Abstract
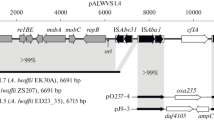
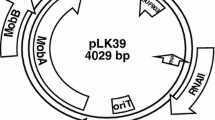
Results of studying plasmid pAH36 in strain Aeronomas hydrophila IBRB-36 4CPA are presented. Plasmid pAH36 possesses BamHI, PstI, and HindIII restriction sites and is 5.4 kb in size. The plasmid was shown to contain genes for catabolism of chlor-substituted phenoxyacetic acids.
Similar content being viewed by others
REFERENCES
Bergey's Manual of Determinative Bacteriology, Kreig, N.R. and Holt, J.G., Eds., Baltimore: Williams Wilkins, 1994.
Manual of Methods for General Bacteriology, Gerhard, P., Murray, R.G.E., Costilow, R.N., et al., Eds., Washington: Am. Soc. Microbiol., 1981.
Maniatis T., Fritsch, E.E., and Sambrook, J., Molecular Cloning: A Laboratory Manual, Cold Spring Harbor, New York: Cold Spring Harbor Lab., 1982.
Metody opredeleniya mikrokolichestv pestitsidov (Methods for Determining Pesticide Traces), Moscow: Meditsina, 1984.
Vedler, E., Koiv, V., and Heinaru, A., TfdR, the LysR-Type Transcriptional Activator, Is Responsible for the Activation of the tfdCB Operon of Pseudomonas putida 2,4-Dichlorophenoxyacetic Acid Degradative Plasmid pEST4011, Gene, 2000, vol. 245, no. 1, pp. 161–168.
Don, R.H. and Weightman, A.J., Transposon Mutagenesis and Cloning Analysis of the Pathways for Degradation of 2,4-Dichlorophenoxyacetic Acid and 3-Chlo-robenzoate in Alcaligenes eutrophus JMP134 (pJP4), J. Bacteriol., 1985, vol. 161, pp. 85–90.
Harker, A.R., Olsen, R.H., and Seidler, R.J., Phenoxy-acetic Acid Degradation by the 2,4-Dichlorophenoxy-acetic Acid (TFD) Pathway of Plasmid JP4: Mapping and Characterization of the 2,4-D Regulatory Gene tfdR, J. Bacteriol., 1989, vol. 171, no. 1, pp. 314–320.
Sangodkar, U.M.X., Aldrich, T.L., Haugland, R.A., et al., Molecular Basis of Biodegradation of Chloroaromatic Compounds, Acta Biotechnol., 1989, vol. 9, no. 4, pp. 301–316.
Perkins, E.J., Gordon, M.P., Caceres, O., and Lurquin, P.F., Organization and Sequence Analysis of the 2,4-Dichlo-rophenol Hydroxylase and Dichlorocatechol Oxidative Operons of Plasmid pJP4, J. Bacteriol., 1990, vol. 172, no. 5, pp. 2351–2359.
Balajee, S. and Mahadevan, A., Influence of Chloroaro-matic Substances on the Biological Activity of Azoto-bacter chroococcum, Chemosphere, 1990, vol. 21, nos. 1–2, pp. 51–56.
Daubaras, D.L., Saido, K., and Chakrabarty, A.M., Purification of Hydroxyquinol and Maleylacetate Reductase: The Lower Pathway of 2,4,5-Trichlorophenoxyacetic Acid Metabolism by Burkholderia cepacia AC1100, Appl. Environ. Microbiol., 1996, vol. 62, no. 11, pp. 4276–4279.
Bhat, M.A., Tsuda, M., Horiike, K., et al., Identification and Characterization of a New Plasmid Carrying Genes for Degradation of 2,4-Dichlorophenoxyacetate from Pseudomonas cepacia CSV90, Appl. Environ. Microbiol., 1994, vol. 60, no. 1, pp. 307–312.
Short, K.A., Seidler, R.J., and Olsen, R.H., Survial and Degradative Capacity of Pseudomonas putida Induced or Constitutively Expressing Plasmid-Mediated Degradation of 2.4-Dichlorophenoxyacetate (TFD) in Soil, Can. J. Microbiol., 1990, vol. 36, pp. 821–829.
Herrmann, H., Muller, C., Schmidt, I., et al., Localization and Organization of Phenol Degradation Genes of Pseudomonas putida Strain H, Mol. Gen. Genet., 1995, vol. 247, no. 2, pp. 240–246.
Ausmees, N.R. and Neinaru, A.L., New Plasmids Determining Biodegradation of Herbicide 2,4-D, Genetika (Moscow), 1990, vol. 26, no. 4, pp. 770–772.
Markusheva, T.V., Zhurenko, E.Yu., and Kusova, I.V., Bakterii-destruktory fenola i ego khlorirovannykh proiz-vodnykh (Bacteria Destructing Phenol and Its Chorinated Derivatives), Ufa: Gilem, 2002.
Neilson, A.H., The Biodegradation of Halogenated Organic Compounds, J. Appl. Bacteriol., 1990, vol. 69, pp. 445–470.
Perez-Pantoja, D., Guzmin, L., Manzano, M., et al., Role of tfdC(I)D(I)E(I)F(I) and tfdD(II)C(II)E(II)F(II) Gene Modules in Catabolism of 3-Chlorobenzoate by Ralsto-nia eutropha JMP134 (pJP4), Appl. Environ. Microbiol., 2000, vol. 66, no. 4, pp. 1602–1608.
Top, E.M., Maltseva, O.V., and Forney, L.J., Capture of a Catabolic Plasmid That Encodes Only 2,4-Dichlo-rophenoxyacetic Acid: α-Ketoglutaric Acid Dioxygenase (TfdA) by Genetic Complementation, Appl. Environ. Microbiol., 1996, vol. 62, no. 7, pp. 2470–2476.
Golovleva, L.A. and Pertsova, R.N., Complete Degradation and Dechlorination of 2,4,5-Trichlorophenoxyace-tic Acid by Nocardioides simplex Strain 3E, Dokl. Akad. Nauk SSSR, 1990, vol. 314, no. 4, pp. 981–983.
Arai, H., Akahira, S., Ohishi, T., et al., Adaptation of Comamonas testosteroni TA441 to Utilize Phenol: Organization and Regulation of the Genes Involved in Phenol Degradation, Microbiology, 1998, vol. 144, pp. 2895–2903.
Kulakova, A.N., Kulakov, L.A., Naumov, A.V., et al., Cloning of the Phenol Degradation Genes from Pseudomonas sp. 5-T, Genetika (Moscow), 1992, vol. 28, no. 2, pp. 35–42.
Nordlund, I., Powlowski, J., Hagstrom, A., and Shingler, V., Conservation of Regulatory and Structural Genes for Multicomponent Phenol Hydroxylase within Phenol-Catabolizing Bacteria That Utilize a Meta-Cleavage Pathway, J. Gen. Microbiol., 1993, vol. 139, pp. 2695–2703.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Markusheva, T.V., Zhurenko, E.Y., Galkin, E.G. et al. Identification and Characterization of a Plasmid in Strain Aeronomas hydrophila IBRB-36 4CPA Carrying Genes for Catabolism of Chlorophenoxyacetic Acids. Russian Journal of Genetics 40, 1210–1214 (2004). https://doi.org/10.1023/B:RUGE.0000048662.23804.41
Issue Date:
DOI: https://doi.org/10.1023/B:RUGE.0000048662.23804.41