supplementary Figure S4).
Polymorphism patterns within 17 Rwandan D. melanogaster lines for coding sequence (CDS) binned by KA value (to D. yakuba), and for sites in the 8–30 bp regions of short introns ⩽65 bp (SI sites). (a) Nucleotide diversity (π) for autosomal CDS-C and (b) X-linked CDS-C regions; (c) Tajima’s D for autosomal CDS-C regions and (d) X-linked CDS-C regions. The x axes show the maximum KA value in each bin. Symbols: 0-fold degenerate sites—open circles; 4-fold degenerate sites—open triangles; SI sites—open red squares.
Figures 2a and b show that evolutionarily conserved genes have significantly smaller  (A: Kendall's τ=0.663, P<0.001; X: Kendall’s τ=0.867, P=0.02). Again, we obtained the same result when using D. simulans as the out-group (Supplementary Figure S5). The pattern remains statistically significant for autosomes when
(A: Kendall's τ=0.663, P<0.001; X: Kendall’s τ=0.867, P=0.02). Again, we obtained the same result when using D. simulans as the out-group (Supplementary Figure S5). The pattern remains statistically significant for autosomes when  was considered (Supplementary Figure S6). The reduction in F0 for genes with smaller KA is associated with a strong reduction in MAF0 (Figures 2c and d) and an increase in the proportion of 0-fold SNPs that are private to one of the two populations (Figures 2e and f), both of which are hallmarks of selection against deleterious mutations (cf., recent findings in humans; Nelson et al., 2012; Fu et al., 2013), and are expected to drive both FU and FW downwards, as shown in Materials and Methods (see also Maruki et al., 2012; Bhatia et al., 2013; Jakobsson et al., 2013). For the 4-fold sites on both A and X, no correlation with KA was observed for FU, FW, MAF and the proportion of private SNPs (Figure 2; P>0.1 in all cases based on Kendall's τ).
was considered (Supplementary Figure S6). The reduction in F0 for genes with smaller KA is associated with a strong reduction in MAF0 (Figures 2c and d) and an increase in the proportion of 0-fold SNPs that are private to one of the two populations (Figures 2e and f), both of which are hallmarks of selection against deleterious mutations (cf., recent findings in humans; Nelson et al., 2012; Fu et al., 2013), and are expected to drive both FU and FW downwards, as shown in Materials and Methods (see also Maruki et al., 2012; Bhatia et al., 2013; Jakobsson et al., 2013). For the 4-fold sites on both A and X, no correlation with KA was observed for FU, FW, MAF and the proportion of private SNPs (Figure 2; P>0.1 in all cases based on Kendall's τ).
Differentiation patterns between 7 French and 17 Rwandan D. melanogaster lines for coding sequence (CDS) binned by KA value (to D. yakuba), and for SI sites. (a) Unweighted mean FST (FU; Equation (5)) for autosomal coding CDS-C and (b) X-linked CDS-C regions; (c) population-average MAF for autosomal CDS-C regions and (d) X-linked CDS-C regions; (e) the proportion of SNPs per bin in which one allele was private to one of the D. melanogaster populations for autosomal CDS-C regions and (f) X-linked CDS-C regions. Symbols: 0-fold degenerate sites—open circles; 4-fold degenerate sites—open triangles; SI sites—open red squares.
The data presented in Figures 1 and 2 suggest that the lack of difference between  and either
and either  or
or  reported in the previous section is probably because of the fact that FW gives more weight to SNPs with higher expected levels of polymorphism (for example, nearly neutral variants), as we have shown in Materials and Methods. In other words, when all 0-fold sites in C regions were analysed together (Table 1), the effects of purifying selection on a substantial fraction of 0-fold sites were probably masked by those 0-fold sites that are nearly neutrally evolving. Consequently, the overall distribution of
reported in the previous section is probably because of the fact that FW gives more weight to SNPs with higher expected levels of polymorphism (for example, nearly neutral variants), as we have shown in Materials and Methods. In other words, when all 0-fold sites in C regions were analysed together (Table 1), the effects of purifying selection on a substantial fraction of 0-fold sites were probably masked by those 0-fold sites that are nearly neutrally evolving. Consequently, the overall distribution of  appears non-distinguishable from those of
appears non-distinguishable from those of  and
and  . In contrast, FU gives equal weight to all SNPs. Considering that the value of F when calculated using a single SNP is constrained by MAF (see Equation (3) in Materials and Methods), FU is expected to be more sensitive to the action of purifying selection than FW, consistent with the observation reported above. In the Discussion, we will further explore the implications of these statistical properties of F, which arise when information from multiple SNPs is combined.
