Abstract
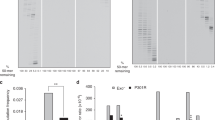
ERRORS in the replication of DNA are a major source of spontaneous mutations, and a number of cellular functions are involved in correction of these errors to keep the frequency of spontaneous mutations very low1. We report here a novel mechanism which prevents replicational errors by degrading a potent mutagenic substrate for DNA synthesis. This error-avoiding process is catalysed by a protein encoded by themutT gene of Escherichia coli, mutations of which increase the occurrence of A · T-→ C · G transversions 100 to 10,000 times the level of the wild type2. Spontaneous oxidation of dGTP forms 8-oxo-7,8-dihydro-2'-dGTP (8-oxodGTP), which is inserted opposite dA and dC residues of template DNA with almost equal efficiency, and the MutT protein specifically degrades 8-oxodGTP to the monophosphate. This indicates that elimination from the nucleotide pool of the oxidized form of guanine nucleotide is important for the high fidelity of DNA synthesis.
Similar content being viewed by others
References
Horiuchi, T., Maki, H. & Sekiguchi, M. Bull. Inst. Pasteur 87, 309–336 (1989).
Yanofsky, C., Cox, E. C. & Horn, V. Proc. natn. Acad. Sci. U.S.A. 55, 274–281 (1966).
Akiyama, M., Maki, H., Sekiguchi, M. & Horiuchi, T. Proc. natn. Acad. Sci. U.S.A. 86, 3949–3952 (1989).
Bhatnagar, S. K. & Bessman, M. J. biol. Chem. 263, 8953–8957 (1988).
Kasai, H. & Nishimura, S. Nucleic Acids Res. 12, 2137–2145 (1984).
Kasai, H. & Nishimura, S. in Oxidative Stress: Oxidants and Antioxidants (ed. Sies, H.), 99–116 (Academic, London, 1991).
Shibutani, S., Takeshita, M. & Grollman, A. P. Nature 349, 431–434 (1991).
Maki, H. & Kornberg, A. J. biol. Chem. 260, 12987–12992 (1985).
Sloane, D. L., Goodman, M. F. & Echols, H. Nucleic Acids Res. 16, 6465–6475 (1988).
Kasai, H., Tanooka, H. & Nishimura, S. Gann 75, 1037–1039 (1984).
Wood, M. L., Dizdaroglu, M., Gajewski, E. & Essigman, J. M. Biochemistry 29, 7024–7032 (1990).
Moriya, M. et al. Mutation Res. 254, 281–288 (1991).
Tchou, J. et al. Proc. natn. Acad. Sci. U.S.A. 88, 4690–4694 (1991).
Cabrera, M., Nghiem, Y. & Miller, J. H. J. Bact. 170, 5405–5407 (1988).
Michaels, M. L., Pham, L., Cruz, C. & Miller, J. Nucleic Acids Res. 19, 3629–3632 (1988).
Boosalis, M. S., Petruska, J. & Goodman, M. F. J. biol. Chem. 262, 14689–14696 (1987).
Fersht, A. in Enzyme Structure and Mechanism 91–92 (Freeman, San Francisco, 1977).
Akiyama, M., Horiuchi, T. & Sekiguchi, M. Molec. gen. Genet. 206, 9–16 (1987).
Bradford, M. M. Analyt. Biochem. 72, 248–254 (1976).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Maki, H., Sekiguchi, M. MutT protein specifically hydrolyses a potent mutagenic substrate for DNA synthesis. Nature 355, 273–275 (1992). https://doi.org/10.1038/355273a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/355273a0
- Springer Nature Limited
This article is cited by
-
MiR-146a rs2910164 (G/C) polymorphism is associated with the development and prognosis of acute coronary syndromes: an observational study including case control and validation cohort
Journal of Translational Medicine (2023)
-
MTH1 protects platelet mitochondria from oxidative damage and regulates platelet function and thrombosis
Nature Communications (2023)
-
Bacterial DNA excision repair pathways
Nature Reviews Microbiology (2022)
-
Impact of Zero-Valent Iron on Freshwater Bacterioplankton Metabolism as Predicted from 16S rRNA Gene Sequence Libraries
Current Microbiology (2021)
-
Reactive Oxygen Species-Related Nanoparticle Toxicity in the Biomedical Field
Nanoscale Research Letters (2020)