. In contrast, FU gives equal weight to all SNPs. Considering that the value of F when calculated using a single SNP is constrained by MAF (see Equation (3) in Materials and Methods), FU is expected to be more sensitive to the action of purifying selection than FW, consistent with the observation reported above. In the Discussion, we will further explore the implications of these statistical properties of F, which arise when information from multiple SNPs is combined.
Longer introns are under stronger selective constraints and are less differentiated
In agreement with earlier findings (Haddrill et al., 2005a; Halligan and Keightley, 2006), longer introns tend to have lower divergence (K) between D. melanogaster and D. simulans (A: Kendall’s τ=−0.635, P<0.001; X: Kendall's τ=−0.486, P<0.001; Figures 3a and b), probably as a result of the presence of functional elements that are subject to purifying selection (Bergman and Kreitman, 2001; Parsch, 2003; Andolfatto, 2005; Haddrill et al., 2005a; Halligan and Keightley, 2006; Casillas et al., 2007; Roy et al., 2010). Here, we report further support for this hypothesis by examining within-population polymorphism patterns as a function of intron length. Consistent with the action of purifying selection, longer introns have lower π (Figures 3c and d) and more negative Tajima’s D (Figures 3e and f) compared with 4-fold and SI sites (similar results were observed in the FR sample; see Supplementary Figure S7). Interestingly, the patterns of divergence and polymorphism level off for introns longer than 2000 bp. Using the RG sample, the values of π and Tajima's D obtained from introns longer than 2000 bp are 0.0072 and −0.5476 for A, and 0.0076 and −0.9013 for X, respectively; all these values are substantially lower than the corresponding values observed at 4-fold and SI sites, but are higher than those obtained from 0-fold sites (see Table 1). Furthermore, the KA values for CDS in C regions between D. melanogaster and D. simulans are 0.015 and 0.018 for A and X, respectively, which are significantly smaller than the values of K for long introns >2000 bp on A and X, which are 0.061 and 0.074, respectively (Mann–Whitney U test, P<0.001). These results imply that long introns, especially those >2000 bp, are more constrained than the 4-fold and SI sites, but probably contain fewer strongly selected sites than 0-fold sites.
Divergence and polymorphism patterns for intronic sites binned by intron length. (a) Divergence (K) between D. melanogaster and D. simulans for autosomal introns and (b) X-linked introns; (c) nucleotide diversity (π) for autosomal introns and (d) X-linked introns; (e) Tajima’s D for autosomal introns and (f) X-linked introns. The x axes display the maximum intron length in each bin. Note that the number of SNPs in each autosomal intron bin is roughly the same as that in the autosomal SI bin; the same applies to the X-linked data. Symbols: Long intronic sites—open circles; positions 8–30 bp sites of short introns ⩽65 bp (SI sites)—open red squares.
Estimates of FW, when calculated using sites from introns more than 65 bp in length, were 0.171 and 0.283 for A and for X, respectively. None of these was found to be statistically different from the corresponding values estimating using 4-fold and SI sites reported in Table 1 (Ppermutation>0.1 in all cases). FU for introns >65 bp were 0.157 and 0.174 for A and X, respectively, both of which were significantly smaller than both  and
and  (Ppermutation<0.001 in all cases). There is a clear negative relationship between FU and intron length (Figures 4a and b; for A and X, Kendall's τ=−0.356 and −0.364; P=0.010 and P<0.001, respectively), which mirrors that between MAF (or the prevalence of private SNPs) and intron length (Supplementary Figure S8), and is consistent with the expected effect of purifying selection on genetic differentiation between populations. The relationship between differentiation and intron length is weaker when FW was analysed (Supplementary Figure S8; for A and X, Kendall's τ=−0.271 and −0.146, and P=0.05 and 0.16, respectively). These differences between FW and FU can be explained by the fact that fewer sites in introns >65 bp are expected to be strongly selected compared with 0-fold sites. As discussed in the previous section, FW, which tends to reflect differentiation patterns at neutral sites in the data, is less likely to recover signatures of purifying selection compared to FU.
(Ppermutation<0.001 in all cases). There is a clear negative relationship between FU and intron length (Figures 4a and b; for A and X, Kendall's τ=−0.356 and −0.364; P=0.010 and P<0.001, respectively), which mirrors that between MAF (or the prevalence of private SNPs) and intron length (Supplementary Figure S8), and is consistent with the expected effect of purifying selection on genetic differentiation between populations. The relationship between differentiation and intron length is weaker when FW was analysed (Supplementary Figure S8; for A and X, Kendall's τ=−0.271 and −0.146, and P=0.05 and 0.16, respectively). These differences between FW and FU can be explained by the fact that fewer sites in introns >65 bp are expected to be strongly selected compared with 0-fold sites. As discussed in the previous section, FW, which tends to reflect differentiation patterns at neutral sites in the data, is less likely to recover signatures of purifying selection compared to FU.
Regions with reduced recombination tend to have higher F
It is known that genomic regions that lack crossing over (NC regions) have very different patterns of divergence and polymorphism than those seen in C regions (Haddrill et al., 2007; Betancourt et al., 2009; Arguello et al., 2010; Campos et al., 2012; Campos et al., 2014). In Table 2, we present summary statistics of the NC data pertinent to the current study (see Materials and Methods for a list of the NC regions considered). It can be seen that, for both 0-fold and 4-fold sites, values of F in NC regions are generally higher than those obtained using the same type of site in C regions, regardless of the way in which information from multiple SNPs was combined. Specifically, the average KA to D. yakuba is about 0.05 for the NC loci (Campos et al., 2012).  calculated using autosomal and X-linked NC data are 0.1817 and 0.3012, respectively (Table 2), higher than the values of 0.1569 and 0.1685 for autosomal and X-linked genes in C regions spanning the same KA values (Figures 2a and b; Ppermutation=0.05 for A and Ppermutation<0.001 for X).
calculated using autosomal and X-linked NC data are 0.1817 and 0.3012, respectively (Table 2), higher than the values of 0.1569 and 0.1685 for autosomal and X-linked genes in C regions spanning the same KA values (Figures 2a and b; Ppermutation=0.05 for A and Ppermutation<0.001 for X).
It should be noted that the elevation in F in NC regions is probably caused by an extreme reduction in within-population diversity induced by tight linkage between a large number of selected sites (Table 2; Kaiser and Charlesworth, 2009; O'Fallon et al., 2010; Seger et al., 2010; Zeng and Charlesworth, 2010). This is because F is a relative measure of differentiation (see Equation (1)), and therefore all else being equal, F is expected to be elevated by forces that reduce within-population diversity (that is, πS in Equation (1)), irrespective of whether diversifying selection or reduced gene flow has affected the genomic region under study (Charlesworth, 1998; Noor and Bennett, 2009).
To further examine the effects of selection at linked sites, we inspect the correlation between F at putatively neutral sites and local recombination rates in C regions. Figure 5 presents results based on autosomal loci, where it can be seen that  is reduced with more frequent recombination (Kendall's τ=−0.474, P=0.004; the data point obtained from the NC regions was not included in the calculation). However, there is no statistically significant relationship between recombination rate and
is reduced with more frequent recombination (Kendall's τ=−0.474, P=0.004; the data point obtained from the NC regions was not included in the calculation). However, there is no statistically significant relationship between recombination rate and  (Figure 5b; Kendall’s τ=−0.179 and P=0.28). Weak negative correlations were also found on the X chromosome for 4-fold and SI sites (Supplementary Figure S9). The patterns remained unchanged when FW was used (Supplementary Figure S10).
(Figure 5b; Kendall’s τ=−0.179 and P=0.28). Weak negative correlations were also found on the X chromosome for 4-fold and SI sites (Supplementary Figure S9). The patterns remained unchanged when FW was used (Supplementary Figure S10).
Discussion
By using the high-quality data provided by the Drosophila Population Genomics Project, we have found that evolutionary conserved regions (that is, genes with lower KA and longer introns) show clear evidence of more intense on-going purifying selection than less conserved genomic regions, which can be detected by analysing patterns of genetic variation both within and between subpopulations. The negative correlation between π and intron length reported in Figure 3 extends the study by Parsch et al. (2010) who examined a much smaller data set and did not find evidence of such a correlation. Because we did not find support for a correlation between local recombination rate and intron length (Kendall's τ=−0.004 and 0.011 for A and X, respectively, and P>0.1 in both cases; Supplementary Figure S11) (cf., Carvalho and Clark, 1999; Comeron and Kreitman, 2000), the relationship is unlikely to be driven by the well-known positive correlation between diversity and recombination. It is unclear why the effect of intron length levels off for introns longer than 2000 bp. Analysis of theoretical models (for example, Ometto et al. 2005) and improved annotation of non-coding functional elements (for example, Roy et al. 2010) are both needed to solve this problem. Finally, there is evidence that the severe reduction in within-population diversity in NC regions of the genome induced by tight linkage between selected sites has led to elevated FST values, but there is limited support for this effect in C regions.
Purifying selection as a major determinant of population differentiation
Our analysis reveals (i) a positive correlation between KA and F0 (Figure 2) and (ii) a negative correlation between intron length and F calculated using intronic sites (Figure 4). After examining other aspects of polymorphism and differentiation patterns (Figures 1 and 3), we suggest that the observations can be most readily explained by differential intensity of purifying selection acting on different parts of the genome. Similar observations have also been reported in humans (Barreiro et al., 2008; Maruki et al., 2012), suggesting the universal importance of purifying selection as a factor that shapes genetic differentiation between populations.
It should be noted that the above conclusion is not inconsistent with the existence of outlier loci with unusually high F, which may have been caused by diversifying selection (Turner et al., 2008; Yukilevich et al., 2010; Kolaczkowski et al., 2011; Fabian et al., 2012; Langley et al., 2012; Pool et al., 2012; Campo et al., 2013). Our analysis intends to detect forces with large-scale effects (there are typically hundreds of genes in each of the bins in our analysis), and is therefore unlikely to respond to processes that have more localised effects in the genome. In fact, it has been suggested that the number of loci contributing to differences between populations may be relatively small (Yukilevich et al., 2010; Fabian et al., 2012). For example, after taking into account the confounding effects of complex demography and correcting for multiple testing, only four loci had strong statistical support for being driven to high levels of differentiation by diversifying selection between North American and African populations of D. melanogaster (Yukilevich et al., 2010). Furthermore, in line with the low level of linkage disequilibrium in the D. melanogaster genome (for example, Pool et al., 2012), previous genome scan studies have shown that most candidate variants that show evidence of involving in local adaptation only affect differentiation patterns in its immediate neighbourhood, typically on the order of the size of a gene (Kolaczkowski et al., 2011; Fabian et al., 2012). Finally, we have focussed on protein-coding regions and introns, whereas a substantial number of previously found candidate loci fall within intergenic regions.
A noticeable exception is chromosome 3R, in which the cosmopolitan inversion In(3R)P is situated. Multiple studies concerning differentiation between various D. melanogaster populations have found that chromosome 3R has a disproportionally large number of candidate loci, especially within the In(3R)P region, and that these candidate variants tend to affect differentiation patterns in a larger genomic neighbourhood (Kolaczkowski et al., 2011; Fabian et al., 2012). To further test the robustness of our results, we repeated the analysis leading to Figure 2a by removing all genes on chromosome 3R, and found that the pattern remains unchanged (Supplementary Figure S12). In summary, it is unlikely that highly differentiated regions driven by adaptive changes have made a substantial contribution to our observations.
The relationship between F and recombination
As pointed out previously (Charlesworth, 1998; Noor and Bennett, 2009), forces that reduce within-population diversity can lead to elevated FST values in the absence of diversifying selection and restricted gene flow. Hence, in light of the lack of evidence of adaptive evolution in NC regions of the D. melanogaster genome (Betancourt et al., 2009; Arguello et al., 2010; Campos et al., 2014), the high F values obtained from NC regions is probably a result of the diversity-reducing effect of linkage between selected sites, which is often referred to Hill–Robertson interference or HRI (Hill and Robertson, 1966; Comeron et al., 2008; Sella et al., 2009; Charlesworth, 2012a; Cutter and Payseur, 2013). Within the C regions, although negative correlations between F at putatively neutral sites and local recombination rate, as predicted by the HRI theory, were observed (Figure 5, Supplementary Figures S9 and S10), these patterns are weak and often non-significant. Langley et al. (2012) also reported weak negative correlations between a different measure of genetic differentiation and fine-scale recombination rates estimated from linkage disequilibrium patterns, but the relationship was inconsistent between chromosome arms and was sometimes weakly positive when broad-scale recombination rates were used.
The weak association between F and recombination in C regions is somewhat surprising given that both π4 and πSI are clearly positively correlated with local recombination rates in both the RG and FR populations (Supplementary Figure S13). A possible explanation is that, because hitchhiking effects induced by both positive and negative selection can lead to an excess of low-frequency variants at linked neutral sites (Charlesworth et al., 1993; Braverman et al., 1995; Zeng and Charlesworth, 2011), the negative correlation between F and recombination may be weakened, if rare variants are more common in low-recombination regions, as these variants tend to lower F (see Equation (3)). Tajima's D is somewhat more negative in autosomal C regions with reduced recombination (Supplementary Figure S14), but it is hard to determine to what extent this has contributed to the observations in Figure 5 and Supplementary Figure S9, especially when noting that NC regions have more negative Tajima's D and yet higher FST values. Further research that takes into account HRI, demography and statistical properties of estimators of F (see below) is needed to clarify the matter.
The importance of sampling strategy regarding using FST to study population differentiation
As is the case for other definitions of FST (Maruki et al., 2012; Jakobsson et al., 2013), Weir and Cockerham's FST can only take a very restricted range of values when MAF is small (max(FST)⩽2 MAF; Supplementary Figure S1). When information is combined across SNPs, the weighted mean FST (FW) is likely to be dominated by SNPs that are more polymorphic (that is, those having a higher expected MAF). This can lead to the masking of signals of purifying selection, as we have shown above. Thus, FW may be a better choice when the intention is to ascertain the overall level of genetic differentiation. In this case, as long as the data contain a substantial number of putatively neutrally evolving variants, a reasonably accurate estimate can be obtained, even in the presence of sites under strong selective constraints. In contrast, the unweighted mean FST (FU) gives equal weight to all SNPs, and is more responsive to the presence of rare variants (for example, those under purifying selection). These considerations, as well as the recommendations proposed by Bhatia et al. (2013), suggest that care should be exercised when deciding which sampling strategy is most appropriate for the question in hand.
Data Archiving
All genetic data analysed in this study are publically available.
References
Andolfatto P . (2005). Adaptive evolution of non-coding DNA in Drosophila. Nature 437: 1149–1152.
Andolfatto P . (2007). Hitchhiking effects of recurrent beneficial amino acid substitutions in the Drosophila melanogaster genome. Genome Res 17: 1755–1762.
Arguello JR, Zhang Y, Kado T, Fan C, Zhao R, Innan H et al. (2010). Recombination yet inefficient selection along the Drosophila melanogaster subgroup's fourth chromosome. Mol Biol Evol 27: 848–861.
Barreiro LB, Laval G, Quach H, Patin E, Quintana-Murci L . (2008). Natural selection has driven population differentiation in modern humans. Nature Genet 40: 340–345.
Beaumont MA . (2005). Adaptation and speciation: what can FST tell us? Trends Ecol Evol 20: 435–440.
Beaumont MA, Balding DJ . (2004). Identifying adaptive genetic divergence among populations from genome scans. Mol Ecol 13: 969–980.
Beaumont MA, Nichols RA . (1996). Evaluating loci for use in the genetic analysis of population structure. P Roy Soc B-Biol Sci 263: 1619–1626.
Bergman CM, Kreitman M . (2001). Analysis of conserved noncoding DNA in Drosophila reveals similar constraints in intergenic and intronic sequences. Genome Res 11: 1335–1345.
Betancourt AJ, Welch JJ, Charlesworth B . (2009). Reduced effectiveness of selection caused by a lack of recombination. Curr Biol 19: 655–660.
Bhatia G, Patterson N, Sankararaman S, Price AL . (2013). Estimating and interpreting FST: the impact of rare variants. Genome Res 23: 1514–1521.
Braverman JM, Hudson RR, Kaplan NL, Langley CH, Stephan W . (1995). The hitchhiking effect on the site frequency spectrum of DNA polymorphisms. Genetics 140: 783–796.
Campo D, Lehmann K, Fjeldsted C, Souaiaia T, Kao J, Nuzhdin SV . (2013). Whole-genome sequencing of two North American Drosophila melanogaster populations reveals genetic differentiation and positive selection. Mol Ecol 22: 5084–5097.
Campos JL, Charlesworth B, Haddrill PR . (2012). Molecular evolution in nonrecombining regions of the Drosophila melanogaster genome. Genome Biol Evol 4: 278–288.
Campos JL, Halligan DL, Haddrill PR, Charlesworth B . (2014). The relation between recombination rate and patterns of molecular evolution and variation in Drosophila melanogaster. Mol Biol Evol 31: 1010–1028.
Campos JL, Zeng K, Parker DJ, Charlesworth B, Haddrill PR . (2013). Codon usage bias and effective population sizes on the X chromosome versus the autosomes in Drosophila melanogaster. Mol Biol Evol 30: 811–823.
Caracristi G, Schlotterer C . (2003). Genetic differentiation between American and European Drosophila melanogaster populations could be attributed to admixture of African alleles. Mol Biol Evol 20: 792–799.
Carvalho AB, Clark AG . (1999). Intron size and natural selection. Nature 401: 344.
Casillas S, Barbadilla A, Bergman CM . (2007). Purifying selection maintains highly conserved noncoding sequences in Drosophila. Mol Biol Evol 24: 2222–2234.
Charif D, Lobry J . (2007). SeqinR 1.0-2: A contributed package to the R Project for Statistical Computing devoted to biological sequences retrieval and analysis. In: Bastolla U, Porto M, Roman HE, Vendruscolo M (eds). Structural Approaches to Sequence Evolution. Springer: Berlin Heidelberg. pp 207–232.
Charlesworth B . (1998). Measures of divergence between populations and the effect of forces that reduce variability. Mol Biol Evol 15: 538–543.
Charlesworth B . (2001). The effect of life-history and mode of inheritance on neutral genetic variability. Genet Res 77: 153–166.
Charlesworth B . (2012a). The effects of deleterious mutations on evolution at linked sites. Genetics 190: 5–22.
Charlesworth B . (2012b). The role of background selection in shaping patterns of molecular evolution and variation: evidence from variability on the Drosophila X chromosome. Genetics 191: 233–246.
Charlesworth B, Charlesworth D . (2010) Elements of evolutionary genetics. Roberts & Company Publishers: Greenwood Village.
Charlesworth B, Charlesworth D, Barton NH . (2003). The effects of genetic and geographic structure on neutral variation. Annu Rev Ecol Evol Syst 34: 99–125.
Charlesworth B, Morgan MT, Charlesworth D . (1993). The effect of deleterious mutations on neutral molecular variation. Genetics 134: 1289–1303.
Comeron JM, Kreitman M . (2000). The correlation between intron length and recombination in Drosophila. Dynamic equilibrium between mutational and selective forces. Genetics 156: 1175–1190.
Comeron JM, Ratnappan R, Bailin S . (2012). The many landscapes of recombination in Drosophila melanogaster. PLoS Genet 8: e1002905.
Comeron JM, Williford A, Kliman RM . (2008). The Hill-Robertson effect: evolutionary consequences of weak selection and linkage in finite populations. Heredity 100: 19–31.
Cutter AD, Choi JY . (2010). Natural selection shapes nucleotide polymorphism across the genome of the nematode Caenorhabditis briggsae. Genome Res 20: 1103–1111.
Cutter AD, Payseur BA . (2013). Genomic signatures of selection at linked sites: unifying the disparity among species. Nat Rev Genet 14: 262–274.
David JR, Capy P . (1988). Genetic variation of Drosophila melanogaster natural populations. Trends Genet 4: 106–111.
Duchen P, Zivkovic D, Hutter S, Stephan W, Laurent S . (2013). Demographic inference reveals African and European admixture in the North American Drosophila melanogaster population. Genetics 193: 291–301.
Excoffier L, Hofer T, Foll M . (2009). Detecting loci under selection in a hierarchically structured population. Heredity 103: 285–298.
Excoffier L, Smouse PE, Quattro JM . (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131: 479–491.
Eyre-Walker A, Keightley PD . (2007). The distribution of fitness effects of new mutations. Nat Rev Genet 8: 610–618.
Eyre-Walker A, Keightley PD . (2009). Estimating the rate of adaptive molecular evolution in the presence of slightly deleterious mutations and population size change. Mol Biol Evol 26: 2097–2108.
Fabian DK, Kapun M, Nolte V, Kofler R, Schmidt PS, Schlotterer C et al. (2012). Genome-wide patterns of latitudinal differentiation among populations of Drosophila melanogaster from North America. Mol Ecol 21: 4748–4769.
Fiston-Lavier AS, Singh ND, Lipatov M, Petrov DA . (2010). Drosophila melanogaster recombination rate calculator. Gene 463: 18–20.
Foll M, Gaggiotti O . (2008). A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180: 977–993.
Fu W, O'Connor TD, Jun G, Kang HM, Abecasis G, Leal SM et al. (2013). Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493: 216–220.
Haddrill PR, Charlesworth B, Halligan DL, Andolfatto P . (2005a). Patterns of intron sequence evolution in Drosophila are dependent upon length and GC content. Genome Biol 6: R67.
Haddrill PR, Halligan DL, Tomaras D, Charlesworth B . (2007). Reduced efficacy of selection in regions of the Drosophila genome that lack crossing over. Genome Biol 8: R18.
Haddrill PR, Thornton KR, Charlesworth B, Andolfatto P . (2005b). Multilocus patterns of nucleotide variability and the demographic and selection history of Drosophila melanogaster populations. Genome Res 15: 790–799.
Haddrill PR, Zeng K, Charlesworth B . (2011). Determinants of synonymous and nonsynonymous variability in three species of Drosophila. Mol Biol Evol 28: 1731–1743.
Halligan DL, Keightley PD . (2006). Ubiquitous selective constraints in the Drosophila genome revealed by a genome-wide interspecies comparison. Genome Res 16: 875–884.
Hershberg R, Petrov DA . (2008). Selection on codon bias. Annu Rev Genet 42: 287–299.
Hey J, Machado CA . (2003). The study of structured populations—new hope for a difficult and divided science. Nat Rev Genet 4: 535–543.
Hill WG, Robertson A . (1966). The effect of linkage on limits to artificial selection. Genet Res 8: 269–294.
Holsinger KE, Weir BS . (2009). Genetics in geographically structured populations: defining, estimating and interpreting FST . Nat Rev Genet 10: 639–650.
Hu TT, Eisen MB, Thornton KR, Andolfatto P . (2013). A second-generation assembly of the Drosophila simulans genome provides new insights into patterns of lineage-specific divergence. Genome Res 23: 89–98.
Hudson RR, Slatkin M, Maddison WP . (1992). Estimation of levels of gene flow from DNA sequence data. Genetics 132: 583–589.
Hutter S, Li H, Beisswanger S, De Lorenzo D, Stephan W . (2007). Distinctly different sex ratios in African and European populations of Drosophila melanogaster inferred from chromosomewide single nucleotide polymorphism data. Genetics 177: 469–480.
Jakobsson M, Edge MD, Rosenberg NA . (2013). The relationship between FST and the frequency of the most frequent allele. Genetics 193: 515–528.
Kaiser VB, Charlesworth B . (2009). The effects of deleterious mutations on evolution in non-recombining genomes. Trends Genet 25: 9–12.
Katoh K, Misawa K, Kuma K, Miyata T . (2002). MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30: 3059–3066.
Keightley PD, Eyre-Walker A . (2007). Joint inference of the distribution of fitness effects of deleterious mutations and population demography based on nucleotide polymorphism frequencies. Genetics 177: 2251–2261.
Keightley PD, Trivedi U, Thomson M, Oliver F, Kumar S, Blaxter ML . (2009). Analysis of the genome sequences of three Drosophila melanogaster spontaneous mutation accumulation lines. Genome Res 19: 1195–1201.
Keinan A, Mullikin JC, Patterson N, Reich D . (2007). Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nat Genet 39: 1251–1255.
Keinan A, Mullikin JC, Patterson N, Reich D . (2009). Accelerated genetic drift on chromosome X during the human dispersal out of Africa. Nat Genet 41: 66–70.
Kimura M . (1980). A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16: 111–120.
Kimura M . (1983) The Neutral Theory of Molecular Evolution. Cambridge University Press: Cambridge.
Kolaczkowski B, Kern AD, Holloway AK, Begun DJ . (2011). Genomic differentiation between temperate and tropical Australian populations of Drosophila melanogaster. Genetics 187: 245–260.
Langley CH, Stevens K, Cardeno C, Lee YC, Schrider DR, Pool JE et al. (2012). Genomic variation in natural populations of Drosophila melanogaster. Genetics 192: 533–598.
Li H, Stephan W . (2006). Inferring the demographic history and rate of adaptive substitution in Drosophila. PLoS Genet 2: e166.
Li WH . (1993). Unbiased Estimation of the Rates of Synonymous and Nonsynonymous Substitution. J Mol Evol 36: 96–99.
Maruki T, Kumar S, Kim Y . (2012). Purifying selection modulates the estimates of population differentiation and confounds genome-wide comparisons across single-nucleotide polymorphisms. Mol Biol Evol 29: 3617–3623.
Nagylaki T . (1998). Fixation indices in subdivided populations. Genetics 148: 1325–1332.
Nei M . (1973). Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70: 3321–3323.
Nei M, Miller JC . (1990). A simple method for estimating average number of nucleotide substitutions within and between populations from restriction data. Genetics 125: 873–879.
Nelson MR, Wegmann D, Ehm MG, Kessner D St, Jean P, Verzilli C et al. (2012). An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 people. Science 337: 100–104.
Noor MA, Bennett SM . (2009). Islands of speciation or mirages in the desert? Examining the role of restricted recombination in maintaining species. Heredity 103: 439–444.
O'Fallon BD, Seger J, Adler FR . (2010). A continuous-state coalescent and the impact of weak selection on the structure of gene genealogies. Mol Biol Evol 27: 1162–1172.
Ometto L, Stephan W, De Lorenzo D . (2005). Insertion/deletion and nucleotide polymorphism data reveal contraints in Drosophila melanogaster introns and intergenic regions. Genetics 169: 1521–1527.
Pal C, Papp B, Lercher MJ . (2006). An integrated view of protein evolution. Nat Rev Genet 7: 337–348.
Paradis E . (2010). pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26: 419–420.
Paradis E, Claude J, Strimmer K . (2004). APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20: 289–290.
Parsch J . (2003). Selective constraints on intron evolution in Drosophila. Genetics 165: 1843–1851.
Parsch J, Novozhilov S, Saminadin-Peter SS, Wong KM, Andolfatto P . (2010). On the utility of short intron sequences as a reference for the detection of positive and negative selection in Drosophila. Mol Biol Evol 27: 1226–1234.
Peden JF . (1999) Analysis of codon usage. PhD thesis, University of Nottingham, UK. Available from http://codonw.source-forge.net/.
Pool JE, Corbett-Detig RB, Sugino RP, Stevens KA, Cardeno CM, Crepeau MW et al. (2012). Population Genomics of sub-saharan Drosophila melanogaster: African diversity and non-African admixture. PLoS Genet 8: e1003080.
Pool JE, Nielsen R . (2007). Population size changes reshape genomic patterns of diversity. Evolution 61: 3001–3006.
Pool JE, Nielsen R . (2008). The impact of founder events on chromosomal variability in multiply mating species. Mol Biol Evol 25: 1728–1736.
Roy S, Ernst J, Kharchenko PV, Kheradpour P, Negre N, Eaton ML et al. (2010). Identification of Functional Elements and Regulatory Circuits by Drosophila modENCODE. Science 330: 1787–1797.
Schaeffer SW . (2002). Molecular population genetics of sequence length diversity in the Adh region of Drosophila pseudoobscura. Genet Res 80: 163–175.
Schneider A, Charlesworth B, Eyre-Walker A, Keightley PD . (2011). A method for inferring the rate of occurrence and fitness effects of advantageous mutations. Genetics 189: 1427–1437.
Seger J, Smith WA, Perry JJ, Hunn J, Kaliszewska ZA, Sala LL et al. (2010). Gene genealogies strongly distorted by weakly interfering mutations in constant environments. Genetics 184: 529–545.
Sella G, Petrov DA, Przeworski M, Andolfatto P . (2009). Pervasive natural selection in the Drosophila genome? PLoS Genet 5: e1000495.
Singh ND, Arndt PF, Petrov DA . (2005). Genomic heterogeneity of background substitutional patterns in Drosophila melanogaster. Genetics 169: 709–722.
Singh ND, Macpherson JM, Jensen JD, Petrov DA . (2007). Similar levels of X-linked and autosomal nucleotide variation in African and non-African populations of Drosophila melanogaster. BMC Evol Biol 7: 202.
Slatkin M . (1995). A measure of population subdivision based on microsatellite allele frequencies. Genetics 139: 457–462.
Stephan W, Li H . (2007). The recent demographic and adaptive history of Drosophila melanogaster. Heredity 98: 65–68.
Tajima F . (1989). Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123: 585–595.
Thornton K, Andolfatto P . (2006). Approximate Bayesian inference reveals evidence for a recent, severe bottleneck in a Netherlands population of Drosophila melanogaster. Genetics 172: 1607–1619.
Turner TL, Levine MT, Eckert ML, Begun DJ . (2008). Genomic analysis of adaptive differentiation in Drosophila melanogaster. Genetics 179: 455–473.
Weir BS, Cockerham CC . (1984). Estimating F-Statistics for the Analysis of Population-Structure. Evolution 38: 1358–1370.
Weir BS, Hill WG . (2002). Estimating F-statistics. Annu Rev Genet 36: 721–750.
Wright S . (1951). The Genetical Structure of Populations. Ann Eugenic 15: 323–354.
Wu CI . (2001). The genic view of the process of speciation. J Evolution Biol 14: 851–865.
Yukilevich R, Turner TL, Aoki F, Nuzhdin SV, True JR . (2010). Patterns and processes of genome-wide divergence between North American and African Drosophila melanogaster. Genetics 186: 219–239.
Zeng K . (2013). A coalescent model of background selection with recombination, demography and variation in selection coefficients. Heredity 110: 363–371.
Zeng K, Charlesworth B . (2009). Estimating selection intensity on synonymous codon usage in a nonequilibrium population. Genetics 183: 651–662.
Zeng K, Charlesworth B . (2010). The effects of demography and linkage on the estimation of selection and mutation parameters. Genetics 186: 1411–1424.
Zeng K, Charlesworth B . (2011). The joint effects of background selection and genetic recombination on local gene genealogies. Genetics 189: 251–266.
Acknowledgements
We are grateful to the Drosophila Population Genomics Project for making these data available. We also thank Brian Charlesworth and three anonymous reviewers for useful discussions and comments. BCJ would like to thank Quentin Geissmann for time spent helping him to become a better programmer. This work was supported by a PhD studentship to BCJ, jointly funded by the National Environmental Research Council (NERC) (NE/H524881/1 and NE/K500914/1) and the Department of Animal and Plant Sciences, University of Sheffield; and a Biotechnology and Biological Sciences Research Council (BBSRC) grant (BB/K000209/1) awarded to KZ. This work has made use of High Performance Computing Cluster, Iceberg, at the University of Sheffield.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Jackson, B., Campos, J. & Zeng, K. The effects of purifying selection on patterns of genetic differentiation between Drosophila melanogaster populations. Heredity 114, 163–174 (2015). https://doi.org/10.1038/hdy.2014.80
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2014.80
- Springer Nature Switzerland AG
This article is cited by
-
Genomic differentiation across the speciation continuum in three hummingbird species pairs
BMC Evolutionary Biology (2020)
-
Genetics and evidence for balancing selection of a sex-linked colour polymorphism in a songbird
Nature Communications (2019)